Abstract
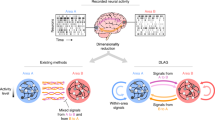
Over the last five decades, progress in neural recording techniques has allowed the number of simultaneously recorded neurons to double approximately every 7 years, mimicking Moore's law. Such exponential growth motivates us to ask how data analysis techniques are affected by progressively larger numbers of recorded neurons. Traditionally, neurons are analyzed independently on the basis of their tuning to stimuli or movement. Although tuning curve approaches are unaffected by growing numbers of simultaneously recorded neurons, newly developed techniques that analyze interactions between neurons become more accurate and more complex as the number of recorded neurons increases. Emerging data analysis techniques should consider both the computational costs and the potential for more accurate models associated with this exponential growth of the number of recorded neurons.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Moore, G.E. Cramming more components onto integrated circuits. Electronics 38 (1965).
Papadimitriou, C.H. Computational Complexity (John Wiley and Sons, 2003).
Nicolelis, M. Methods for Neural Ensemble Recordings 2nd edn (CRC Press, 2007).
Nicolelis, M. et al. Chronic, multisite, multielectrode recordings in macaque monkeys. Proc. Natl. Acad. Sci. USA 100, 11041–11046 (2003).
Kelly, R. et al. Comparison of recordings from microelectrode arrays and single electrodes in the visual cortex. J. Neurosci. 27, 261–264 (2007).
Moore, G. in Understanding Moore's Law: Four Decades of Innovation (ed. Brock, D.C.) Ch. 7 (Chemical Heritage Foundation, 2006).
Harris, K., Henze, D., Csicsvari, J., Hirase, H. & Buzsaki, G. Accuracy of tetrode spike separation as determined by simultaneous intracellular and extracellular measurements. J. Neurophysiol. 84, 401–414 (2000).
Lewicki, M. A review of methods for spike sorting: the detection and classification of neural action potentials. Network 9, R53–R78 (1998).
Brown, E.N., Kass, R.E. & Mitra, P.P. Multiple neural spike train data analysis: state-of-the-art and future challenges. Nat. Neurosci. 7, 456–461 (2004).
Kass, R., Ventura, V. & Brown, E. Statistical issues in the analysis of neuronal data. J. Neurophysiol. 94, 8–25 (2005).
Paninski, L. et al. A new look at state-space models for neural data. J. Comput. Neurosci. 29, 1–20 (2009).
Paninski, L., Pillow, J. & Lewi, J. Statistical models for neural encoding, decoding, and optimal stimulus design. Prog. Brain Res. 165, 493–507 (2007).
Brockwell, A., Rojas, A. & Kass, R. Recursive Bayesian decoding of motor cortical signals by particle filtering. J. Neurophysiol. 91, 1899–1907 (2004).
Okatan, M., Wilson, M.A. & Brown, E.N. Analyzing functional connectivity using a network likelihood model of ensemble neural spiking activity. Neural Comput. 17, 1927–1961 (2005).
Pillow, J.W. et al. Spatio-temporal correlations and visual signaling in a complete neuronal population. Nature 454, 995–999 (2008).
Stevenson, I.H., Rebesco, J.M., Miller, L.E. & Körding, K.P. Inferring functional connections between neurons. Curr. Opin. Neurobiol. 18, 582–588 (2008).
Truccolo, W., Eden, U.T., Fellows, M.R., Donoghue, J.P. & Brown, E.N. A point process framework for relating neural spiking activity to spiking history, neural ensemble and extrinsic covariate effects. J. Neurophysiol. 93, 1074–1089 (2005).
Schneidman, E., Berry, M.J. II, Segev, R. & Bialek, W. Weak pairwise correlations imply strongly correlated network states in a neural population. Nature 440, 1007–1012 (2006).
Maynard, E. et al. Neuronal interactions improve cortical population coding of movement direction. J. Neurosci. 19, 8083–8093 (1999).
Harris, K., Csicsvari, J., Hirase, H., Dragoi, G. & Buzsáki, G. Organization of cell assemblies in the hippocampus. Nature 424, 552–556 (2003).
Paninski, L. Maximum likelihood estimation of cascade point-process neural encoding models. Network 15, 243–262 (2004).
Hatsopoulos, N., Joshi, J. & O'Leary, J.G. Decoding continuous and discrete motor behaviors using motor and premotor cortical ensembles. J. Neurophysiol. 92, 1165–1174 (2004).
Smith, M. & Kohn, A. Spatial and temporal scales of neuronal correlation in primary visual cortex. J. Neurosci. 28, 12591–12603 (2008).
Stevenson, I.H. et al. Bayesian inference of functional connectivity and network structure from spikes. IEEE Trans. Neural Syst. Rehabil. Eng. 17, 203–213 (2009).
Truccolo, W., Hochberg, L. & Donoghue, J. Collective dynamics in human and monkey sensorimotor cortex: predicting single neuron spikes. Nat. Neurosci. 13, 105–111 (2009).
Babadi, B., Casti, A., Xiao, Y., Kaplan, E. & Paninski, L. A generalized linear model of the impact of direct and indirect inputs to the lateral geniculate nucleus. J. Vis. 10, 22 (2010).
Kelly, R., Smith, M., Kass, R. & Lee, T. Local field potentials indicate network state and account for neuronal response variability. J. Comput. Neurosci. 29, 567–579 (2010).
Rebesco, J.M., Stevenson, I.H., Koerding, K., Solla, S.A. & Miller, L.E. Rewiring neural interactions by micro-stimulation. Front. Syst. Neurosci. 4, 39 (2010).
Yu, B. et al. Gaussian-process factor analysis for low-dimensional single-trial analysis of neural population activity. J. Neurophysiol. 102, 614–635 (2009).
Churchland, M., Yu, B., Sahani, M. & Shenoy, K. Techniques for extracting single-trial activity patterns from large-scale neural recordings. Curr. Opin. Neurobiol. 17, 609–618 (2007).
Churchland, M. et al. Stimulus onset quenches neural variability: a widespread cortical phenomenon. Nat. Neurosci. 13, 369–378 (2010).
Vogelstein, J. et al. Spike inference from calcium imaging using sequential monte carlo methods. Biophys. J. 97, 636–655 (2009).
Stosiek, C., Garaschuk, O., Holthoff, K. & Konnerth, A. In vivo two-photon calcium imaging of neuronal networks. Proc. Natl. Acad. Sci. USA 100, 7319–7324 (2003).
Shlens, J. et al. The structure of multi-neuron firing patterns in primate retina. J. Neurosci. 26, 8254–8266 (2006).
Ecker, A. et al. Decorrelated neuronal firing in cortical microcircuits. Science 327, 584–587 (2010).
Vogels, T., Rajan, K. & Abbott, L. Neural network dynamics. Annu. Rev. Neurosci. 28, 357–376 (2005).
Brette, R. et al. Simulation of networks of spiking neurons: a review of tools and strategies. J. Comput. Neurosci. 23, 349–398 (2007).
Averbeck, B., Latham, P. & Pouget, A. Neural correlations, population coding and computation. Nat. Rev. Neurosci. 7, 358–366 (2006).
Pouget, A., Dayan, P. & Zemel, R. Information processing with population codes. Nat. Rev. Neurosci. 1, 125–132 (2000).
Barna, J., Arezzo, J. & Vaughan, H. Jr. A new multielectrode array for the simultaneous recording of field potentials and unit activity. Electroencephalogr. Clin. Neurophysiol. 52, 494–496 (1981).
Krüger, J. & Bach, M. Simultaneous recording with 30 microelectrodes in monkey visual cortex. Exp. Brain Res. 41, 191–194 (1981).
Rousche, P. & Normann, R. Chronic intracortical microstimulation (ICMS) of cat sensory cortex using the Utah Intracortical Electrode Array. IEEE Trans. Rehabil. Eng. 7, 56–68 (2002).
Blanche, T., Spacek, M., Hetke, J. & Swindale, N. Polytrodes: high-density silicon electrode arrays for large-scale multiunit recording. J. Neurophysiol. 93, 2987–3000 (2005).
Acknowledgements
Thanks to A. Kohn and members of the Kohn laboratory for providing data from visual cortex (US National Institutes of Health EY016774) and N. Hatsopoulos and J. Reimer for providing data from motor cortex. All animal use procedures were approved by the institutional animal care and use committees at Albert Einstein College of Medicine and the University of Chicago, respectively. Thanks to B. Yu and J. Cunningham for providing the GPFA code and B. Yu for insightful discussions. This work was supported by the Chicago Community Trust and US National Institutes of Health grants 1R01NS063399 and 2P01NS044393.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Table 1 and Supplementary Methods (PDF 279 kb)
Rights and permissions
About this article
Cite this article
Stevenson, I., Kording, K. How advances in neural recording affect data analysis. Nat Neurosci 14, 139–142 (2011). https://doi.org/10.1038/nn.2731
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.2731
This article is cited by
-
Population coding of strategic variables during foraging in freely moving macaques
Nature Neuroscience (2024)
-
Implantable intracortical microelectrodes: reviewing the present with a focus on the future
Microsystems & Nanoengineering (2023)
-
Large-scale neural recordings call for new insights to link brain and behavior
Nature Neuroscience (2022)
-
A large-scale neural network training framework for generalized estimation of single-trial population dynamics
Nature Methods (2022)
-
Neural tuning and representational geometry
Nature Reviews Neuroscience (2021)