Abstract
Insights into the genomic adaptive traits of Treponema pallidum, the causative bacterium of syphilis, have long been hampered due to the absence of in vitro culture models and the constraints associated with its propagation in rabbits. Here, we have bypassed the culture bottleneck by means of a targeted strategy never applied to uncultivable bacterial human pathogens to directly capture whole-genome T. pallidum data in the context of human infection. This strategy has unveiled a scenario of discreet T. pallidum interstrain single-nucleotide-polymorphism-based microevolution, contrasting with a rampant within-patient genetic heterogeneity mainly targeting multiple phase-variable loci and a major antigen-coding gene (tprK). TprK demonstrated remarkable variability and redundancy, intra- and interpatient, suggesting ongoing parallel adaptive diversification during human infection. Some bacterial functions (for example, flagella- and chemotaxis-associated) were systematically targeted by both inter- and intrastrain single nucleotide polymorphisms, as well as by ongoing within-patient phase variation events. Finally, patient-derived genomes possess mutations targeting a penicillin-binding protein coding gene (mrcA) that had never been reported, unveiling it as a candidate target to investigate the impact on the susceptibility to penicillin. Our findings decode the major genetic mechanisms by which T. pallidum promotes immune evasion and survival, and demonstrate the exceptional power of characterizing evolving pathogen subpopulations during human infection.
Similar content being viewed by others
The spirochaete Treponema pallidum subspecies pallidum is the causative agent of syphilis. This bacterium is usually transmitted by sexual contact or from mother to infant before or at the time of birth, but can also be transmitted by blood transfusion1,2. Syphilis remains a global problem, which may in part be attributed to the absence of a vaccine to prevent infection and transmission3.
The extreme ex vivo fragility of T. pallidum has contributed to the inability to culture it in vitro, with in vivo culture maintenance only being possible following intratesticular or intradermal inoculation of rabbits1. Despite this, in one of the pioneer microbial whole-genome sequencing (WGS) projects, Fraser and colleagues4 in 1998 sequenced the genome of T. pallidum. Almost two decades later, only the genomes of five other strains have been released, all of them obtained using the same restrictive culture strategy5–10. Treponema pallidum was found to possess a small genome of ∼1.1 Mb characterized by a striking lack of metabolic capabilities, indicating that this pathogen is highly adapted and extremely dependent on the mammalian host4. Moreover, there is a high nucleotide similarity among the few sequenced T. pallidum genomes, with no described mechanisms of intra-subspecies horizontal gene transfer11, suggesting that phenotypic differences may arise from subtle genetic changes11,12. In particular, remarkable efforts using the rabbit model have indicated that targeted mechanisms of gene conversion13–15 and in-length variation of homopolymeric tracts16–18 (driving on/off switching phase variation) may be critical for generating the genetic diversity contributing to pathogen survival and adaptation within the host. Other findings have singled out potential determinants of T. pallidum pathogenesis, highlighting the likely surface-exposed proteins encoded by a 12-member paralogue of the T. pallidum repeat (Tpr) gene family16,19,20. Within this family, which accounts for ∼2% of the genome, the antigen-encoding tprK has been the focus of extensive research due to its putative pivotal role in immune evasion and pathogen persistence13–15,21–26. However, because T. pallidum proteins are in general not fully characterized in terms of their structure, topology, location and antigenicity, assumptions regarding their impact on T. pallidum biology and pathogenesis have not always been consensual27–29. Furthermore, little is known about how T. pallidum mediates adaptation and virulence and which pathogen features (genetic and phenotypic) determine some specific traits, such as the invasion of the central nervous system2,11. This lack of knowledge is partially caused by the experimental constraints underlying the acquisition of extensive and consistent genomic data and the identification and genome mapping of allelic variation within human in vivo T. pallidum populations, which has skewed our understanding of the epidemiology and pathobiology of syphilitic strains.
Targeted WGS of uncultured bacteria from complex DNA populations, such as clinical samples, has only recently taken its first steps due to the difficulties associated with purification of the target microorganism. A recently developed cutting-edge methodology, based on selective enrichment of the desired DNA through hybridization with RNA oligonucleotides (‘baits’)30, has only recently been applied to the direct WGS of bacteria from clinical specimens31–33. In the present study, this culture-independent targeted-WGS strategy was successfully applied to fully recover the genome sequence of the syphilis-causing agent T. pallidum directly from multiple clinical samples. This ‘in vivo’ approach constitutes the first large-scale genome-based insight into the genetic diversity of T. pallidum in the context of human infection, revealing extensive within-patient genetic variation of this pathogen, probably as a means to achieve immune evasion and persistence.
Results
WGS of T. pallidum directly from clinical samples
The application of ‘SureSelectXT Target Enrichment’ technology coupled with WGS allowed the full capture and sequencing of 24 of 34 genomes of the uncultivable T. pallidum bacterium directly from clinical samples (Supplementary Table 1). Real-time quantitative PCR (qPCR) data showed that the proportion of reads mapping to the target genome is mainly linked to the number of bacterial copies within the input DNA samples and does not seem to depend on the degree of human DNA contamination (Supplementary Fig. 1a). Enrichment success was obtained exclusively for samples with more than 1 × 104T. pallidum copies (Supplementary Fig. 1a,b). The median depth of coverage for the novel genomes was 131× (ranging from 20× to 1,196×) (Supplementary Table 1).
