Abstract
In neuroscience, experimental designs in which multiple observations are collected from a single research object (for example, multiple neurons from one animal) are common: 53% of 314 reviewed papers from five renowned journals included this type of data. These so-called 'nested designs' yield data that cannot be considered to be independent, and so violate the independency assumption of conventional statistical methods such as the t test. Ignoring this dependency results in a probability of incorrectly concluding that an effect is statistically significant that is far higher (up to 80%) than the nominal α level (usually set at 5%). We discuss the factors affecting the type I error rate and the statistical power in nested data, methods that accommodate dependency between observations and ways to determine the optimal study design when data are nested. Notably, optimization of experimental designs nearly always concerns collection of more truly independent observations, rather than more observations from one research object.
Similar content being viewed by others
Main
Neuroscience has seen major advances in understanding the nervous system over the past decades. Serious concerns have, however, been raised about an excess of false positive results contaminating the neuroscience literature1,2,3,4. Controlling the false positive rate is critical, since theoretical progress in the neuroscience field relies fundamentally on drawing correct conclusions from experimental research. Reported causes of increased levels of false positives range from inadequate sample size (i.e., underpowered studies), to lack of standardization with respect to research design, applied measures and corrections, exclusion/inclusion criteria, and choice of statistical methods. To improve transparency and reproducibility, Nature journals recently developed a checklist to aid authors to report basic methods information5,6. Among things, authors are asked whether the assumptions of chosen statistical methods are met. Here, we show that one of these assumptions, i.e., the assumption of independent observations, is particularly relevant to neuroscience: neuroscience data often show dependency (that is, nesting; Box 1) and failure to accommodate this is another, as yet neglected, cause of false positive results.
Nested designs are not unique to the neuroscience field, but are also encountered, for instance, in the social sciences (for example, children nested in classes, nested in schools), in behavioral genetics (for example, relatives nested in families) and in the field of medicine (for example, patients nested in doctors, nested in hospitals). In biomedical research, nested data are common in electron microscopy studies, with the n often at a subcellular level. In neuroscience, however, studies on neuron morphology and physiology typically give rise to nested data, as technical advances allow researchers to obtain measurements on every dendrite of a neuron and every spine of each dendrite or to acquire multiple recordings of neuronal activity from the same cell.
The problem of nesting
Nested designs are designs in which multiple observations or measurements are collected in each research object (for example, animal, tissue sample or neuron/cell)7. Consider the following fictive, yet representative, research results. “The channel blocker significantly affected Ca2+ signals (n = 120 regions of interest (ROI) from 10 cells, P < 0.01).” “The number of vesicles docked at the active zone was smaller in presynaptic boutons in mutant neurons than in WT neurons (n = 20 and 25 synapses each from 3 neurons for mutant and WT, P < 0.01).” Both statements concern experimental designs involving nested (or clustered) data. These nested designs are particularly common to neuroscience, as many research questions in neuroscience consider multiple layers of complexity: from protein complexes, synapses and neurons, to neuronal networks, connected systems in the brain and behavior. In such multiple layer–crossing designs, careful consideration of the issues that come with nesting is crucial to avoid incorrect inferences. The generality of nested designs in molecular, cellular and developmental neuroscience is apparent from a literature study that we conducted involving research articles published over the last 18 months in Science, Nature, Cell, Nature Neuroscience and every first issue of the month of Neuron (see below): at least 53% of the 314 publications included nested data.
But why is nesting an issue? Given that observations taken from the same research object (for example, brain, animal, cell) tend to be more similar than observations taken from different objects (for example, due to natural variation between objects, and differences in measurement procedures or conditions), nested designs yield clusters of observations that cannot be considered independent. Nevertheless, conventional statistical methods, such as the t test and ANOVA, are often used to analyze these nested data, even though these methods assume observations to be independent. However, the failure to take the dependency among observations into account forms a threat to the validity of the statistical inference. Depending on the number of observations per research object and the degree of dependence, the probability of incorrectly concluding that an effect is statistically significant (that is, type I error rate) can be far higher than the nominal level expressed by α (usually α = 0.05). To illustrate the effect of nesting on results obtained through conventional tests, we conducted a simulation study (Fig. 1 and Supplementary Simulation). Given a nominal α of 0.05, ignoring nesting can result in an actual type I error rate as high as 0.80. That is, if no experimental effect is present, conventional methods that do not accommodate dependency will yield spurious statistically significant results in 80% of the studies (see Box 2 for a detailed discussion of the results and the theoretical proof).
