Abstract
Family study results are consistent with genetic effects making substantial contributions to risk of psychiatric disorders such as schizophrenia, yet robust identification of specific genetic variants that explain variation in population risk had been disappointing until the advent of technologies that assay the entire genome in large samples. We highlight recent progress that has led to a better understanding of the number of risk variants in the population and the interaction of allele frequency and effect size. The emerging genetic architecture implies a large number of contributing loci (that is, a high genome-wide mutational target) and suggests that genetic risk of psychiatric disorders involves the combined effects of many common variants of small effect, as well as rare and de novo variants of large effect. The capture of a substantial proportion of genetic risk facilitates new study designs to investigate the combined effects of genes and the environment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Lichtenstein, P., Carlstrom, E., Rastam, M., Gillberg, C. & Anckarsater, H. The genetics of autism spectrum disorders and related neuropsychiatric disorders in childhood. Am. J. Psychiatry 167, 1357–1363 (2010).
Lichtenstein, P. et al. Common genetic determinants of schizophrenia and bipolar disorder in Swedish families: a population-based study. Lancet 373, 234–239 (2009).
Ronald, A. & Hoekstra, R.A. Autism spectrum disorders and autistic traits: a decade of new twin studies. Am. J. Med. Genet. B. Neuropsychiatr. Genet. 156B, 255–274 (2011).
Sullivan, P.F., Kendler, K.S. & Neale, M.C. Schizophrenia as a complex trait: evidence from a meta-analysis of twin studies. Arch. Gen. Psychiatry 60, 1187–1192 (2003).
Wray, N.R. & Gottesman, I.I. Using summary data from the Danish national registers to estimate heritabilities for schizophrenia, bipolar disorder, and major depressive disorder. Front. Genet. 3, 118 (2012).
Eyre-Walker, A. Evolution in health and medicine Sackler colloquium: Genetic architecture of a complex trait and its implications for fitness and genome-wide association studies. Proc. Natl. Acad. Sci. USA 107 Suppl 1, 1752–1756 (2010).
Keller, M.C. & Miller, G. Resolving the paradox of common, harmful, heritable mental disorders: which evolutionary genetic models work best? Behav. Brain Sci. 29, 385–452 (2006).
McClellan, J. & King, M.-C. Genetic heterogeneity in human disease. Cell 141, 210–217 (2010).
Hill, W.G., Goddard, M.E. & Visscher, P.M. Data and theory point to mainly additive genetic variance for complex traits. PLoS Genet. 4, e1000008 (2008).
Corvin, A., Craddock, N. & Sullivan, P.F. Genome-wide association studies: a primer. Psychol. Med. 40, 1063–1077 (2010).
Visscher, P.M., Brown, M.A., McCarthy, M.I. & Yang, J. Five years of GWAS discovery. Am. J. Hum. Genet. 90, 7–24 (2012).
McCarroll, S.A., Feng, G. & Hyman, S.E. Genome-scale neurogenetics: methodology and meaning. Nat. Neurosci. 17, 756–763 (2014).
Psychiatric Genomics Consortium Schizophrenia Working Group. Psychiatric Genomics Consortium quadruples schizophrenia GWAS sample-size to 35,000 cases and 47,000 controls. in XXIst World Congress of Psychiatric Genetics: Redefining Mental Illness Through Genetics (Boston, 2013). This conference paper reported the identification of more than 100 loci for schizophrenia, the 'flagship' disorder of the Psychiatric Genomics Consortium. The results show what might be achieved for other psychiatric disorders as sample sizes increase.
European Alzheimer's Disease Initiative et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer's disease. Nat. Genet. 45, 1452–1458 (2013).
Charney, A.W. et al. Bipolar disorder GWAS of 13,741 cases and 19,762 controls identifies eight genome-wide significant hits and implicates genes targeted by FMRP protein. in XXIst World Congress of Psychiatric Genetics: Redefining Mental Illness Through Genetics (Boston, 2013).
Anney, R. et al. Meta-analysis of European ancestry individuals with autism spectrum disorder reveals strong association 3′ of the astrotactin 2 (ASTN2) gene locus on chromosome 9. in XXIst World Congress of Psychiatric Genetics: Redefining Mental Illness Through Genetics (Boston, 2013).
Neale, B.M. et al. Case-control genome-wide association study of attention-deficit/hyperactivity disorder. J. Am. Acad. Child Adolesc. Psychiatry 49, 906–920 (2010).