T. pallidum pallidum tree and diversity
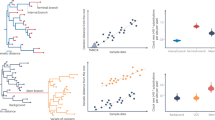
Phylogenetic reconstruction within the T. pallidum pallidum was performed using PT_SIF genomes (this study) and six reference genomes (obtained after bacterial propagation in rabbit testis) available at GenBank4–10 (Fig. 1a). Phylogeny comprised whole-genome sequences excluding arp and tprK, due to their well-described sequence repetitions34 and gene conversion mechanisms13–15, respectively. tprK clearly biases distance estimations and phylogenetic inferences6,21 (Supplementary Fig. 2). All clinical strains were found to be segregated in a separate branch (clade I) and to be very closely related to the Street Strain 14 (SS14) isolate (collected in 1977 in Atlanta, USA) (Fig. 1a). Clade I, which also includes the Mexico A strain, diverges from the two other clades (II and III) by more than ∼700 nucleotide differences (Supplementary Table 2). Nevertheless, about half of these polymorphisms are due to the differential presence of highly heterogeneous tprD alleles between strains6,35. Of note, and in contrast, for instance, to the reference genome Nichols-RS, which harbours identical copies of tprC and tprD (ref. 6), PT_SIF clinical strains carry the well-described tprD2 allele, which is non-identical to tprC. Single nucleotide polymorphism (SNP) density analysis throughout the genome revealed that the segregation of clades is mainly supported by mutations in a restricted set of 13 genes (that constitute ∼1% of all T. pallidum genes but account for ∼70% of all single nucleotide variant sites (SNVs)) (Fig. 2). Overall, we found a single non-synonymous variant site that distinguishes all clinical strains from all reference strains (including the closely related SS14) (Fig. 1b). This variant site affects TPASS_20705/mrcA, which codes for a bifunctional transglycosylase/transpeptidase penicillin-binding protein. Regarding the divergence within clade I, whereas SS14 branch segregation is marked by an overrepresentation of synonymous (12/19) and homoplasic mutations (15/19), microevolution of the clinical strains unveiled a mutational signature that probably reflects pathogen adaptation to its human host.
a, Whole-genome based Neighbor-Joining phylogenetic tree, showing three main clades (I, II and III). The genetic relationships within the novel PT_SIF genomes sequenced directly from clinical samples is zoomed in (bootstrap values with 1,000 replicates are shown next to the branch nodes; arp and tprK genes were excluded from the analysis). The only SNP (targeting the TPASS_20705/mrcA) that was found to segregate all clinical strains from all reference strains is depicted near the tree. b,c, Profile of genetic diversity within PT_SIF genomes showing SNV sites with both fixed (b) or probable emerging (c) mutations, when compared with the genome sequence of the most closely related ancestral strain (SS14 RS). To avoid duplication of SNV sites, probable emerging mutations occurring in the same site as fixed mutations are displayed in b rather than in c. Gene designations (above the chart) and nucleotide positions (below the chart) of the SNVs are relative to the SS14 RS genome (accession no. CP004011.1). Genes are ordered according to the putative functional categories, and names highlighted in blue refer to genes displaying more than one SNV site among the PT_SIF genomes and appear repeated in b and/or c. *Homoplasic SNV sites shared with a strain from clusters II and/or III. †Given that the ancestral genome of the SS14 strain already carried the mutation (A2058G) in both copies of the 23SrRNA locus associated with macrolides resistance6, the mutational profile refers to the presence of these mutations within the PT_SIF group. IGR; intergenic region.
SNV site density across a whole-genome alignment enrolling all T. pallidum pallidum genomes sequenced directly from clinical samples and reference genomes (excluding tprK and arp genes) over a sliding window. Genes highlighted above the graph represent top polymorphic genes. For each gene, polymorphism was essentially given by strains from *clade I, II and III; †clade II/III or I; ‡clade I, only Mexico A strain; §clade I, SS14 plus Mexico A strains; ||clade III. As the Mexico A genome carries tRNA annotations remarkably divergent from the other strains, suggesting potential misannotation, these were discarded from the analysis. tprJ polymorphim is particularly inflated by 275 nucleotide differences occurring exclusively for the Sea 81-4 strain. The huge amount of nucleotide differences occurring in tprD is due to the existence of two distinct alleles for this gene within T. pallidum pallidum.
Microevolutionary expansion of the patient-derived PT_SIF ‘clone’
SNP-based diversity revealed a high genetic homogeneity among the PT_SIF clinical strains' genomes marked by 35 SNV sites, of which 32 are PT_SIF-specific (Fig. 1b). They revealed a mean pairwise nucleotide distance of 9 ± 2 SNPs (maximum of 18 SNPs). However, their microevolutionary expansion mainly relied on the fixation (or near-fixation) of non-synonymous mutations (26/32 SNVs occurring in coding regions). Some strains could also be discriminated by three small indels, seven silent SNPs and one nucleotide replacement (A2058G) in each of the two copies of the 23SrRNA (Fig. 1b). It is noteworthy that the latter, which is associated with bacterial resistance to macrolides36, emerged in separate phylogenetic branches, being already fixed or nearly fixed in 19 of the 25 clinical strains probably reflecting antibiotic-driven selective pressure. The set of genes targeted by microevolution essentially involves genes encoding known or putative membrane proteins, flagella-associated proteins, chemotaxis-associated proteins and proteins without predicted function4,11,37. Besides this, from inspection of SNP-based intrastrain heterogeneity, which probably reflects ongoing adaptive diversification, 16 of the 25 PT_SIF populations revealed allelic variation affecting single nucleotide sites (excluding homopolymeric tracts and tprK) (Fig. 1b,c). Overall, 63 intrastrain heterogeneous sites were detected, where eight in every ten of the potentially emerging mutations are non-synonymous or inactivating mutations. In contrast to the emerging mutation in 23SrRNA that is probably under fixation, the same cannot be stated for the remaining mutations, particularly for those leading to protein truncation, as the genes' essentiality would be questioned. Nevertheless, all reported intrastrain mutations are already present with more than 10% frequency, and the observed scenario (Fig. 1c) remarkably parallels the scenario seen for the pool of fixed or nearly fixed mutations (Fig. 1b), as the likely emerging mutations targeted the same genes or gene categories and are clearly overrepresented by potentially adaptive non-synonymous mutations. In support of this, we also observed (1) genes targeted by more than one mutation; (2) genes presenting both fixed and emerging mutations affecting distinct nucleotide positions, and (3) genes carrying both intra- and interstrain mutations. Interestingly, TPASS_20705/mrcA not only determines the segregation of all clinical strains (as described above), but has also been targeted during the ongoing expansion of the T. pallidum PT_SIF ‘clone’ within the human host, already revealing two additional non-synonymous mutations either fixed (in 20 of the 25 strains) or probably emerging (Fig. 1b). It cannot be discounted that, similar to the scenario observed for 23SrRNA mutations, the over-targeting of mrcA may rely on its function, as it encodes a penicillin-binding protein. Interestingly, the already fixed non-synonymous mutation affects the C-terminus transpeptidase domain, which includes the beta-lactamase active-site serine in other bacteria38.