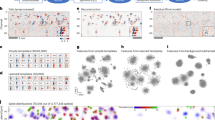
(a) Under two conditions (unexplained ICC = 0.10 or 0.50), nested data were simulated for two experimental groups (for example, knockout versus wild type), with 25 clusters per group. The groups did not differ with respect to their means (that is, no experimental effect). These nested data were analyzed using either a conventional t test or multilevel analysis. When using a t test, the type I error increased steadily as the number of observations per cluster increased. The yellow bars with accompanying right y axis represent the average number of observations per cluster from 314 research articles published in Science, Nature, Cell, Nature Neuroscience and Neuron is shown. The vertical gray line represents the median number of observations per cluster reported in the literature. (b) Under two conditions (unexplained ICC = 0.10 or 0.50), nested data were simulated for two experimental groups with a small, medium or large experimental effect (Cohen's d = 0.20, 0.50 or 0.80, respectively). Compared with multilevel analysis, the loss in power when analyzing summary statistics is larger when the number of clusters is smaller. The vertical gray line represents the median number of clusters observed in the 7% of published papers that reported analyses on cluster-based summary statistics in which multilevel analysis could have been used.
The distinction between the observed and the effective sample size is essential for understanding why clustering affects the type I error rate. The core of this distinction is whether each individual observation contributes unique information. This can be inferred from the degree of relative similarity between observations obtained from the same research object. This similarity is expressed in the intracluster correlation (ICC), which ranges from 0 to 1 (Fig. 2a–c). If clustering is absent (ICC = 0), all observations obtained from a research object are independent, that is, contribute fully unique information. In the extreme case of ICC = 1, all observations obtained from the same research objects are equal and therefore convey the very same information.
(a–c) Graphical representation of three data sets with overall intracluster correlations ICC of 0.00 (a), 0.50 (b), fully explained by experimental condition, and 0.50 (c), partly explained by experimental condition, respectively. The ICC is calculated from the variance between clusters (inferred from the deviations of the cluster means from the grand mean, represented by the horizontal dashed line) and the total variance (that is, the sum of the variance between clusters and the variance within clusters, calculated from the deviations of individual observations from their cluster mean). (d) Effective sample size as function of the ICC under two conditions (number of observations per clusters was 5 or 50, total number of observations was always 500). The higher the unexplained ICC, the larger the difference between the observed sample size (Ntotal = 500) and the effective sample size. The difference between the observed and effective sample size increases faster when the number of observations per cluster is higher.
The experimental variable (for example, genotype) can contribute to the dissimilarity of observations from different objects, and thereby to the relative similarity of observations from the same object. The part of the relative similarity that is attributable to the experimental variable is referred to as the explained ICC, whereas the part of the ICC that is attributable to other, unknown factors is called the unexplained ICC. We use the term ICC to indicate the unexplained ICC, unless stated otherwise. Notably, the unexplained part of the ICC causes inflation of the type I error rate.
In the extreme case of ICC = 1, the observed sample size may be N, but the effective sample size, that is, the number of unique information units, equals the number of research objects (that is, the number of clusters). For example, given five measurements on ten cells, ICC = 0 implies a sample size of 5 × 10 = 50, but as the ICC tends to 1, the effective sample size tends to 10 (Fig. 2d). In terms of variation, correlation between observations from the same research objects (ICC > 0) reduces the variation in the total sample, compared with the variation expected in a random sample (ICC = 0; Fig. 2a). Because conventional statistical analyses are based on the observed rather than the effective sample size, standard errors of parameters are underestimated and test statistics are overestimated. As a result, the associated P values are too low, which results in an excessive type I error rate8.
