Abstract
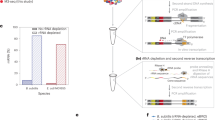
Intracellular bacterial pathogens can exhibit large heterogeneity in growth rate inside host cells, with major consequences for the infection outcome. If and how the host responds to this heterogeneity remains poorly understood. Here, we combined a fluorescent reporter of bacterial cell division with single-cell RNA-sequencing analysis to study the macrophage response to different intracellular states of the model pathogen Salmonella enterica serovar Typhimurium. The transcriptomes of individual infected macrophages revealed a spectrum of functional host response states to growing and non-growing bacteria. Intriguingly, macrophages harbouring non-growing Salmonella display hallmarks of the proinflammatory M1 polarization state and differ little from bystander cells, suggesting that non-growing bacteria evade recognition by intracellular immune receptors. By contrast, macrophages containing growing bacteria have turned into an anti-inflammatory, M2-like state, as if fast-growing intracellular Salmonella overcome host defence by reprogramming macrophage polarization. Additionally, our clustering approach reveals intermediate host functional states between these extremes. Altogether, our data suggest that gene expression variability in infected host cells shapes different cellular environments, some of which may favour a growth arrest of Salmonella facilitating immune evasion and the establishment of a long-term niche, while others allow Salmonella to escape intracellular antimicrobial activity and proliferate.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
14 July 2017
In the PDF version of this article previously published, the year of publication provided in the footer of each page and in the 'How to cite' section was erroneously given as 2017, it should have been 2016. This error has now been corrected. The HTML version of the article was not affected.
References
Bumann, D. Heterogeneous host–pathogen encounters: act locally, think globally. Cell Host Microbe 17, 13–19 (2015).
Price, J. V. & Vance, R. E. The macrophage paradox. Immunity 41, 685–693 (2014).
Avraham, R. et al. Pathogen cell-to-cell variability drives heterogeneity in host immune responses. Cell 162, 1309–1321 (2015).
Monack, D. M., Mueller, A. & Falkow, S. Persistent bacterial infections: the interface of the pathogen and the host immune system. Nat. Rev. Microbiol. 2, 747–765 (2004).
Claudi, B. et al. Phenotypic variation of salmonella in host tissues delays eradication by antimicrobial chemotherapy. Cell 158, 722–733 (2014).
Helaine, S. et al. Internalization of Salmonella by macrophages induces formation of nonreplicating persisters. Science 343, 204–208 (2014).
Helaine, S. et al. Dynamics of intracellular bacterial replication at the single cell level. Proc. Natl Acad. Sci. USA 107, 3746–3751 (2010).
Jenner, R. G. & Young, R. A. Insights into host responses against pathogens from transcriptional profiling. Nat. Rev. Microbiol. 3, 281–294 (2005).
Saliba, A. E., Westermann, A. J., Gorski, S. A. & Vogel, J. Single-cell RNA-seq: advances and future challenges. Nucleic Acids Res. 42, 8845–8860 (2014).
Shalek, A. K. et al. Single-cell RNA-seq reveals dynamic paracrine control of cellular variation. Nature 510, 363–369 (2014).
Keestra-Gounder, A. M., Tsolis, R. M. & Bäumler, A. J. Now you see me, now you don't: the interaction of Salmonella with innate immune receptors. Nat. Rev. Microbiol. 13, 206–216 (2015).
Eisele, N. A. et al. Salmonella require the fatty acid regulator PPARδ for the establishment of a metabolic environment essential for long-term persistence. Cell Host Microbe 14, 171–182 (2013).
Kreibich, S. & Hardt, W. D. Experimental approaches to phenotypic diversity in infection. Curr. Opin. Microbiol. 27, 25–36 (2015).
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096–1098 (2013).
Eisenreich, W., Heesemann, J., Rudel, T. & Goebel, W. Metabolic host responses to infection by intracellular bacterial pathogens. Front. Cell Infect. Microbiol. 3, 24 (2013).
Lawrence, T. & Natoli, G. Transcriptional regulation of macrophage polarization: enabling diversity with identity. Nat. Rev. Immunol. 11, 750–761 (2011).
Kohyama, M. et al. Role for Spi-C in the development of red pulp macrophages and splenic iron homeostasis. Nature 457, 318–321 (2009).
Nix, R. N., Altschuler, S. E., Henson, P. M. & Detweiler, C. S. Hemophagocytic macrophages harbor Salmonella enterica during persistent infection. PLoS Pathog. 3, e193 (2007).
Guilliams, M., Bruhns, P., Saeys, Y., Hammad, H. & Lambrecht, B. N. The function of Fcγ receptors in dendritic cells and macrophages. Nat. Rev. Immunol. 14, 94–108 (2014).