Phase variation mediated by variable homopolymeric tracts
Cumulative evidence supports that reversible expansion and contraction of homopolymeric tracts (poly(G) and poly(C)) underlies phase variation mechanisms in T. pallidum, as a mean of this pathogen to quickly generate phenotypic diversity towards host adaptation16–18. As such, 46 chromosome-dispersed homopolymeric tracts were identified and analysed, including poly(G) and poly(C) strings falling equally within coding regions (probably mediating a classical on/off switching mechanism) and putative regulatory regions (probably affecting expression at the transcriptional or translational level) (Table 1). As indicated in Table 1 and Supplementary Table 3, it should be taken into account that the precise location of some poly(G/C) tracts in relation to the potential target gene is not always straightforward, as a consequence of both annotation incongruence and the general lack of in-depth characterization of T. pallidum genes. Thirty-one out of the 46 tracts revealed intrastrain in-length variability in at least one clinical strain, with 17 of those tracts being variable in more than 50% of clinical strains (Fig. 3; Table 1). In ensuring the reliability of these results, (1) homopolymer-associated errors have been shown to be tremendously minimized using Illumina technology39–42, (2) similar strategies have been used to support that these variable regions mediate relevant alterations in virulence-associated genes39,40, and (3) 15 T. pallidum poly(G/C) tracts have consistently been found to present no variation across all clinical strains, which excludes Illumina bias and constitutes a good proof of principle for our approach. It is noteworthy that a vast number of cases have been found where, although a given homopolymeric tract was conserved within a particular T. pallidum population, it remarkably displayed different nucleotide lengths between populations (Fig. 3 and Supplementary Table 3). For instance, for TPANIC_0126, which was recently described to be transcriptionally regulated by phase variation18, the poly(G/C) presents conserved lengths within all populations (except one) but the dominant base count is different between clinical strains (Fig. 3 and Supplementary Table 3). Additionally, for variable poly(G/C) tracts directly affecting annotated coding regions, we observed that the dominant count in several populations probably yields protein truncation. This phenomenon occurred for 11 distinct homopolymeric tracts, and the affected genes mainly encode hypothetical proteins, a putative methyl-accepting chemotaxis protein (TPANIC_0040/mcp1) and putative membrane proteins (Table 1). Among the genes whose transcription levels are potentially modulated by phase variation, we highlight genes from the tpr family, putative outer and inner membrane protein-encoding genes and a gene encoding a flagellar motor switch protein (TPANIC_0026/fliG1). Altogether, although some of these genes have been described to be regulated by poly(G/C)-driven phase-variation mechanisms, detected either in vitro or using the rabbit model16–18, our findings report, for the first time, the existence of heterogeneous homopolymeric tracts, probably yielding T. pallidum phenotypic diversity within the human host.
Data are presented for 31 intrastrain variable homopolymeric poly(G/C) tracts, where each graph displays the relative percentage of sequence reads with a particular base count for five representative T. pallidum in vivo populations (for complete data see Supplementary Table 3). A fully conserved (both intra- and interstrain) homopolymeric tract (targeting TPANIC_0059) is also shown for comparison purposes. Names for genes (displayed above the graphs) potentially targeted by phase variation are based on the Nichols RS genome (accession no. CP004010.2) and are coloured according to the relative position of the poly(G/C): in the putative regulatory region (blue), within the coding region (red) or an unpredictable target (black) (for the latter, the Nichols RS genome position is indicated). *Due to incongruences in gene annotations among reference genomes, the impact of these poly(G/C) tracts on the potential target gene may not be the one suggested, because the poly(G/C) falls either on the putative regulatory region or within the coding sequence depending on the annotated genome. †No annotation in the Nichols RS genome, so the gene name refers to the genome of the Chicago strain (accession no. CP001752.1). ‡Although base counts had an average counting coverage of 153× per tract across the 24 T. pallidum populations, particular base counts relying on a count coverage <20 are labelled and should be viewed with caution. §Based on the tprL extended annotation19.
Antigenic variation of TprK in in vivo human infection
Antigenic variation of the T. pallidum TprK antigen was known (in the rabbit model) to be mediated by a gene conversion mechanism specifically targeting seven variable (V) gene regions (V1–V7)13–15 that yields multiple distinct tprK sequences within single bacterial populations21,26,43, as a major adaptive mechanism for treponemal immune evasion and persistence. In this regard, we captured the sequences within each V region directly from clinical samples to evaluate intrastrain tprK variability and then ranked them according to their relative frequency within each population (Supplementary Table 4). Our in silico strategy proved to be more reliable and sensitive than classical Sanger sequencing, although Sanger results corroborated the existence of a sequence mixture and allowed the identification of the predominant sequences within each population (Supplementary Fig. 3). Sequence variability was found to be rampant both in content and length, with V3, V6 and V7 being particularly variable in length (Fig. 4a). In total, 230 distinct nucleotide sequences were captured, with a range of 13–54 distinct sequences (either nucleotide or amino acid) within each V region across all populations (Fig. 4b and Supplementary Fig. 4). Astonishingly, only one of the 230 sequences yields a tprK frameshift. Also, the captured nucleotide sequences rarely yielded the same peptide sequence within a single population; that is, for the same V region, synonymous nucleotide sequences were very infrequent within the same sample (Supplementary Table 4). In fact, we found a strong parallel scenario of sequence diversity within populations at both the nucleotide and amino acid levels (Fig. 5). All seven V regions showed variability within at least one-third of the 24 clinical populations, with a maximum obtained for the V7 region, for which we simultaneously detected up to nine amino acid sequences within the same population (Fig. 5) and variability within 16 of the 24 populations (Fig. 4c). Noteworthy, V6 region presented the highest number of distinct sequences across all populations, although sequences captured within this region were least prone to be shared between bacterial populations. In fact, within all other V regions, at least half of the populations carry sequences that are shared by other populations (Fig. 4c), pointing to a remarkable scenario of inter-population redundancy. In support of this scenario, sequences found to be redundant between populations were the ones that mostly dominate within single populations (regardless of the V region). Altogether, sequence analysis of the antigen-encoding tprK revealed a remarkable variability and redundancy within and between populations, respectively, which points to an ongoing adaptive diversification of TprK during infection where some specific sequences may be more advantageous in the context of T. pallidum interaction with the human host.