To correctly handle dependence in nested designs, multilevel models (also known as hierarchical or random effects models) can be used. These models produce correct type I error rates (Fig. 1a). Alternatively, multilevel analysis can be circumvented by conducting conventional analyses on cluster-based summary statistics, for example, by performing a t test on the means or medians calculated in each cluster. Although this strategy is statistically valid, information contributed by the individual observations is lost, and, relative to multilevel analysis, statistical power to detect the experimental effect of interest decreases7,9,10. Conducting t tests on cluster-based means instead of multilevel analysis on all observations results in up to a 40% loss of statistical power, depending on the number of clusters in the study and the ICC (Fig. 1b, and Supplementary Simulation).
The prevalence of nesting in neuroscience studies
To assess the prevalence of nested data and the ensuing problem of inflated type I error rate in neuroscience, we scrutinized all molecular, cellular and developmental neuroscience research articles published in five renowned journals (Science, Nature, Cell, Nature Neuroscience and every month's first issue of Neuron) in 2012 and the first six months of 2013. Unfortunately, precise evaluation of the prevalence of nesting in the literature is hampered by incomplete reporting: not all studies report whether multiple measurements were taken from each research object and, if so, how many. Still, at least 53% of the 314 examined articles clearly concerned nested data, of which 44% specifically reported the number of observations per cluster with a minimum of five observations per cluster (that is, for robust multilevel analysis a minimum of five observations per cluster is required11,12). The median number of observations per cluster, as reported in literature, was 13 (Fig. 1a), yet conventional analysis methods were used in all of these reports.
The studies reporting nested data typically do not provide information on the ICC, which is required to evaluate the extent to which clustering affected the type I error rates of these studies. To assess the range of ICCs that can be expected, we analyzed 36 research questions in 18 neuroscience data sets from varying disciplines. In these data, unexplained ICCs ranged between 0.00 and 0.74, with a mean of 0.19 (Supplementary Table 1). As even a low degree of dependency (for example, ICC = 0.10) increases the type I error rate from 5% to nearly 20% when the number of observations per cluster is 13 (the median number of observations per cluster observed in our literature study; Fig. 1a), an excess number of false positive results is to be expected. It should be noted that differences in the statistical significance of the findings between multilevel analysis and conventional testing are a result of the unexplained ICC only, not the total ICC (see, for example, results of analysis 7, where all of the ICC is explained). Further inspection of the research articles that reported nested data with a minimum of five observations per cluster showed that 25% of the P values were between 0.01 and 0.001, and 31% between 0.05 and 0.01. False positive effects are to be expected in at least some of these articles. Moreover, another 7% of the examined papers used cluster-based summary statistics in which multilevel analysis could have been applied, resulting in a loss of power to detect experimental effects (see Fig. 1b and Supplementary Table 1 for examples of non-significant results obtained with pooled t tests, which actually prove significant when multilevel analysis is used).
Multilevel analysis
Multilevel models can be used to statistically accommodate dependence between observations in nested designs. The basics of multilevel analysis are readily explained with reference to the conventional two-group t test. Suppose we studied whether characteristic X of the cell is affected by a specific gene mutation. In 15 mice carrying the mutation, we collect ten cells (15 × 10 observations) and we do the same in 15 mice that do not carry the mutation, resulting in 300 observations in total. A standard t test on these data can be carried out by regressing X on the dummy coded (0/1) experimental variable. Significance of the slope parameter, representing the differences in means, can be tested using a t test (Fig. 3a). In this conventional analysis, cluster information is discarded: all 15 × 10 observations in each group are simply pooled. In contrast, in multilevel analysis, the individual observations (the cells) are regarded as level 1 units, which are nested in the level 2 units: the clusters (the mice). Multilevel analysis retains cluster-membership information by conducting the t test on the cluster-level (mouse) means while retaining the distinction between the variance within clusters (differences between cells within a mouse) and variance between clusters (differences between the mice in cluster-level means; Fig. 3b). Multilevel analysis therefore effectively accommodates the possibly increased similarity of observations taken from the same research object by retaining cluster-membership information of each individual observation when evaluating parameters such as group differences.