Wang, K. et al. A genome-wide association study on common SNPs and rare CNVs in anorexia nervosa. Mol. Psychiatry 16, 949–959 (2011).
Ripke, S. et al. A mega-analysis of genome-wide association studies for major depressive disorder. Mol. Psychiatry 18, 497–511 (2013).
Davis, L.K. et al. Partitioning the heritability of tourette syndrome and obsessive compulsive disorder reveals differences in genetic architecture. PLoS Genet. 9, e1003864 (2013).
Schizophrenia Psychiatric Genome-Wide Association Study (GWAS) Consortium et al. Genome-wide association study identifies five new schizophrenia loci. Nat. Genet. 43, 969–976 (2011).
Cross-Disorder Group of the Psychiatric Genomics Consortium et al. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet 381, 1371–1379 (2013).
Pe'er, I., Yelensky, R., Altshuler, D. & Daly, M.J. Estimation of the multiple testing burden for genome-wide association studies of nearly all common variants. Genet. Epidemiol. 32, 381–385 (2008).
Sullivan, P.F. The psychiatric GWAS consortium: big science comes to psychiatry. Neuron 68, 182–186 (2010).
Wray, N.R. et al. Genome-wide association study of major depressive disorder: new results, meta-analysis and lessons learned. Mol. Psychiatry 17, 36–48 (2012).
Purcell, S.M. et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460, 748–752 (2009). This is a landmark study that demonstrated a substantial contribution to risk of schizophrenia from common variation distributed throughout the genome. It also revealed evidence for genetic overlap between schizophrenia and bipolar disorder involving many common loci of small effect.
Shi, J. et al. Common variants on chromosome 6p22.1 are associated with schizophrenia. Nature 460, 753–757 (2009).
Ripke, S. et al. Genome-wide association analysis identifies 13 new risk loci for schizophrenia. Nat. Genet. 45, 1150–1159 (2013).
Anderson-Schmidt, H. et al. Selected rapporteur summaries from the XX World Congress of Psychiatric Genetics, Hamburg, Germany, October 14–18, 2012. Am. J. Med. Genet. B. Neuropsychiatr. Genet. 162B, 96–121 (2013).
Corder, E.H. et al. Gene dose of apolipoprotein E type 4 allele and the risk of Alzheimer's disease in late onset families. Science 261, 921–923 (1993).
Klein, R.J. et al. Complement factor H polymorphism in age-related macular degeneration. Science 308, 385–389 (2005).
Wellcome Trust Case Control Consortium. Genome-wide association study of 14,000 cases of seven common diseases and 3,000 shared controls. Nature 447, 661–678 (2007).
Sullivan, P. Don't give up on GWAS. Mol. Psychiatry 17, 2–3 (2012).
Morris, A.P. et al. Large-scale association analysis provides insights into the genetic architecture and pathophysiology of type 2 diabetes. Nat. Genet. 44, 981–990 (2012).
Plenge, R.M., Scolnick, E.M. & Altshuler, D. Validating therapeutic targets through human genetics. Nat. Rev. Drug Discov. 12, 581–594 (2013).
Sanseau, P. et al. Use of genome-wide association studies for drug repositioning. Nat. Biotechnol. 30, 317–320 (2012).
Okada, Y. et al. Genetics of rheumatoid arthritis contributes to biology and drug discovery. Nature 506, 376–381 (2014).
Teslovich, T.M. et al. Biological, clinical and population relevance of 95 loci for blood lipids. Nature 466, 707–713 (2010).
Jostins, L. et al. Host-microbe interactions have shaped the genetic architecture of inflammatory bowel disease. Nature 491, 119–124 (2012).
Lango Allen, H. et al. Hundreds of variants clustered in genomic loci and biological pathways affect human height. Nature 467, 832–838 (2010).
Gilman, S.R. et al. Rare de novo variants associated with autism implicate a large functional network of genes involved in formation and function of synapses. Neuron 70, 898–907 (2011).
Sebat, J. et al. Strong association of de novo copy number mutations with autism. Science 316, 445–449 (2007).
Kirov, G. et al. De novo CNV analysis implicates specific abnormalities of postsynaptic signaling complexes in the pathogenesis of schizophrenia. Mol. Psychiatry 17, 142–153 (2012).
Xu, B. et al. Strong association of de novo copy number mutations with sporadic schizophrenia. Nat. Genet. 40, 880–885 (2008).
Williams, N.M. et al. Rare chromosomal deletions and duplications in attention-deficit hyperactivity disorder: a genome-wide analysis. Lancet 376, 1401–1408 (2010).
Malhotra, D. et al. High frequencies of de novo CNVs in bipolar disorder and schizophrenia. Neuron 72, 951–963 (2011).
Bergen, S.E. et al. Genome-wide association study in a Swedish population yields support for greater CNV and MHC involvement in schizophrenia compared with bipolar disorder. Mol. Psychiatry 17, 880–886 (2012).
Levinson, D.F. et al. Copy number variants in schizophrenia: confirmation of five previous findings and new evidence for 3q29 microdeletions and VIPR2 duplications. Am. J. Psychiatry 168, 302–316 (2011). This is the largest meta-analysis of copy number variation in schizophrenia.
Sanders, S.J. et al. Multiple recurrent de novo CNVs, including duplications of the 7q11.23 Williams syndrome region, are strongly associated with autism. Neuron 70, 863–885 (2011).
Vacic, V. et al. Duplications of the neuropeptide receptor gene VIPR2 confer significant risk for schizophrenia. Nature 471, 499–503 (2011).
Williams, N.M. et al. Genome-wide analysis of copy number variants in attention deficit hyperactivity disorder: the role of rare variants and duplications at 15q13.3. Am. J. Psychiatry 169, 195–204 (2012).
Crespi, B., Stead, P. & Elliot, M. Evolution in health and medicine Sackler colloquium: comparative genomics of autism and schizophrenia. Proc. Natl. Acad. Sci. USA 107 Suppl 1, 1736–1741 (2010).
Levy, D. et al. Rare de novo and transmitted copy-number variation in autistic spectrum disorders. Neuron 70, 886–897 (2011).
Iossifov, I. et al. De novo gene disruptions in children on the autistic spectrum. Neuron 74, 285–99 (2012). This is the largest of four recent studies reporting the results of WES in autism families, and the only study other than Sanders et al . (2012) to use a large sample of family controls.
Neale, B.M. et al. Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature 485, 242–245 (2012). This is one of four studies reporting the results of WES in autism families. Collectively, these studies have identified six brain-expressed genes harboring recurrent de novo gene-disrupting mutations that are now strong candidate genes for autism.
O'Roak, B.J. et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Nature 485, 246–50 (2012). This is one of four studies reporting the results of WES in autism families. The authors report recurrent gene-disrupting mutations in the CHD8 gene in unrelated probands, an observation that is highly unlikely to have occurred by chance.
Sanders, S.J. et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature 485, 237–241 (2012). This study, one of four that report results from WES in autism families, used a discordant sibling family design to demonstrate an enrichment of de novo gene-disrupting mutations in autism.
Fromer, M. et al. De novo mutations in schizophrenia implicate synaptic networks. Nature 506, 179–184 (2014). This is the largest WES study of de novo mutations in schizophrenia to date. The authors report an enrichment of de novo mutations in genes encoding for post-synaptic proteins.
Purcell, S. et al. A polygenic burden of rare disruptive mutations in schizophrenia. Nature 506, 185–190 (2014). Together with the parallel family-based study by Fromer et al . (2014)58, this large case-control WES study demonstrated a burden of rare mutations in specific sets of post-synaptic genes in schizophrenia.
Xu, B. et al. De novo gene mutations highlight patterns of genetic and neural complexity in schizophrenia. Nat. Genet. 44, 1365–1369 (2012).
Hoischen, A., Krumm, N. & Eichler, E.E. Prioritization of neurodevelopmental disease genes by discovery of new mutations. Nat. Neurosci. 17, 764–772 (2014).
de Ligt, J. et al. Diagnostic exome sequencing in persons with severe intellectual disability. N. Engl. J. Med. 367, 1921–1929 (2012).
Kong, A. et al. Rate of de novo mutations and the importance of father's age to disease risk. Nature 488, 471–475 (2012). This paper demonstrated that the majority of de novo point mutations originate in the paternal germline, and that the de novo mutation rate is positively correlated with paternal age.
Michaelson, J.J. et al. Whole-genome sequencing in autism identifies hot spots for de novo germline mutation. Cell 151, 1431–1442 (2012).
McGrath, J.J. et al. A comprehensive assessment of parental age and psychiatric disorders. JAMA Psychiatry 71, 301–309 (2014).
Petersen, L., Mortensen, P.B. & Pedersen, C.B. Paternal age at birth of first child and risk of schizophrenia. Am. J. Psychiatry 168, 82–88 (2011).
Perrin, M.C., Brown, A.S. & Malaspina, D. Aberrant epigenetic regulation could explain the relationship of paternal age to schizophrenia. Schizophr. Bull. 33, 1270–1273 (2007).
Samocha, K.E., Robinson, E., Neale, B. & Daly, M. Statistical evaluation of de novo variation implicates a distinct etiologic subtype of autism. in XXIst World Congress of Psychiatric Genetics: Redefining Mental Illness Through Genetics (Boston, 2013).
Liu, L. et al. Analysis of rare, exonic variation amongst subjects with autism spectrum disorders and population controls. PLoS Genet. 9, e1003443 (2013).
Ionita-Laza, I. et al. Scan statistic-based analysis of exome sequencing data identifies FAN1 at 15q13.3 as a susceptibility gene for schizophrenia and autism. Proc. Natl. Acad. Sci. USA 111, 343–348 (2014).
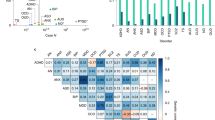
Lee, S.H. et al. Genetic relationship between five psychiatric disorders estimated from genome-wide SNPs. Nat. Genet. 45, 984–994 (2013). This paper reported evidence for overlap of common genetic variation (that is, pleiotropy) between key psychiatric disorders, including SCZ and BIP (high), SCZ and MDD (moderate), MDD and BIP (moderate), MDD and ADHD (moderate) and SCZ and ASD (low).
Lee, S.H. et al. Estimation and partitioning of polygenic variation captured by common SNPs for Alzheimer's disease, multiple sclerosis and endometriosis. Hum. Mol. Genet. 22, 832–841 (2013).
Yang, J. et al. Common SNPs explain a large proportion of the heritability for human height. Nat. Genet. 42, 565–569 (2010). This is a landmark paper that described methods for estimation of heritability for quantitative traits based on genetic relationships among individuals inferred from genome-wide data on common SNPs. Applied to height, the classic model trait for human quantitative genetics, the authors demonstrate that a substantial proportion of the heritability can be explained by many common loci throughout the genome.
Lee, S.H., Wray, N.R., Goddard, M.E. & Visscher, P.M. Estimating missing heritability for disease from genome-wide association studies. Am. J. Hum. Genet. 88, 294–305 (2011).
Yang, J. et al. Ubiquitous polygenicity of human complex traits: genome-wide analysis of 49 traits in Koreans. PLoS Genet. 9, e1003355 (2013).
Gratten, J., Visscher, P.M., Mowry, B.J. & Wray, N.R. Interpreting the role of de novo protein-coding mutations in neuropsychiatric disease. Nat. Genet. 45, 234–238 (2013).
Visscher, P.M., Goddard, M.E., Derks, E.M. & Wray, N.R. Evidence-based psychiatric genetics, AKA the false dichotomy between common and rare variant hypotheses. Mol. Psychiatry 17, 474–485 (2012).
Kapur, S., Phillips, A.G. & Insel, T.R. Why has it taken so long for biological psychiatry to develop clinical tests and what to do about it? Mol. Psychiatry 17, 1174–1179 (2012).
Allardyce, J., Suppes, T. & Van Os, J. Dimensions and the psychosis phenotype. Int. J. Methods Psychiatr. Res. 16 Suppl 1, S34–S40 (2007).
Kendler, K.S. & Gardner, C.O. The risk for psychiatric disorders in relatives of schizophrenic and control probands: a comparison of three independent studies. Psychol. Med. 27, 411–419 (1997).
Lee, S.H., Yang, J., Goddard, M.E., Visscher, P.M. & Wray, N.R. Estimation of pleiotropy between complex diseases using single-nucleotide polymorphism–derived genomic relationships and restricted maximum likelihood. Bioinformatics 28, 2540–2542 (2012).
Bromet, E.J. et al. Diagnostic shifts during the decade following first admission for psychosis. Am. J. Psychiatry 168, 1186–1194 (2011).
Laursen, T.M., Agerbo, E. & Pedersen, C.B. Bipolar disorder, schizoaffective disorder, and schizophrenia overlap: a new comorbidity index. J. Clin. Psychiatry 70, 1432–1438 (2009).
Wray, N.R., Lee, S.H. & Kendler, K.S. Impact of diagnostic misclassification on estimation of genetic correlations using genome-wide genotypes. Eur. J. Hum. Genet. 20, 668–674 (2012).
Lynch, M. & Walsh, B. Genetics and Analysis of Quantitative Traits (Sinauer Associates, 1998).
Ji, J. et al. Incidence of cancer in patients with schizophrenia and their first-degree relatives: a population-based study in Sweden. Schizophr. Bull. 39, 527–536 (2013).
Duncan, L.E. & Keller, M.C. A critical review of the first 10 years of candidate gene-by-environment interaction research in psychiatry. Am. J. Psychiatry 168, 1041–1049 (2011).
McGrath, J.J., Mortensen, P.B., Visscher, P.M. & Wray, N.R. Where GWAS and epidemiology meet: opportunities for the simultaneous study of genetic and environmental risk factors in schizophrenia. Schizophr. Bull. 39, 955–959 (2013).
Medland, S.E., Jahanshad, N., Neale, B.M. & Thompson, P.M. Whole-genome analyses of whole-brain data: working within an expanded search space. Nat. Neurosci. 17, 791–800 (2014).
WHO. The Global Burden of Disease: 2004 Update (WHO Press, 2008).
de Candia, T.R. et al. Additive genetic variation in schizophrenia risk is shared by populations of African and European descent. Am. J. Hum. Genet. 93, 463–470 (2013).
Yang, L. et al. Polygenic transmission and complex neuro developmental network for attention deficit hyperactivity disorder: genome-wide association study of both common and rare variants. Am. J. Med. Genet. B. Neuropsychiatr. Genet. 162B, 419–430 (2013).
Sullivan, P.F. The genetics of schizophrenia. PLoS Med. 2, e212 (2005).
Yang, J., Visscher, P.M. & Wray, N.R. Sporadic cases are the norm for complex disease. Eur. J. Hum. Genet. 18, 1039–1043 (2010).
Turner, M.R. et al. Controversies and priorities in amyotrophic lateral sclerosis. Lancet Neurol. 12, 310–322 (2013).
Sebat, J., Levy, D.L. & McCarthy, S.E. Rare structural variants in schizophrenia: one disorder, multiple mutations; one mutation, multiple disorders. Trends Genet. 25, 528–535 (2009).
Acknowledgements
This work was supported by the US National Institutes of Health (GM099568 and GM075091 to P.M.V.), the National Institute of Mental Health (K01MH085812 and R01MH100141 to M.C.K.), the Australian Research Council (FT0991360 to N.R.W.) and the Australian National Health and Medical Research Council (APP1011506, APP1048853 and APP1067795 to P.M.V., APP1011506 and APP1047956 to N.R.W., APP1067795 to J.G.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Locus
-
A locus is a place on a chromosome. A locus may contain one gene, multiple genes or no genes at all.
- Allele
-
An allele is one of a number of alternative forms of a gene or locus. The minor allele is the less frequent allele at a locus and the major allele is the more frequent allele.
- Whole-genome sequencing (WGS)
-
Whole-genome sequencing (WGS) is the sequencing of all of the DNA in an individual's genome.
- Exome
-
The exome is the part of a genome that encodes proteins, approximately 1% of the human genome.
- Common variant
-
Common variant generally refers to an allele that segregates in a population at an allele frequency of at least 5%.
- Rare-variant association study (RVAS)
-
A rare-variant association study (RVAS) is a genome-wide association study to discover rare variants that, as a group, present at different frequencies in affected and unaffected individuals. RVAS can be performed by whole-genome or whole-exome sequencing.
- de novo mutation (DNM)
-
A de novo mutation (DNM) is a mutation that is part of an individual's genome that is not detected in the genome of either parent (although it may have arisen from a mutation in the parental germline). With the exception of de novo mutations in monozygotic twins, or those shared by siblings as a result of germline mosaicism, most new mutations are not shared by relatives and do not contribute to heritability estimates.
- Genome-wide association study (GWAS)
-
A genome-wide association study (GWAS) is an unbiased screen of the genome for genetic variants that present at different frequencies in affected and unaffected individuals, that is, that associate with a phenotype. Although either rare or common variants can now be studied and analyzed for association in a genome-wide way, GWAS has historically referred to a specific, early type of genome-wide study in which a genome-wide set of common polymorphisms (single nucleotide polymorphisms) is analyzed using microarray-based technologies to find disease-associated common alleles.
- Copy number variation (CNV)
-
A copy number variation (CNV) is a type of submicroscopic genetic variation involving the deletion or duplication of a genomic region. Although CNVs can involve genomic segments as small as a kilobase or as large as several megabases, most CNVs detected are relatively large (100 kilobases or larger) because of the resolution of genotyping arrays; in the future, sequencing-based studies may analyze many smaller CNVs.
- Whole-exome sequencing (WES)
-
Whole-exome sequencing (WES) is the targeted enrichment and sequencing of the set of all protein-coding exons and non-coding RNAs in the genome (the exome). WES is performed by selectively capturing the protein-coding part of the genome by hybridization to pre-designed oligonucleotide 'baits'. The captured DNA is then sequenced. Although WES offers a less-complete view of an individual's genome sequence than whole-genome sequencing, WES has been more frequently used because of its substantially lower cost. As the price of sequencing continues to fall, WES may be gradually replaced by whole-genome sequencing.
- Candidate gene
-
A candidate gene is a pre-specified gene of potential interest. Candidate gene studies are often distinguished from unbiased genome-wide studies that analyze variation in all or most genes simultaneously.
- Common variant
-
Common variant generally refers to an allele that segregates in a population at an allele frequency of at least 5%.
- Polygenic
-
Polygenic is a term meaning "many genes". A polygenic phenotype is influenced by more than one gene and can refer to common variants with small effects or rare variants with larger effects.
- Complex disease
-
A complex disease describes a disorder caused by many contributing factors, both genetic and non-genetic, and does not display a simple pattern of inheritance.
- Genotyping arrays
-
Genotyping arrays are a microarray-based technology that allows the inexpensive typing of hundreds of thousands of single-nucleotide variants (single nucleotide polymorphisms) and the ascertainment of simpler, larger forms of copy number variation. Because common variants can be systematically cataloged and assays for them placed on microarray platforms, genotyping arrays are used to perform genome-wide association studies for common variants.
- Odds ratio (OR)
-
An odds ratio (OR) measures the effect size of a genetic variant. An OR of 1 means that the variant has no effect, OR > 1 is risk-conferring and OR < 1 is protective.
- Proband
-
A proband is an individual being studied or reported on. The term is often used to refer to an individual affected with a disease or disorder, as distinct from their unaffected relatives.
- Trio family study
-
A trio family study is an analysis of probands and both of their parents. Sequencing-based trio studies often focus on de novo mutations that are present in the proband's genome, but are not detected in the genomes of his or her parents.
- Case-control study
-
A case-control study is a study design that compares the distribution of a genetic or other variable between individuals affected with a disease (cases) and unaffected individuals (controls).
- Heritability
-
Heritability refers to the proportion of phenotypic variance of a trait, such as disease liability, that can be attributed to genetic factors.
- Mutational target
-
Mutational target refers to the proportion of the genome, or the number of genes, that harbor causal genetic variation for a complex trait or disease.
- Low-frequency variant
-
Low-frequency variant refers to a variant that is not common, but still segregates in a population at an appreciable frequency (for example, 0.1 to 5%) and is observed in the genomes of multiple individuals who are not closely related.
- Single-nucleotide polymorphism (SNP)
-
A single-nucleotide polymorphism (SNP) is a single base-pair position in the genome that varies between members of a species. The terms polymorphism and SNP generally refer to sequence variations that segregate in a population at an allele frequency of at least 1%.
- Mendelian disease
-
A Mendelian disease describes a single gene disorder that is caused by the presence of one (dominant) or two (recessive) alleles.
Rights and permissions
About this article
Cite this article
Gratten, J., Wray, N., Keller, M. et al. Large-scale genomics unveils the genetic architecture of psychiatric disorders. Nat Neurosci 17, 782–790 (2014). https://doi.org/10.1038/nn.3708
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.3708
This article is cited by
-
A robust immune-related gene pairs signature for predicting the overall survival of esophageal cancer
BMC Genomics (2023)
-
The molecular genetic basis of creativity: a mini review and perspectives
Psychological Research (2023)
-
Analysis of the Relationship between Genetic Factors and the Risk of Schizophrenia
Neuroscience and Behavioral Physiology (2023)
-
Polygenic risk scores for schizophrenia and major depression are associated with socio-economic indicators of adversity in two British community samples
Translational Psychiatry (2022)
-
Induced neural progenitor cells and iPS-neurons from major depressive disorder patients show altered bioenergetics and electrophysiological properties
Molecular Psychiatry (2022)