a, Schematic representation of the antigen-coding gene tprK showing the seven variable regions (V1–V7). The in-length variation range detected for each is also depicted above each V region. The V regions are specifically located between previously defined 4 bp conserved nucleotides strings15 (highlighted in blue). b, Scenario of TprK sequence diversity and redundancy within and between PT_SIF populations. The height of each bar proportionally represents the number of distinct amino acid sequences detected for all 24 populations within each V region, where distinct sequences are grouped according to their redundancy profile (that is, the number of populations sharing a particular sequence; see colour code). Numbers next to the bars refer to the number of sequences within each colour-coded group. c, The criteria and respective ratios enabling variability analysis within and across populations are presented below each bar. All detected nucleotide and amino acid sequences captured for each V region as well as their relative frequency (and ranking) within each population are presented in Supplementary Table 4.
Diversity of the antigenic repeats within the acidic repeat protein (Arp) of T. pallidum clinical populations
The T. pallidum potential virulence protein Arp possesses a central region composed of 20-amino-acid repeat motifs that were found to be immunogenic and variable in number both within the species and subspecies44,45. Our in silico capture of the repetitive motifs within each clinical population revealed the presence of all four motif types (I, II, III and II/III) previously described within the T. pallidum susbp. pallidum44,45. Curiously, all clinical populations displayed a very similar motif frequency hierarchy, although their frequency within each population was found to be different (Supplementary Fig. 5). Repeat type I was found to be the most frequent in all populations, followed by type II, III and II/III and, although inferences cannot be made regarding diversity within population for the reference strains, it is worth noting that type I is not the most common repeat for four of the five reference strains. Nevertheless, it must be stated that the presented frequencies may be a result of both within-chromosome variation in repeat number and clonal variation within the same population.
Discussion
The present study reports the full recovery and sequencing of multiple high-quality genomes of the syphilis-causing agent T. pallidum directly from clinical samples, thus bypassing the cultivation bottleneck that has hindered large-scale genomic analyses of this human pathogen. We demonstrate that this highly genomic conserved (>99.8%) bacterium takes advantage of the hypermutability of the antigen-coding tprK as well as of the in-length variation of poly(G/C) tracts mediating phase variation as major mechanisms for introducing rampant intrastrain genetic diversity within each patient. In fact, the TprK antigen revealed astonishing intrapatient variability targeting seven discrete hypervariable protein regions (Figs 4 and 5). TprK variability is known to be generated by a segmental gene conversion mechanism involving the unidirectional transfer of donor sequences into the seven tprK V regions13,15. Importantly, although the antigenicity character and surface exposure of TprK remains to be fully confirmed and is the subject of controversy27,28, it has been suggested that B-cell epitopes are located within the likely surface-exposed V regions, being primarily targeted by antibody responses following T. pallidum infection in rabbits24. This heterogeneity is believed to be responsible for both the lack of heterologous protection and the occurrence of reinfection in rabbits and humans24,25. Using the rabbit model, it was also found that antibodies recognize different epitopes at different times post-infection24 and that variability within V regions increases during active infection and passage15,22, with each V region accumulating diversity in an asynchronous fashion; that is, while V1 remained unchanged, the V6 region started diverging early after infection15. Our results corroborate this trend, as more distinct sequences were found for the V6 region (in contrast to the least variable, V1) (Figs 4 and 5), potentially indicating that hypermutability within V6 may be an initial trigger for TprK-mediated humoral immune evasion. Another noteworthy observation can be made for V7, which was found to be as heterogeneous as V6 (Figs 4 and 5), although it seems to evolve later on and to reach less sequence diversity than V6 during rabbit infection15. Another finding concerns the uncovering of TprK inter-population redundancy, where multiple sequences (usually the dominant alleles) were simultaneously found in distinct patients, pointing to parallel antigenic evolution in independent infection contexts. Overall, these observations raised the question of whether the TprK adaptive diversification pathway (that is, the timeline of V region evolution) is similar throughout infection in different hosts, and also whether host-specific immunological pressures ultimately determine the sequence outcome (that is, the most fitted TprK epitopes profile). Although our data may be consistent with the latter hypothesis, as comparisons between more than 150 distinct tprK V region sequences obtained from T. pallidum isolates cultured in rabbits15 and the 230 distinct sequences captured directly from clinical samples (our study) revealed a small overlap (45 sequences), parallel evolution experiments between humans and rabbits would be needed to validate it. Additionally, previous studies14,15,22,23 have supported the supposition that the ability of T. pallidum to escape immune clearance and become more invasive (reaching the cardiovascular and/or central nervous systems) may be seeded by a small and nearly clonal population that presumably harbours an advantageous ‘TprK-escaping variant’23. As such, our approach of capturing in vivo TprK diversity could ultimately lead to the confirmation of its critical role in T. pallidum virulence and tropism, while our results constitute per se a major database of TprK B-cell epitopes to be explored in future human immunological studies.
All clinical strains revealed intrastrain heterogeneity in at least eight poly(G/C) tracts (Supplementary Table 3). These reversible in-length genetic alterations are known to mediate quick phenotypic/adaptive changes through phase variation, either by modulating gene expression or by alternating the protein status (functional – on; truncated – off)46. Among the 31 poly(G/C) tracts revealing intrastrain allelic variation, more than half were consistently variable within most of the patient-derived strains (Fig. 3). Proteins whose expression may be potentially affected by phase variation include putative membrane proteins (most Tpr), several proteins with unknown function, a flagellar-associated protein and a chemotaxis-associated protein (Table 1). The variation of homopolymeric tracts has previously been demonstrated to impact the protein expression of the virulence-associated Tpr family16,17 and of a putative homologue of OmpW-family porins18. Our data constitute an important ‘population snapshot’ of ongoing phase variation within patients as well as the widest genome-scale mapping of potential targets of phase variation that are likely to be relevant to T. pallidum biology and syphilis pathogenesis. Further experimental validation is necessary at the gene and protein expression levels regarding whether these targets are regulated by phase variation. In particular, for the several proteins found to be in an ‘off’ state, especially for those where this state dominates within most patients (Table 1), it will be interesting to confirm their non-essentiality for T. pallidum in vivo growth and to investigate whether their activity is contingent on the infection context (for example, anatomical niches or differential hosts).
A detailed analysis of the phylogenetic branch enrolling all clinical strains revealed a very discrete SNP-based diversification, as they are separated by fewer than 20 SNPs. Additionally, this PT_SIF ‘clone’ (collected in Lisbon) was found to be highly related to the SS14 isolate (Fig. 1a), consistent with the expectation that SS14-like strains probably prevail in Europe6. Our data suggest a microevolutionary expansion of the circulating PT_SIF ‘clone’ from an SS14-like common ancestor (Fig. 1b). Comparison of the SS14 and PT_SIF genomic backbones revealed rather distinct mutational signatures. In contrast to the SS14 phylogenetic separation (the genome may have been affected during rabbit propagation), microevolution of the PT_SIF clinical strains suggests pathoadaptation as a result of human-derived selective pressures, as about eight of ten of the already fixed or potentially emerging SNPs are non-synonymous (Fig. 1b,c). A previous study that focused on comparing the genomes of two closely related strains also singled out that SNPs emerging during microevolution are essentially non-synonymous47. In the present study, the SNP-based microevolution was highly targeted, as some genes were affected more than once and very few functional categories were enrolled. As these protein categories are essentially the same as those involved in phase variation, this highlights a scenario of parallel evolution of T. pallidum populations in the context of the human–pathogen arms race. Furthermore, we described an intriguing mutational profile for a gene encoding a potential penicillin-binding protein (TPASS_20705/mrcA), whose putative involvement in a decrease in susceptibility to penicillin warrants investigation. Corroborating this, a different mutation in this gene (also designated pbp2) as well as other mutations in pbp genes have recently been reported in isolates from China48. Although the 23SrRNA A2058G resistance-associated mutation is commonly monitored11,49 and potentially results from macrolide administration to treat concomitant infections, for penicillin G (the current antibiotic of choice for syphilis treatment), evidence of drug resistance or of reduced susceptibility to penicillin is, to our knowledge, not known.
Our insights into the global genetic diversity within the T. pallidum pallidum (reviewed by Smajs and collegues11) supported the finding that the segregation of clades (Fig. 1a) relies on mutations essentially concentrated within a very limited gene set that enrols ∼70% of the ∼1,300 chromosome variant sites (excluding tprK and arp) (Fig. 2). The existence of the two main clades I and II (clade III is represented by a single genome) has been suggested by other genomic studies6,11,50, which have essentially distinguished the SS14-like (I) and Nichols-like (II) groups. However, it is worth noting that as all samples in this study were collected in a single town (Lisbon), we cannot discount the hypothesis that the studied T. pallidum clones are circulating in limited sexual transmission networks. As such, additional country-spread WGS studies will be needed to generalize our findings to the global picture of the molecular epidemiology of syphilis. In the current era of transition to WGS-based typing methodologies for epidemiological surveillance of infectious diseases, our results support that the in silico prediction of traditional T. pallidum subtypes is challenging, as one of the current typing genes (arp) carries multiple immunogenic repetitive motifs, the exact number of which is not predictable using the widely used short-read sequencing chemistry. However, our in silico strategy to retrieve the frequency hierarchy of arp motifs from short-read pools revealed that all clinical strains are particularly enriched by the immunogenic type I motif, which is believed to be exclusive to venereal T. pallidum subspecies45, in contrast to what is annotated in the reference genomes. Considering this trend and the fact that T. pallidum molecular evolution most probably relies on adaptive allelic variation targeting tprK and poly(G/C)s rather than on SNP fixation, a demanding design of WGS-based typing strategies will be required to guarantee an effective molecular surveillance of syphilis.
In summary, although this culture-independent targeted-WGS approach may contribute to the generation of relevant data on the worldwide geographic distribution of syphilitic strains, routes of transmission or the potential emergence of antibiotic-resistant strains, the results reported here have already unveiled that T. pallidum generates extensive within-patient subpopulation diversity, probably as a means to evade the host immune system and thus promote its survival, dissemination and persistence. We anticipate that the worldwide scale-up of our strategy for straightforwardly capturing T. pallidum in vivo genetic diversity will constitute a critical step towards unravelling genotype–phenotype associations, prioritizing candidates for vaccine development and ultimately decoding syphilis pathogenesis and dissimilar clinical outcomes.
Methods
T. pallidum clinical samples
Thirty-five T. pallidum positive DNA samples from the collection of the Reference Laboratory of Sexually Transmitted Infections at the Portuguese NIH were enrolled in the present study. These samples were obtained from clinical specimens of individuals attending the major Portuguese sexually transmitted disease clinic (Lapa Health Centre, Lisbon, Portugal) (Supplementary Table 1). To potentiate the success of the culture-independent targeted WGS approach, DNA samples were chosen according to the T. pallidum load (defined by real-time qPCR). Some samples with low T. pallidum copy number were also included to evaluate the success threshold of the applied strategy. DNA extraction was performed using the QIAamp DNA Mini Kit (Qiagen) according to the manufacturer's instructions. Each sample was characterized by quantifying both the number of T. pallidum (targeting the single-copy tprB) and human genome copies (targeting β-actin) through qPCR using LightCycler 480 SYBR Green chemistry and optical plates (Roche Diagnostics). Absolute quantification was possible by using, as standard curves, cloned plasmids (for both tprB and β-actin) generated through the TOPO TA technology (Invitrogen) and transformation of DH5α Escherichia coli cells, as described for other pathogens51.
SureSelectXTT. pallidum enrichment and WGS directly from clinical samples
RNA oligonucleotide baits (120 bp in size; total of 19,094) were designed to span the ∼1.1 Mb of the T. pallidum genome. To ensure sensitivity and specificity, bait design accounted for genetic variability among the six publicly available T. pallidum genomes, and baits with considerable homology to the human genome (after BLASTn search against the Human Genomic + Transcript database) were excluded. This custom bait library was then uploaded to the SureDesign software and synthesized by Agilent Technologies. Before enrichment and WGS, DNA samples were quantified using Qubit HS kit (Invitrogen, Life Technologies) to calibrate input to 200 ng DNA. Genomic DNA samples were sheared at 4 °C on a Bioruptor Next Gen (diagenode) sonicator using 35 cycles of 30 s each to fragment DNA to ∼300–400 bp. The enrichment protocol was performed according to the Agilent Technologies' SureSelectXT Target Enrichment System for Illumina Paired-End Sequencing Library protocol (version B.1, December 2014), with two modifications. Briefly, the hybridization and capture steps were performed twice, consecutively, and the number of cycles of the post-capture PCR was increased from 14 to 23 to maximize the yield of target DNA. In fact, preliminary testing (Supplementary Fig. 6) revealed that the two-step approach yielded at least a twofold increase in the subsequently obtained T. pallidum specific (on-target) reads. This strategy was applied to all enrolled clinical samples except one (PT_SIF1127), which was instead subjected to immunomagnetic separation (IMS) followed by multiple displacement amplification (MDA) before WGS. This consisted of a previous attempt at sequencing T. pallidum directly from clinical samples, which proved to be unsuccessful, as an extremely uneven coverage was generated across the chromosome. Nonetheless, after dedicating more than 14 million reads, we were still able to obtain a nearly-complete genome sequence (>99%) for this single IMS-MDA-processed sample (PT_SIF1127). This genome was included in this study exclusively for SNP-based comparative genomics.
T. pallidum enriched libraries were subjected to cluster generation and paired-end sequencing (2 × 250 bp) in Illumina MiSeq equipment (Illumina), according to the manufacturer's instructions. FastQC (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/) and FASTX (http://hannonlab.cshl.edu/fastx_toolkit/) tools were applied to check and improve the quality of the raw sequence data, respectively. Reads were mapped against the T. pallidum SS14 re-sequenced (RS) chromosome sequence (CP004011.1) available at GenBank6,7 using Bowtie2 (ref. 52) (version 2.1.0). Median depth of coverage (Supplementary Table 1) was highly homogeneous across the chromosome, with no bias towards particular regions or genes. SAMtools/BCFtools53 were applied to call SNPs, and insertions/deletions (indels) and inter- and intrastrain variant nucleotide sites were carefully confirmed by visual inspection using the Integrative Genomics Viewer54 (version 2.3.59). Although the reference-based approach seemed highly accurate for this study (T. pallidum pan- and core-genome overlap), de novo assembly of T. pallidum genomes, after subtraction of reads mapping to the human genome, was also performed for confirmation purposes using Velvet version 1.2.10 (ref. 55) optimized with VelvetOptimiser script. Given the strict clonality detected among the sequenced genomes, final genome sequences were assembled and closed by replacing variant bases in the reference genome sequence, where, in cases of allelic mixtures (affecting single nucleotide sites, variable homopolymeric tracts and tprK gene; see section ‘Evaluation of intrastrain genetic heterogeneity'), the highest frequent allele was annotated. For the arp gene, which has multiple 60 bp tandem repeats34, although the exact number cannot be predicted using the currently available Illumina short-read-based technology, the unique sequences of the most abundant 60 bp repeats within T. pallidum populations could be extracted and thus were annotated. The additional non-annotated 60 bp repeats, as well as chromosome regions not covered (only for three samples, representing less than 0.06% of the chromosome length), were reported as undefined bases. The closed genome sequences (designated PT_SIF plus a specific number code) had a mean assembled genome size of 1,139,136 bp (Supplementary Table 1).
Whole-genome-based comparative genomics
Alignments of the closed genome sequences were performed using the progressive algorithm of Mauve software56 (version 2.3.1). All T. pallidum pallidum genomes available at GenBank (at the time the study was conducted) were included in these analyses (strains SS14 RS (ref. 6; accession no. CP004011.1), Nichols RS (ref. 6; accession no. CP004010.2), Mexico A (ref. 5; accession no. CP003064.1), DAL-1 (ref. 10; accession no. CP003115.1), Chicago (ref. 8; accession no. CP001752.1) and Sea 81-4 (ref. 9; accession no. CP003679.1). Identification of regions with high SNP density across the T. pallidum genome was performed through DnaSP v5 analysis57 using a window size and a step size of 1,000 base pairs each. MEGA 5 software58 was applied to determine the overall mean distances and matrices of pairwise comparisons at the nucleotide level and to scrutinize the mutational dynamics underlying the T. pallidum microevolution. Whole-genome-based phylogenetic trees were inferred by using the Neighbor-Joining method59,60 (bootstrap = 1,000). For the nucleotide sequences, the evolutionary distances were computed using the Kimura 2-parameter method61.
Evaluation of intrastrain genetic heterogeneity
To disclose the within-patient genetic diversity of T. pallidum, a detailed search was performed for genomic regions displaying intrastrain heterogeneity, through the inspection of single nucleotide sites with allelic variation, in-length variable DNA homopolymeric tracts (poly-G and poly-C) and tprK gene variable (V) regions (V1–V7)14,15.
Single nucleotide sites were validated as intrastrain heterogeneous when the following conditions were simultaneously verified: (1) they were supported by at least 50 reads; (2) the less frequent base displayed a frequency above 10%; and (3) the less frequent base was supported by at least eight unique reads. Regarding variation within DNA homopolymeric tracts and tprK gene regions, an in-house Python script was developed and applied to extract and count (directly from raw reads, both forward and reverse) DNA sequences that are contiguously flanked by two conserved (among all reference and our clinical strains), user-defined, small DNA strings. Thus, for a given variable region (in size and/or in content), the script retrieves the exact number of base counts and sequences, allowing determination of the precise relative frequency of clones carrying specific base counts or distinct sequence within an intrapopulation variable region. For poly(G/C) tracts analysis, the following approach was applied for variability/conservation validation, thus improving the analysis quality: (1) homopolymeric tracts were considered ‘conserved’ if the dominant ‘count’ represents more than 90% of all respective reads counted in that region; (2) nucleotide strings containing bases other than the expected bases, for a given homopolymeric tract, were excluded from the final count (although it must be noted that they represent a mean of 1 ± 0.4% of the total counts); (3) strand bias was not accounted for, as we were counting nucleotide strings (that is, the homopolymeric tract plus flanking regions) rather than single nucleotide positions; and (4) specifically for poly(G/C) tracts falling within promoter regions of the tpr paralogues (which are essentially conserved across tpr subfamilies), the flanking regions are not immediately contiguous to the respective tracts (although this impacted the coverage, the counting confidence is enhanced as larger sequences are considered). Following this approach, all results were validated, regardless of the ‘counting coverage’ (an average of 153-fold per tract was obtained across populations), although results with low coverage (<20) were labelled throughout the text as they should be viewed with caution. The criteria for tprK intrastrain variability analysis were as follows: (1) only variants with at least ten reads counted were validated, which means, for instance, that we excluded variants below 5% when the ‘counting coverage’ was 200 (because the probability of the occurrence of a single base error increases proportionally with increasing sequence length, this conservative criterion minimizes the likelihood that reads with random single base errors are assumed to be variant alleles); (2) exceptionally, variants supported by fewer than ten reads were validated if they represented 25% of all reads counted (although this only happened for 6 of the 370 total sequences captured); and (3) strand bias was not accounted for, because we were counting nucleotide strings (that is, the V region plus flanking regions) rather than single nucleotide positions. Counting quality was also reinforced when using the script by the fact that both flanking regions are conserved among all reported counts/sequences. Finally, the in-house Python script was also applied to determine the relative frequency of the four known 60 bp repetitive motifs (types I, II, III and II/III) of the T. pallidum typing gene arp within each clinical sample.
Ensuring that the observed intrapopulation T. pallidum heterogeneity matched the one present in the clinical specimens, we obtained similar results for the relative frequency of the clones within the population after performing independent SureSelect procedures on the same sample. Thus, the population profiles (that is, the proportion of clones within the population), which were also supported by Sanger sequencing of the tprK V2 region before the enrichment process, were probably not affected by our culture-independent targeted-WGS strategy.
Data availability
A NCBI Bioproject was created to group all reads and assemblies associated with the genomes sequenced in this study and is available using accession code PRJNA322283. Closed genome sequences have been deposited in GenBank and annotated by the NCBI Prokaryotic Genomes Annotation Pipeline 2.3. Raw sequence data (after exclusion of reads matching the human genome, for ethical purposes) were uploaded to the Short Read Archive (SRA). All accession codes are listed in Supplementary Table 1. Data sets generated during this study are included in the Supplementary Information. Code for the in-house Python script is available at https://github.com/monsanto-pinheiro/countDNABox.
References
LaFond, R. E. & Lukehart, S. A. Biological basis for syphilis. Clin. Microbiol. Rev. 19, 29–49 (2006).
Ho, E. L. & Lukehart, S. A. Syphilis: using modern approaches to understand an old disease. J. Clin. Invest. 121, 4584–4592 (2011).
Cullen, P. A. & Cameron, C. E. Progress towards an effective syphilis vaccine: the past, present and future. Exp. Rev. Vaccines 5, 67–80 (2006).
Fraser, C. M. et al. Complete genome sequence of Treponema pallidum, the syphilis spirochete. Science 281, 375–388 (1998).
Pětrošová, H. et al. Whole genome sequence of Treponema pallidum ssp. pallidum, strain Mexico A, suggests recombination between yaws and syphilis strains. PLoS Negl. Trop. Dis. 6, e1832 (2012).
Pětrošová, H. et al. Resequencing of Treponema pallidum ssp. pallidum strains Nichols and SS14: correction of sequencing errors resulted in increased separation of syphilis treponeme subclusters. PLoS ONE 8, e74319 (2013).
Matejková, P. et al. Complete genome sequence of Treponema pallidum ssp. pallidum strain SS14 determined with oligonucleotide arrays. BMC Microbiol. 8, 76 (2008).
Giacani, L. et al. Complete genome sequence and annotation of the Treponema pallidum subsp. pallidum Chicago strain. J. Bacteriol. 192, 2645–2646 (2010).
Giacani, L. et al. Complete genome sequence of the Treponema pallidum subsp. pallidum Sea81-4 strain. Genome Announce. 2, e00333 (2014).
Zobaníková, M. et al. Complete genome sequence of Treponema pallidum strain DAL-1. Stand Genomic Sci. 7, 12–21 (2012).
Smajs, D., Norris, S. J. & Weinstock, G. M. Genetic diversity in Treponema pallidum: implications for pathogenesis, evolution and molecular diagnostics of syphilis and yaws. Infect Genet. Evol. 12, 191–202 (2012).
Čejková, D., Strouhal, M., Norris, S. J., Weinstock, G. M. & Šmajs, D. A retrospective study on genetic heterogeneity within Treponema strains subpopulations are genetically distinct in a limited number of positions. PLoS Negl. Trop. Dis. 9, e0004110 (2015).
Giacani, L. et al. Comparative investigation of the genomic regions involved in antigenic variation of the TprK antigen among treponemal species, subspecies, and strains. J. Bacteriol. 194, 4208–4225 (2012).
Centurion-Lara, A., Godornes, C., Castro, C., Van Voorhis, W. C. & Lukehart, S. A. The tprK gene is heterogeneous among Treponema pallidum strains and has multiple alleles. Infect. Immun. 68, 824–831 (2000).
Centurion-Lara, A. et al. Gene conversion: a mechanism for generation of heterogeneity in the tprK gene of Treponema pallidum during infection. Mol. Microbiol. 52, 1579–1596 (2004).
Giacani, L., Hevner, K. & Centurion-Lara, A. Gene organization and transcriptional analysis of the tprJ, tprI, tprG, and tprF loci in Treponema pallidum strains Nichols and Sea 81-4. J. Bacteriol. 187, 6084–6093 (2005).
Giacani, L., Lukehart, S. & Centurion-Lara, A. Length of guanosine homopolymeric repeats modulates promoter activity of subfamily II tpr genes of Treponema pallidum ssp. pallidum. FEMS Immunol. Med. Microbiol. 51, 289–301 (2007).
Giacani, L. et al. Transcription of TP0126, Treponema pallidum putative OmpW homolog, is regulated by the length of a homopolymeric guanosine repeat. Infect. Immun. 83, 2275–2289 (2015).
Centurion-Lara, A. et al. Fine analysis of genetic diversity of the tpr gene family among treponemal species, subspecies and strains. PLoS Negl. Trop. Dis. 7, e2222 (2013).
Gray, R. R. et al. Molecular evolution of the tprC, D, I, K, G, and J genes in the pathogenic genus Treponema. Mol. Biol. Evol. 23, 2220–2233 (2006).
LaFond, R. E. et al. Sequence diversity of Treponema pallidum subsp. pallidum tprK in human syphilis lesions and rabbit-propagated isolates. J. Bacteriol. 185, 6262–6268 (2003).
LaFond, R. E., Centurion-Lara, A., Godornes, C., Van Voorhis, W. C. & Lukehart, S. A. TprK sequence diversity accumulates during infection of rabbits with Treponema pallidum subsp. pallidum Nichols strain. Infect. Immun. 74, 1896–1906 (2006).
Reid, T. B., Molini, B. J., Fernandez, M. C. & Lukehart, S. A. Antigenic variation of TprK facilitates development of secondary syphilis. Infect. Immun. 82, 4959–4967 (2014).
Morgan, C. A., Molini, B. J., Lukehart, S. A. & Van Voorhis, W. C. Segregation of B and T cell epitopes of Treponema pallidum repeat protein K to variable and conserved regions during experimental syphilis infection. J. Immunol. 169, 952–957 (2002).
Morgan, C. A., Lukehart, S. A. & Van Voorhis, W. C. Protection against syphilis correlates with specificity of antibodies to the variable regions of Treponema pallidum repeat protein K. Infect. Immun. 71, 5605–5612 (2003).
Stamm, L. V. & Bergen, H. L. The sequence-variable, single-copy tprK gene of Treponema pallidum Nichols strain UNC and Street strain 14 encodes heterogeneous TprK proteins. Infect. Immun. 68, 6482–6486 (2000).
Hazlett, K. R. et al. The TprK protein of Treponema pallidum is periplasmic and is not a target of opsonic antibody or protective immunity. J. Exp. Med. 193, 1015–1026 (2001).
Cox, D. L. et al. Surface immunolabeling and consensus computational framework to identify candidate rare outer membrane proteins of Treponema pallidum. Infect. Immun. 78, 5178–5194 (2010).
Anand, A. et al. The major outer sheath protein (Msp) of Treponema denticola has a bipartite domain architecture and exists as periplasmic and outer membrane-spanning conformers. J. Bacteriol. 195, 2060–2071 (2013).
Gnirke, A. et al. Solution hybrid selection with ultra-long oligonucleotides for massively parallel targeted sequencing. Nat. Biotechnol. 27, 182–189 (2009).
Geniez, S. et al. Targeted genome enrichment for efficient purification of endosymbiont DNA from host DNA. Symbiosis 58, 201–207 (2012).
Brown, A. C. et al. Rapid whole-genome sequencing of Mycobacterium tuberculosis isolates directly from clinical samples. J. Clin. Microbiol. 53, 2230–2237 (2015).
Christiansen, M. T. et al. Whole-genome enrichment and sequencing of Chlamydia trachomatis directly from clinical samples. BMC Infect. Dis. 14, 591 (2014).
Pillay, A. et al. Molecular subtyping of Treponema pallidum subspecies pallidum. Sex. Transm. Dis. 25, 408–414 (1998).
Centurion-Lara, A. et al. Multiple alleles of Treponema pallidum repeat gene D in Treponema pallidum isolates. J. Bacteriol. 182, 2332–2335 (2000).
Stamm, L. V. & Bergen, H. L. A point mutation associated with bacterial macrolide resistance is present in both 23S rRNA genes of an erythromycin-resistant Treponema pallidum clinical isolate. Antimicrob. Agents Chemother. 44, 806–807 (2000).
Naqvi, A. A., Shahbaaz, M., Ahmad, F. & Hassan, M. I. Identification of functional candidates amongst hypothetical proteins of Treponema pallidum ssp. pallidum. PLoS ONE 10, e0124177 (2015).
Pares, S., Mouz, N., Pétillot, Y., Hakenbeck, R. & Dideberg, O. X-ray structure of Streptococcus pneumoniae PBP2x, a primary penicillin target enzyme. Nat. Struct. Biol. 3, 284–289 (1996).
Jerome, J. P. et al. Standing genetic variation in contingency loci drives the rapid adaptation of Campylobacter jejuni to a novel host. PLoS ONE 6, e16399 (2011).
Dohm, J. C., Lottaz, C., Borodina, T. & Himmelbauer, H. Substantial biases in ultra-short read data sets from high-throughput DNA sequencing. Nucleic Acids Res. 36, e105 (2008).
Loman, N. J. et al. Performance comparison of benchtop high-throughput sequencing platforms. Nat. Biotechnol. 30, 434–439 (2012).
Quail, M. A. et al. A tale of three next generation sequencing platforms: comparison of Ion Torrent, Pacific Biosciences and Illumina MiSeq sequencers. BMC Genomics 13, 341 (2012).
Heymans, R., Kolader, M. E., van der Helm, J. J., Coutinho, R. A. & Bruisten, S. M. TprK gene regions are not suitable for epidemiological syphilis typing. Eur. J. Clin. Microbiol. Infect. Dis. 28, 875–878 (2009).
Liu, H., Rodes, B., George, R. & Steiner, B. Molecular characterization and analysis of a gene encoding the acidic repeat protein (Arp) of Treponema pallidum. J. Med. Microbiol. 56, 715–721 (2007).
Harper, K. N. et al. The sequence of the acidic repeat protein (arp) gene differentiates venereal from nonvenereal Treponema pallidum subspecies, and the gene has evolved under strong positive selection in the subspecies that causes syphilis. FEMS Immunol. Med. Microbiol. 53, 322–332 (2008).
Moxon, R., Bayliss, C. & Hood, D. Bacterial contingency loci: the role of simple sequence DNA repeats in bacterial adaptation. Annu. Rev. Genet. 40, 307–333 (2006).
Giacani, L. et al. Footprint of positive selection in Treponema pallidum subsp. pallidum genome sequences suggests adaptive microevolution of the syphilis pathogen. PLoS Negl. Trop. