Abstract
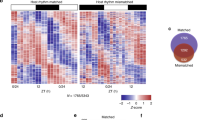
The Earth's rotation forced life to evolve under cyclic day and night environmental changes. To anticipate such daily cycles, prokaryote and eukaryote free-living organisms evolved intrinsic clocks that regulate physiological and behavioural processes. Daily rhythms have been observed in organisms living within hosts, such as parasites. Whether parasites have intrinsic molecular clocks or whether they simply respond to host rhythmic physiological cues remains unknown. Here, we show that Trypanosoma brucei, the causative agent of human sleeping sickness, has an intrinsic circadian clock that regulates its metabolism in two different stages of the life cycle. We found that, in vitro, ∼10% of genes in T. brucei are expressed with a circadian rhythm. The maximum expression of these genes occurs at two different phases of the day and may depend on a post-transcriptional mechanism. Circadian genes are enriched in cellular metabolic pathways and coincide with two peaks of intracellular adenosine triphosphate concentration. Moreover, daily changes in the parasite population lead to differences in suramin sensitivity, a drug commonly used to treat this infection. These results demonstrate that parasites have an intrinsic circadian clock that is independent of the host, and which regulates parasite biology throughout the day.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Young, M. W. & Kay, S. A. Time zones: a comparative genetics of circadian clocks. Nat. Rev. Genet. 2, 702–715 (2001).
Rosbash, M. The implications of multiple circadian clock origins. PLoS Biol. 7, e1000062 (2009).
Bell-Pedersen, D. et al. Circadian rhythms from multiple oscillators: lessons from diverse organisms. Nat. Rev. Genet. 6, 544–556 (2005).
Welsh, D. K., Yoo, S. H., Liu, A. C., Takahashi, J. S. & Kay, S. A. Bioluminescence imaging of individual fibroblasts reveals persistent, independently phased circadian rhythms of clock gene expression. Curr. Biol. 14, 2289–2295 (2004).
Balsalobre, A., Damiola, F. & Schibler, U. A serum shock induces circadian gene expression in mammalian tissue culture cells. Cell 93, 929–937 (1998).
Buhr, E. D., Yoo, S. H. & Takahashi, J. S. Temperature as a universal resetting cue for mammalian circadian oscillators. Science 330, 379–385 (2010).
Refinetti, R. & Menaker, M. The circadian rhythm of body temperature. Physiol. Behav. 51, 613–637 (1992).
Zhang, R., Lahens, N. F., Ballance, H. I., Hughes, M. E. & Hogenesch, J. B. A circadian gene expression atlas in mammals: implications for biology and medicine. Proc. Natl Acad. Sci. USA 111, 16219–16224 (2014).
van der Linden, A. M. et al. Genome-wide analysis of light- and temperature-entrained circadian transcripts in Caenorhabditis elegans. PLoS Biol. 8, e1000503 (2010).
Hughes, M. E., Grant, G. R., Paquin, C., Qian, J. & Nitabach, M. N. Deep sequencing the circadian and diurnal transcriptome of Drosophila brain. Genome Res. 22, 1266–1281 (2012).
Mony, B. M. et al. Genome-wide dissection of the quorum sensing signalling pathway in Trypanosoma brucei. Nature 505, 681–685 (2013).
Dejung, M. et al. Quantitative proteomics uncovers novel factors involved in developmental differentiation of Trypanosoma brucei. PLoS Pathog. 12, e1005439 (2016).
Kabani, S. et al. Genome-wide expression profiling of in vivo-derived bloodstream parasite stages and dynamic analysis of mRNA alterations during synchronous differentiation in Trypanosoma brucei. BMC Genomics 10, 427 (2009).
Trindade, S. et al. Trypanosoma brucei parasites occupy and functionally adapt to the adipose tissue in mice. Cell Host Microbe 19, 837–848 (2016).
Nilsson, D. et al. Spliced leader trapping reveals widespread alternative splicing patterns in the highly dynamic transcriptome of Trypanosoma brucei. PLoS Pathog. 6, e1001037 (2010).
Koike, N. et al. Transcriptional architecture and chromatin landscape of the core circadian clock in mammals. Science 338, 349–354 (2012).
Hoffmann, J. et al. Non-circadian expression masking clock-driven weak transcription rhythms in U2OS cells. PLoS ONE 9, e102238 (2014).
Hughes, M. E. et al. Harmonics of circadian gene transcription in mammals. PLoS Genet. 5, e1000442 (2009).
Izumo, M. et al. Differential effects of light and feeding on circadian organization of peripheral clocks in a forebrain Bmal1 mutant. eLife 3, e04617 (2014).
Reddy, A. B. & Rey, G. Metabolic and nontranscriptional circadian clocks: eukaryotes. Annu. Rev. Biochem. 83, 165–189 (2014).
Figueiredo, L. M., Cross, G. A. & Janzen, C. J. Epigenetic regulation in African trypanosomes: a new kid on the block. Nat. Rev. Microbiol. 7, 504–513 (2009).
Clayton, C. E. Life without transcriptional control? From fly to man and back again. EMBO J. 21, 1881–1888 (2002).
Asio, S. M., Simonsen, P. E. & Onapa, A. W. Analysis of the 24-h microfilarial periodicity of Mansonella perstans. Parasitol. Res. 104, 945–948 (2009).
Lindstrom, K. M. et al. Feather mites and internal parasites in small ground finches (Geospiza fuliginosa, Emberizidae) from the Galapagos Islands (Equador). J. Parasitol. 95, 39–45 (2009).
Gryczynska, A., Dolnik, O. & Mazgajski, T. D. Parasites of chaffinch (Fringilla coelebs) population. Part I. Coccidia (Protozoa, Apicomplexa). Wiad. Parazytol. 45, 495–500 (1999).
Dolnik, O. V., Metzger, B. J. & Loonen, M. J. Keeping the clock set under the midnight sun: diurnal periodicity and synchrony of avian Isospora parasites cycle in the high Arctic. Parasitology 138, 1077–1081 (2011).
Hawking, F. The clock of the malaria parasite. Sci. Am. 222, 123–131 (1970).
Thaiss, C. A. et al. Transkingdom control of microbiota diurnal oscillations promotes metabolic homeostasis. Cell 159, 514–529 (2014).
Zarrinpar, A., Chaix, A., Yooseph, S. & Panda, S. Diet and feeding pattern affect the diurnal dynamics of the gut microbiome. Cell Metab. 20, 1006–1017 (2014).
Curtis, A. M., Bellet, M. M., Sassone-Corsi, P. & O'Neill, L. A. Circadian clock proteins and immunity. Immunity 40, 178–186 (2014).
Brady, J. & Crump, A. J. The control of circadian activity rhythms in tsetse flies: environment or physiological clock? Physiol. Entomol. 3, 177–190 (1978).
MacGregor, P., Savill, N. J., Hall, D. & Matthews, K. R. Transmission stages dominate trypanosome within-host dynamics during chronic infections. Cell Host Microbe. 9, 310–318 (2011).
Bass, J. & Takahashi, J. S. Circadian integration of metabolism and energetics. Science 330, 1349–1354 (2010).
Engstler, M. & Boshart, M. Cold shock and regulation of surface protein trafficking convey sensitization to inducers of stage differentiation in Trypanosoma brucei. Genes Dev. 18, 2798–2811 (2004).
Hirumi, H. & Hirumi, K. Continuous cultivation of Trypanosoma brucei blood stream forms in a medium containing a low concentration of serum protein without feeder cell layers. J. Parasitol. 75, 985–989 (1989).
Knusel, S. & Roditi, I. Insights into the regulation of GPEET procyclin during differentiation from early to late procyclic forms of Trypanosoma brucei. Mol. Biochem. Parasitol. 191, 66–74 (2013).
Cox, M. P., Peterson, D. A. & Biggs, P. J. SolexaQA: at-a-glance quality assessment of Illumina second-generation sequencing data. BMC Bioinformatics 11, 485 (2010).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Lawrence, M. et al. Software for computing and annotating genomic ranges. PLoS Comput. Biol. 9, e1003118 (2013).
Pages, H., Aboyoun, P., Gentleman, R. & DebRoy, S. Biostrings: String Objects Representing Biological Sequences, and Matching Algorithms. R package v.2.38.4 (Bioconductor, 2016); https://bioconductor.org/packages/release/bioc/html/Biostrings.html
Lawrence, M., Gentleman, R. & Carey, V. Rtracklayer: an R package for interfacing with genome browsers. Bioinformatics 25, 1841–1842 (2009).
Huber, W. et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat. Methods 12, 115–121 (2015).
Warnes, G. R. et al. gplots: Various R Programming Tools for Plotting Data. R package v.3.0.1 (CRAN, 2016); https://cran.r-project.org/web/packages/gplots/index.html
Wichert, S., Fokianos, K. & Strimmer, K. Identifying periodically expressed transcripts in microarray time series data. Bioinformatics 20, 5–20 (2004).
Hughes, M. E., Hogenesch, J. B. & Kornacker, K. JTK_CYCLE: an efficient nonparametric algorithm for detecting rhythmic components in genome-scale data sets. J. Biol. Rhythms 25, 372–380 (2010).
Yang, R. & Su, Z. Analyzing circadian expression data by harmonic regression based on autoregressive spectral estimation. Bioinformatics 26, i168–i174 (2010).
Kurt Hornik, B. G. movMF: an R package for fitting mixtures of von Mises–Fisher distributions. J. Stat. Softw. 58, 1–31 (2014).
Siepka, S. M. & Takahashi, J. S. Methods to record circadian rhythm wheel running activity in mice. Methods Enzymol. 393, 230–239 (2005).
Siepka, S. M. et al. Circadian mutant overtime reveals F-box protein FBXL3 regulation of cryptochrome and period gene expression. Cell 129, 1011–1023 (2007).
Falcon, S. & Gentleman, R. Using GOstats to test gene lists for GO term association. Bioinformatics 23, 257–258 (2007).
Acknowledgements
The authors thank M. Phillips for help with starting parasite cultures at UT Southwestern and advice on the suramin sensitivity experiments, P. Nakashe for the library preparations of the light/dark data sets, M. Broderick for help during the oxidative stress experiment, L. Pinho, I. Kornblum and G. Kilaru for technical support, J. Stubblefield for help during the blood collection, D. Barry, M. Phillips, C. Green and M. Vaz for reading the manuscript and C. Janzen for the GFP-PAD1utr cell line. The work was supported by an HHMI International Early Career Scientist award (55007419, to L.M.F.), EMBO Installation Grant (2151) to L.M.F., Fundação para a Ciência e Tecnologia awards (SFRH/BD/51286/2010 to F.R.-F. and IF/00595/2014 to N.L.B.-M.). J.S.T. is an Investigator in the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
F.R-F., L.M.F. and J.S.T. designed the study. F.R.-F. performed the experiments. D.P.-N., F.R.-F., N.L.B.-M., L.M.F. and J.S.T analysed the data. F.R.-F. wrote the manuscript and all authors contributed to reviewing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1–10 and Supplementary Tables 1 and 2. (PDF 11397 kb)
Supplementary Data 1
List of genes, their RPKM counts and circadian algorithms analyses for bloodstream forms entrained with temperature. (XLSX 1626 kb)
Supplementary Data 2
List of genes, their RPKM counts and circadian algorithms analyses for insect procyclic forms entrained with temperature. (XLSX 1241 kb)
Supplementary Data 3
List of genes, their RPKM counts and circadian algorithms analyses for bloodstream forms entrained with light. (XLSX 382 kb)
Supplementary Data 4
GO term enrichment analysis of bloodstream and procyclic forms throughout the day. (XLSX 39 kb)
Rights and permissions
About this article
Cite this article
Rijo-Ferreira, F., Pinto-Neves, D., Barbosa-Morais, N. et al. Trypanosoma brucei metabolism is under circadian control. Nat Microbiol 2, 17032 (2017). https://doi.org/10.1038/nmicrobiol.2017.32
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2017.32
This article is cited by
-
Ecology of asynchronous asexual replication: the intraerythrocytic development cycle of Plasmodium berghei is resistant to host rhythms
Malaria Journal (2021)
-
Daily rhythms in gene expression of the human parasite Schistosoma mansoni
BMC Biology (2021)
-
Malaria parasites regulate intra-erythrocytic development duration via serpentine receptor 10 to coordinate with host rhythms
Nature Communications (2020)
-
Genomics of circadian rhythms in health and disease
Genome Medicine (2019)
-
African trypanosomes
Parasites & Vectors (2019)