Abstract
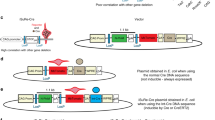
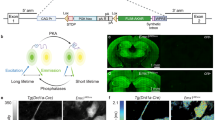
The Cre/lox system is widely used in mice to achieve cell-type-specific gene expression. However, a strong and universally responding system to express genes under Cre control is still lacking. We have generated a set of Cre reporter mice with strong, ubiquitous expression of fluorescent proteins of different spectra. The robust native fluorescence of these reporters enables direct visualization of fine dendritic structures and axonal projections of the labeled neurons, which is useful in mapping neuronal circuitry, imaging and tracking specific cell populations in vivo. Using these reporters and a high-throughput in situ hybridization platform, we are systematically profiling Cre-directed gene expression throughout the mouse brain in several Cre-driver lines, including new Cre lines targeting different cell types in the cortex. Our expression data are displayed in a public online database to help researchers assess the utility of various Cre-driver lines for cell-type-specific genetic manipulation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Feng, G. et al. Imaging neuronal subsets in transgenic mice expressing multiple spectral variants of GFP. Neuron 28, 41–51 (2000).
Sugino, K. et al. Molecular taxonomy of major neuronal classes in the adult mouse forebrain. Nat. Neurosci. 9, 99–107 (2006).
Zong, H., Espinosa, J.S., Su, H.H., Muzumdar, M.D. & Luo, L. Mosaic analysis with double markers in mice. Cell 121, 479–492 (2005).
Balthasar, N. et al. Leptin receptor signaling in POMC neurons is required for normal body weight homeostasis. Neuron 42, 983–991 (2004).
Kellendonk, C. et al. Transient and selective overexpression of dopamine D2 receptors in the striatum causes persistent abnormalities in prefrontal cortex functioning. Neuron 49, 603–615 (2006).
McHugh, T.J. et al. Dentate gyrus NMDA receptors mediate rapid pattern separation in the hippocampal network. Science 317, 94–99 (2007).
Karpova, A.Y., Tervo, D.G., Gray, N.W. & Svoboda, K. Rapid and reversible chemical inactivation of synaptic transmission in genetically targeted neurons. Neuron 48, 727–735 (2005).
Luquet, S., Perez, F.A., Hnasko, T.S. & Palmiter, R.D. NPY/AgRP neurons are essential for feeding in adult mice but can be ablated in neonates. Science 310, 683–685 (2005).
Nakashiba, T., Young, J.Z., McHugh, T.J., Buhl, D.L. & Tonegawa, S. Transgenic inhibition of synaptic transmission reveals role of CA3 output in hippocampal learning. Science 319, 1260–1264 (2008).
Barth, A.L. Visualizing circuits and systems using transgenic reporters of neural activity. Curr. Opin. Neurobiol. 17, 567–571 (2007).
Luo, L., Callaway, E.M. & Svoboda, K. Genetic dissection of neural circuits. Neuron 57, 634–660 (2008).
Xu, X., Roby, K.D. & Callaway, E.M. Mouse cortical inhibitory neuron type that coexpresses somatostatin and calretinin. J. Comp. Neurol. 499, 144–160 (2006).
Holtmaat, A., Wilbrecht, L., Knott, G.W., Welker, E. & Svoboda, K. Experience-dependent and cell-type-specific spine growth in the neocortex. Nature 441, 979–983 (2006).
Livet, J. et al. Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature 450, 56–62 (2007).
Boyden, E.S., Zhang, F., Bamberg, E., Nagel, G. & Deisseroth, K. Millisecond-timescale, genetically targeted optical control of neural activity. Nat. Neurosci. 8, 1263–1268 (2005).
Arenkiel, B.R. et al. In vivo light-induced activation of neural circuitry in transgenic mice expressing channelrhodopsin-2. Neuron 54, 205–218 (2007).
Soriano, P. Generalized lacZ expression with the ROSA26 Cre reporter strain. Nat. Genet. 21, 70–71 (1999).
Srinivas, S. et al. Cre reporter strains produced by targeted insertion of EYFP and ECFP into the ROSA26 locus. BMC Dev. Biol. 1, 4 (2001).
Novak, A., Guo, C., Yang, W., Nagy, A. & Lobe, C.G. Z/EG, a double reporter mouse line that expresses enhanced green fluorescent protein upon Cre-mediated excision. Genesis 28, 147–155 (2000).
Muzumdar, M.D., Tasic, B., Miyamichi, K., Li, L. & Luo, L. A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007).
Shaner, N.C., Patterson, G.H. & Davidson, M.W. Advances in fluorescent protein technology. J. Cell Sci. 120, 4247–4260 (2007).
Zufferey, R., Donello, J.E., Trono, D. & Hope, T.J. Woodchuck hepatitis virus posttranscriptional regulatory element enhances expression of transgenes delivered by retroviral vectors. J. Virol. 73, 2886–2892 (1999).
Raymond, C.S. & Soriano, P. High-efficiency FLP and PhiC31 site-specific recombination in mammalian cells. PLoS ONE 2, e162 (2007).
Lein, E.S. et al. Genome-wide atlas of gene expression in the adult mouse brain. Nature 445, 168–176 (2007).
Zhuang, X., Masson, J., Gingrich, J.A., Rayport, S. & Hen, R. Targeted gene expression in dopamine and serotonin neurons of the mouse brain. J. Neurosci. Methods 143, 27–32 (2005).
Lewis, P.M., Gritli-Linde, A., Smeyne, R., Kottmann, A. & McMahon, A.P. Sonic hedgehog signaling is required for expansion of granule neuron precursors and patterning of the mouse cerebellum. Dev. Biol. 270, 393–410 (2004).
Gong, S. et al. Targeting Cre recombinase to specific neuron populations with bacterial artificial chromosome constructs. J. Neurosci. 27, 9817–9823 (2007).
Dong, H.W. The Allen Reference Atlas: A Digital Color Brain Atlas of the C57BL/6J Male Mouse (Wiley, Hoboken, New Jersey, USA, 2008).
Ng, L. et al. Neuroinformatics for genome-wide 3D gene expression mapping in the mouse brain. IEEE/ACM Trans. Comput. Biol. Bioinform. 4, 382–393 (2007).
Lau, C. et al. Exploration and visualization of gene expression with neuroanatomy in the adult mouse brain. BMC Bioinformatics 9, 153 (2008).
Coulter, C.L., Happe, H.K. & Murrin, L.C. Postnatal development of the dopamine transporter: a quantitative autoradiographic study. Brain Res. Dev. Brain Res. 92, 172–181 (1996).
Furuta, Y., Lagutin, O., Hogan, B.L. & Oliver, G.C. Retina- and ventral forebrain-specific Cre recombinase activity in transgenic mice. Genesis 26, 130–132 (2000).
Szymczak, A.L. et al. Correction of multi-gene deficiency in vivo using a single 'self-cleaving' 2A peptide-based retroviral vector. Nat. Biotechnol. 22, 589–594 (2004).
Hippenmeyer, S. et al. A developmental switch in the response of DRG neurons to ETS transcription factor signaling. PLoS Biol. 3, e159 (2005).
Atasoy, D., Aponte, Y., Su, H.H. & Sternson, S.M.A. FLEX switch targets Channelrhodopsin-2 to multiple cell types for imaging and long-range circuit mapping. J. Neurosci. 28, 7025–7030 (2008).
Kuhlman, S.J. & Huang, Z.J. High-resolution labeling and functional manipulation of specific neuron types in mouse brain by Cre-activated viral gene expression. PLoS ONE 3, e2005 (2008).
Rotolo, T., Smallwood, P.M., Williams, J. & Nathans, J. Genetically-directed, cell type-specific sparse labeling for the analysis of neuronal morphology. PLoS ONE 3, e4099 (2008).
Tsien, J.Z. et al. Subregion- and cell type-restricted gene knockout in mouse brain. Cell 87, 1317–1326 (1996).
Heusner, C.L., Beutler, L.R., Houser, C.R. & Palmiter, R.D. Deletion of GAD67 in dopamine receptor-1 expressing cells causes specific motor deficits. Genesis 46, 357–367 (2008).
Gorski, J.A. et al. Cortical excitatory neurons and glia, but not GABAergic neurons, are produced in the Emx1-expressing lineage. J. Neurosci. 22, 6309–6314 (2002).
Zhuo, L. et al. hGFAP-cre transgenic mice for manipulation of glial and neuronal function in vivo. Genesis 31, 85–94 (2001).
George, S.H. et al. Developmental and adult phenotyping directly from mutant embryonic stem cells. Proc. Natl. Acad. Sci. USA 104, 4455–4460 (2007).
Zhang, Y., Buchholz, F., Muyrers, J.P. & Stewart, A.F. A new logic for DNA engineering using recombination in Escherichia coli. Nat. Genet. 20, 123–128 (1998).
Thompson, C.L. et al. Genomic anatomy of the hippocampus. Neuron 60, 1010–1021 (2008).
Tsai, P.S. et al. Principle, design and construction of a two photon laser-scanning microscope for in vitro and in vivo brain imaging. in In Vivo Optical Imaging of Brain Function (ed. Frostig, R.D.) 113–172 (CRC, Boca Raton, Florida, USA, 2002).
Garaschuk, O., Milos, R.I. & Konnerth, A. Targeted bulk-loading of fluorescent indicators for two-photon brain imaging in vivo. Nat. Protoc. 1, 380–386 (2006).
Acknowledgements
We are grateful for the professional support of the entire Atlas production team, led by P. Wohnoutka, and the Information Technology team, led by C. Dang, at the Allen Institute, without which the work would have not been possible to accomplish. We are thankful to A. Bernard for her contribution in establishing the DFISH process, L. Kuan for ISH data quantification and R. Hunter for coordinating transgenic mice production. We also gratefully acknowledge the following researchers for providing various research materials: R. Tsien (University of California at San Diego) for the tdTomato DNA construct, L. Luo (Stanford University) for the Rosa-CAG targeting construct, K. Deisseroth (Stanford University) for the WPRE-containing DNA construct, B. Sauer (National Institute of Diabetes and Digestive and Kidney Diseases) via Addgene for the Cre and EGFP-Cre DNA constructs, P. Chambon (Institut de Génétique et de Biologie Moléculaire et Cellulaire) for the CreERT2 DNA construct, P. Soriano (Fred Hutchinson Cancer Research Center) via Addgene for the PhiC31o, FLPo and pPGKneotpAlox2 DNA constructs, A. Nagy (Mount Sinai Hospital in Toronto) for the G4 ES cell line, G. Oliver (St. Jude's Children's Research Hospital) for the Six3-Cre mice, N. Heintz (Rockefeller University) via Mutant Mouse Regional Resource Centers (MMRRC) for the Ntsr1-Cre mice and X. Zhuang (University of Chicago) for the Slc6a3-Cre mice. The authors thank the Allen Institute founders, P.G. Allen and J. Patton, for their vision, encouragement and support. This work was funded by the Allen Institute for Brain Science and a US National Institutes of Health grant (MH085500) to H.Z.
Author information
Authors and Affiliations
Contributions
H.Z. designed the study, analyzed data and wrote the paper. L.M. generated the Cre reporter lines and the knock-in Cre driver lines. T.A.Z. and E.S.L. designed and generated the BAC transgenic Cre driver lines. H.Z., S.M.S. and T.A.Z. set up the characterization pipeline and database. S.W.O. produced and performed experiments with rAAVs. H.A.Z. performed in vivo two-photon imaging experiments. M.J.H. and L.L.N. conducted informatics analysis. H.G. assisted with transgenic mice production and characterization. R.D.P. provided lab resources and scientific advice. A.R.J. provided institutional support.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 (PDF 5331 kb)
Rights and permissions
About this article
Cite this article
Madisen, L., Zwingman, T., Sunkin, S. et al. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat Neurosci 13, 133–140 (2010). https://doi.org/10.1038/nn.2467
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.2467
This article is cited by
-
Neurotrophin-3 from the dentate gyrus supports postsynaptic sites of mossy fiber-CA3 synapses and hippocampus-dependent cognitive functions
Molecular Psychiatry (2024)
-
Acoustically targeted noninvasive gene therapy in large brain volumes
Gene Therapy (2024)
-
Basal ganglia–spinal cord pathway that commands locomotor gait asymmetries in mice
Nature Neuroscience (2024)
-
Long-term labeling and imaging of synaptically connected neuronal networks in vivo using double-deletion-mutant rabies viruses
Nature Neuroscience (2024)
-
Optogenetic recruitment of hypothalamic corticotrophin-releasing-hormone (CRH) neurons reduces motivational drive
Translational Psychiatry (2024)