Martinez, F. O., Helming, L. & Gordon, S. Alternative activation of macrophages: an immunologic functional perspective. Annu. Rev. Immunol. 27, 451–483 (2009).
Bronte, V. & Zanovello, P. Regulation of immune responses by l-arginine metabolism. Nat. Rev. Immunol. 5, 641–654 (2005).
De Jong, H. K. et al. Expression and function of S100A8/A9 (calprotectin) in human typhoid fever and the murine Salmonella model. PLOS Negl. Trop. Dis. 9, e0003663 (2015).
Ashkar, S. et al. Eta-1 (osteopontin): an early component of type-1 (cell-mediated) immunity. Science 287, 860–864 (2000).
Benoit, M., Desnues, B. & Mege, J. L. Macrophage polarization in bacterial infections. J. Immunol. 181, 3733–3739 (2008).
McCoy, M. W., Moreland, S. M. & Detweiler, C. S. Hemophagocytic macrophages in murine typhoid fever have an anti-inflammatory phenotype. Infect. Immun. 80, 3642–3649 (2012).
Vazquez-Torres, A. et al. Salmonella pathogenicity island 2-dependent evasion of the phagocyte NADPH oxidase. Science 287, 1655–1658 (2000).
Burton, N. A. et al. Disparate impact of oxidative host defenses determines the fate of Salmonella during systemic infection in mice. Cell Host Microbe 15, 72–83 (2014).
Tahoun, A. et al. Salmonella transforms follicle-associated epithelial cells into M cells to promote intestinal invasion. Cell Host Microbe 12, 645–656 (2012).
Masaki, T. et al. Reprogramming adult Schwann cells to stem cell-like cells by leprosy bacilli promotes dissemination of infection. Cell 152, 51–67 (2013).
Westermann, A. J. et al. Dual RNA-seq unveils noncoding RNA functions in host–pathogen interactions. Nature 529, 496–501 (2016).
Figueira, R., Watson, K. G., Holden, D. W. & Helaine, S. Identification of Salmonella pathogenicity Island-2 type III secretion system effectors involved in intramacrophage replication of S. enterica serovar typhimurium: implications for rational vaccine design. mBio 4, e00065-13 (2013).
Picelli, S. et al. Full-length RNA-seq from single cells using smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Kapteyn, J., He, R., McDowell, E. T. & Gang, D. R. Incorporation of non-natural nucleotides into template-switching oligonucleotides reduces background and improves cDNA synthesis from very small RNA samples. BMC Genomics 11, 413 (2010).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with bowtie 2. Nat. Methods 9, 357–359 (2012).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-seq. Bioinformatics 25, 1105–1111 (2009).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with topHat and cufflinks. Nat. Protoc. 7, 562–578 (2012).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Brennecke, P. et al. Accounting for technical noise in single-cell RNA-seq experiments. Nat. Methods 10, 1093–1095 (2013).
Lê, S., Josse, J. & Husson, F. Factominer: an R package for multivariate analysis. J. Stat. Soft. 25, 1–18 (2008).
Kharchenko, P. V., Silberstein, L. & Scadden, D. T. Bayesian approach to single-cell differential expression analysis. Nat. Methods 11, 740–742 (2014).
Fan, J. et al. Characterizing transcriptional heterogeneity through pathway and gene set overdispersion analysis. Nat. Methods 13, 241–244 (2016).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Grün, D. et al. Single-cell messenger RNA sequencing reveals rare intestinal cell types. Nature 525, 251–255 (2015).
Acknowledgements
The authors thank T. Achmedov, V. McParland, H. Merkert and B. Plaschke for technical support. A.-E.S. was supported by the PostDoc Plus program of the University of Würzburg. A.J.W. was the recipient of an Elite Advancement PhD stipend from Universität Bayern e.V. S.H. was supported by an MRC Career Development Award (MR/M009629/1). D.A.C.S. was the recipient of an EMBO postdoctoral fellowship (ALTF 441-2015).
Author information
Authors and Affiliations
Contributions
A.-E.S., A.J.W. and J.V. designed and A.-E.S. performed the experiments. L.L. and S.A. performed bioinformatics analysis. S.H. and D.A.C.S. performed flow cytometry experiments. A.J.W., S.H. and L.N.S. provided reagents. A.-E.S., A.J.W., S.H. and J.V. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary Figures 1–12, Supplementary References (PDF 1900 kb)
Supplementary Table 1
Sequencing results summary (XLSX 15 kb)
Supplementary Table 2
Cell clustering and differentially expressed genes between subpopulations (XLSX 1371 kb)
Rights and permissions
About this article
Cite this article
Saliba, AE., Li, L., Westermann, A. et al. Single-cell RNA-seq ties macrophage polarization to growth rate of intracellular Salmonella. Nat Microbiol 2, 16206 (2017). https://doi.org/10.1038/nmicrobiol.2016.206
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2016.206