(a) Graphical representation of the conventional t test in regression terms: the individual observations yi are a function of the mean of the control group (that is, the intercept β0) and, when applicable, the estimated deviation from this mean for observations from the experimental group (that is, the slope β1), and an individual error term ei. X is essentially a weight variable that takes on values 0 and 1 for observations from the control and experimental group, respectively. (b) Graphical representation of multilevel analysis. The individual outcomes of observation i from cluster j, yij, are a function of the cluster-specific intercept β0j plus, when applicable, the estimated deviation from this intercept for clusters belonging to the experimental group, β1, and an individual-specific error term eij. The higher the unexplained ICC, the more variation there is in the cluster specific intercepts β0j.
Various standardized effect size measures have been suggested in the context of multilevel analysis13,14. When comparing only two experimental conditions, Cohen's d is a generally accepted index. When comparing more than two experimental conditions, the overall effect size can be represented by the explained variance R2, which equals the explained ICC when the experimental condition only varies over clusters and not within clusters. Cohen15 defined a Cohen's d of 0.20, 0.50 and 0.80, and an explained variance R2 of 0.01, 0.09 and 0.25 as small, medium and large effects, respectively. Note that these two effect sizes are not on the same scale and can therefore not be compared directly. However, d and R2 can be converted into each other16


Note that in equation (1), the sum of the Cohen's d values obtained in pairwise comparisons is used when multiple pair-wise comparisons are combined into one omnibus test. Equations (1) and (2) assume experimental groups with equal sample sizes (see ref. 16 for formulas for unbalanced designs). A worked example of the analysis of nested data, including effect size calculation, is provided in the Supplementary Analysis and Supplementary Tables 2–6.
Power up: determining the optimal study design
Generally, power is increased by increasing the number of observations in a study. In conventional analysis, this is straightforward, but, in multilevel analysis, the relation between sample size and power is more complicated as the total number of observations is distributed over the research objects (clusters). In the allocation of research resources (for example, money and time), a trade-off must be considered between the number of clusters and the number of observations per cluster. In practice, collecting many measures in a few clusters may be easier, faster and cheaper than collecting a few measures in many clusters. But which option confers the greatest power?
In multilevel analysis, power depends essentially on the number of clusters: power steadily increases to 100% as the number of clusters increases (Fig. 4a). In contrast, when increasing the number of observations per cluster, the power curve often approaches an asymptote below 100%, with the maximum level depending on the ICC (Fig. 4b). In general, high ICCs result in lower power and, unless the ICC is low, adding extra observations per cluster does little to increase power (Fig. 4a,b).
Power is depicted under six conditions (Cohen's d of 0.20, 0.50 or 0.80, and unexplained intracluster correlation ICC of 0.10 or 0.50) and as function of the number of clusters (a) or the number of observations per cluster (b). In a, the number of observations per cluster is held constant at 5; in b, the number of clusters is held constant at 10. Evidently, the number of clusters, and not the number of observations per cluster, is essential to increase the statistical power to detect the experimental effect.
Given available resources (for example, money or time), the optimal balance between the number of clusters (N) and the number of observations per cluster (n) can be determined, given a specified level of dependency (ICC). In theory, optimal N and n are dictated by the desired level of statistical power. In practice, however, available resources have a bearing on the attainable values of N and n. Given that including additional observations within a cluster (C1) is usually less costly than including an additional cluster (C2), these two costs are defined distinctly. The total costs of a study are calculated as (3)

while the optimal number of observations per cluster can be obtained by solving9,13 (4)

where  is the residual variance, which equals 1 − overall ICC (note that we make use of the standardized model, that is, the observations are standardized such that they have a mean of 0 and an s.d. of 1). Given the total available resources T and the optimal number of observations per cluster noptimal, the optimal number of clusters N can be obtained by (5)
is the residual variance, which equals 1 − overall ICC (note that we make use of the standardized model, that is, the observations are standardized such that they have a mean of 0 and an s.d. of 1). Given the total available resources T and the optimal number of observations per cluster noptimal, the optimal number of clusters N can be obtained by (5)

The optimal balance between the number of clusters and number of observations per cluster does not guarantee that the subsequent study will have sufficient power to detect the experimental effect of interest. The actual power of the experiment also depends on the chosen α level and on the expected effect size (for example, the magnitude of the difference between the control and experimental group). However, the calculated optimal N and n can be used to estimate the expected power given specific values of the effect size and the ICC (Box 3).
Discussion
Multilevel modeling is relevant to neuroscientific data collected using traditional techniques, such as the analysis of immunofluorescence signal intensity in slices (where the use of cluster-based summary statistics causes a loss of power), and the analysis of electrophysiological parameters, such as excitatory postsynaptic potentials (where the use of conventional statistical models inflates type I error rates). Recent advances in the field of neuroscience, such as optogenetics, super-resolution microscopy, immunogold cytochemistry and optopharmacology, will, if anything, increase the relevance of multilevel modeling17. A common feature of all these techniques is that they shift the n from the animal or tissue level to the cellular or even subcellular level, and invariably yield data with a nested structure. For instance, super-resolution light microscopy allows imaging and advanced understanding of neuronal compartments18, immunogold cytochemistry allows determination of subcellular localization of proteins19, and recent advances in optogenetics and optopharmacology facilitate selective control of electrical and protein activity, respectively, in circuits, individual cells or subcellular compartments20,21. All these techniques concern the collection of multiple observations from one cell, thereby yielding nested data.
To fully exploit the advantages that these techniques offer, neuroscientists should adopt multilevel modeling to avoid the limitations of conventional analyses in this context. In addition, nested data come with specific design issues that are relevant to the statistical power to resolve the effects of interest. Optimization of design in terms of allocation of resources does not guarantee sufficiently powered studies. In terms of power, the ratio of number of research objects (for example, mice) to the number of measurements per object (for example, cells per mouse) is important. We showed that the power increase achievable by increasing the latter is limited (Fig. 4). In addition, to obtain robust and unbiased estimates of variance components in multilevel analysis, sufficient observations on both levels are required. As a rule of thumb, afforded by simulation studies11,12, a minimum of five observations per cluster and ten clusters per experimental group are recommended to obtain a robust and unbiased estimate of the standard error for the experimental effect. To also obtain a robust and unbiased estimate of the intracluster correlation, the number of clusters needs to be increased to 30.
Here we focused on the most common design, that is, data that span two levels (for example, cells in mice) and an experimental variable that does not vary within clusters (for example, in comparing cell characteristic X between mutants and wild types, all cells from one mouse have the same genotype). Other nested designs—featuring three or more levels of nesting, experimental variables that do vary within levels (for example, when investigating whether the number of docked vesicles differs between observations from a dendrite or an axon), nested longitudinal data (data collected on multiple time points describing dynamical processes22,23) or nested non-normally distributed data (for example, binary or Poisson distributed data)—are, however, possible and can be analyzed using multilevel analysis. We refer to previous publications12,13,24 for comprehensive introductions to multilevel modeling and to the Centre for Multilevel Modeling website (http://www.bristol.ac.uk/cmm/learning/mmsoftware/) for a recent overview of existing multilevel software.
Various recent publications force neuroscientists to acknowledge the possibility that the harvest of their hard labor is contaminated by an abundance of false positive effects1,2,3,4. Nested designs are ubiquitous in neuroscience, and an increased awareness of the problem of nesting in both researchers and reviewers will prevent costly and time-consuming quixotic pursuits of spurious effects, thereby assisting progress in the understanding of the nervous system.
References
Button, K.S. et al. Power failure: why small sample size undermines the reliability of neuroscience. Nat. Rev. Neurosci. 14, 365–376 (2013).
Ioannidis, J.P. Why most published research findings are false. PLoS Med. 2, e124 (2005).
Nieuwenhuis, S., Forstmann, B.U. & Wagenmakers, E.J. Erroneous analyses of interactions in neuroscience: a problem of significance. Nat. Neurosci. 14, 1105–1107 (2011).
Tsilidis, K.K. et al. Evaluation of excess significance bias in animal studies of neurological diseases. PLoS Biol. 11, e1001609 (2013).
Raising standards. Nat. Neurosci. 16, 517 (2013).
Making methods clearer. Nat. Neurosci. 16, 1 (2013).
Galbraith, S., Daniel, J.A. & Vissel, B. A study of clustered data and approaches to its analysis. J. Neurosci. 30, 10601–10608 (2010).
Walsh, J.E. Concerning the effects of the intra-class correlation on certain significance tests. Ann. Math. Stat. 18, 88–96 (1947).
Raudenbush, S.W. Statistical analysis and optimal design for cluster randomized trials. Psychol. Methods 2, 173–185 (1997).
Moerbeek, M., van Breukelen, G.J.P. & Berger, M.P. A comparison between traditional methods and multilevel regression for the analysis of multicenter intervention studies. J. Clin. Epidemiol. 56, 341–350 (2003).
Maas, C.J.M. & Hox, J.J. Robustness issues in multilevel regression analysis. Stat. Neerl. 58, 127–137 (2004).
Snijders, T.A.B. & Bosker, R.J. Multilevel Analysis: an Introduction to Basic and Advanced Multilevel Modeling (Sage Publications, London, 2011).
Snijders, T.A.B. & Bosker, R.J. Standard errors and sample sizes for 2-level research. J. Educ. Stat. 18, 237–259 (1993).
Hox, J. Multilevel Analysis: Techniques and Applications (Erlbaum, New Jersey, 2010).
Cohen, J. Statistical Power Analysis for the Behavioral Sciences (Erlbaum, Hillsdale, JN, 1988).
Lipsey, M.W. & Wilson, D.B. Practical Meta-analysis (Sage, Thousand Oaks, CA, 2001).
Focus on neurotechniques. Nat. Neurosci. 16, 771 (2013).
Maglione, M. & Sigrist, S.J. Seeing the forest tree by tree: super-resolution light microscopy meets the neurosciences. Nat. Neurosci. 16, 790–797 (2013).
Amiry-Moghaddam, M. & Ottersen, O.P. Immunogold cytochemistry in neuroscience. Nat. Neurosci. 16, 798–804 (2013).
Packer, A.M., Roska, B. & Hausser, M. Targeting neurons and photons for optogenetics. Nat. Neurosci. 16, 805–815 (2013).
Kramer, R.H., Mourot, A. & Adesnik, H. Optogenetic pharmacology for control of native neuronal signaling proteins. Nat. Neurosci. 16, 816–823 (2013).
Smith, A.C., Stefani, M.R., Moghaddam, B. & Brown, E.N. Analysis and design of behavioral experiments to characterize population learning. J. Neurophysiol. 93, 1776–1792 (2005).
Czanner, G. et al. Analysis of between-trial and within-trial neural spiking dynamics. J. Neurophysiol. 99, 2672–2693 (2008).
Goldstein, H. Multilevel Statistical Models (Edward Arnold, London, 2010).
Acknowledgements
We are very grateful to our colleagues from the VU University/VU Medical Center Functional Genomics department for sharing their data. M.V. is supported by the European Union (ERC Advanced grant 322966; HEALTH–F2–2009–241498 EUROSPIN, and HEALTH–F2– 2009–242167 SynSys) and the Netherlands Organization for Scientific Research (TOP 903–42–095). C.V.D. is supported by the European Research Council (Genetics of Mental Illness, grant number: ERC–230374). S.v.d.S. is supported by the Netherlands Scientific Organization (Nederlandse Organisatie voor Wetenschappelijk Onderzoek, gebied Maatschappij- en Gedragswetenschappen: NWO/MaGW: VIDI–452–12–014).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Simulation, Supplementary Analysis, and Supplementary Tables 2–6 (PDF 510 kb)
Supplementary Table 1
Conventional and multilevel analysis of various neuroscience datasets (XLS 53 kb)
Rights and permissions
About this article
Cite this article
Aarts, E., Verhage, M., Veenvliet, J. et al. A solution to dependency: using multilevel analysis to accommodate nested data. Nat Neurosci 17, 491–496 (2014). https://doi.org/10.1038/nn.3648
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.3648
This article is cited by
-
Triple dissociation of visual, auditory and motor processing in mouse primary visual cortex
Nature Neuroscience (2024)
-
Distinct forms of structural plasticity of adult-born interneuron spines in the mouse olfactory bulb induced by different odor learning paradigms
Communications Biology (2024)
-
Cellular mechanisms underlying carry-over effects after magnetic stimulation
Scientific Reports (2024)
-
Similar Gap-Overlap Profiles in Children with Fragile X Syndrome and IQ-Matched Autism
Journal of Autism and Developmental Disorders (2024)
-
Complete sources of cluster variation on the risk of under-five malaria in Uganda: a multilevel-weighted mixed effects logistic regression model approach
Malaria Journal (2023)





 represents the residual error variance (see
represents the residual error variance (see 
 is actually a composite of the residual error variance and variance resulting from clustering. Also, note that equations (6) and (7) assume a standardized model (all variables have a mean of 0 and s.d. of 1), a balanced design (the number of observations per cluster are equal and the number of clusters are evenly divided over the experimental groups) and absence of covariates.
is actually a composite of the residual error variance and variance resulting from clustering. Also, note that equations (6) and (7) assume a standardized model (all variables have a mean of 0 and s.d. of 1), a balanced design (the number of observations per cluster are equal and the number of clusters are evenly divided over the experimental groups) and absence of covariates.

 to the standard normal distribution. In this Z test, the Z statistic reflects the number of s.d. that β1 deviates from the expected value under the null hypothesis (0), from which a P value for β1 can be calculated. Power can be calculated by obtaining the estimated error variance (SE2) of β1, using the estimated error variance to convert β1 to a Z statistic, and subsequently obtaining the probability that the Z score for β1 exceeds the critical value Z1−α for the noncentral Z distribution given α. Below, we discuss the power calculation stepwise.
to the standard normal distribution. In this Z test, the Z statistic reflects the number of s.d. that β1 deviates from the expected value under the null hypothesis (0), from which a P value for β1 can be calculated. Power can be calculated by obtaining the estimated error variance (SE2) of β1, using the estimated error variance to convert β1 to a Z statistic, and subsequently obtaining the probability that the Z score for β1 exceeds the critical value Z1−α for the noncentral Z distribution given α. Below, we discuss the power calculation stepwise. equals 1− overall ICC. The equation to obtain the estimated error variance SE2 of the experimental effect β1 is equation (6). Next, as
equals 1− overall ICC. The equation to obtain the estimated error variance SE2 of the experimental effect β1 is equation (6). Next, as

 is set to 1− 0.25 − 0.01 = 0.74 and 1− 0.25 − 0.06 = 0.69, respectively. Given that β1 is calculated as d × 0.5, the β1 for the small and medium effects correspond to β1 = 0.2 *× 0.5 = 0.1 and β1 = 0.5 ×* 0.5 = 0.25, respectively.
is set to 1− 0.25 − 0.01 = 0.74 and 1− 0.25 − 0.06 = 0.69, respectively. Given that β1 is calculated as d × 0.5, the β1 for the small and medium effects correspond to β1 = 0.2 *× 0.5 = 0.1 and β1 = 0.5 ×* 0.5 = 0.25, respectively.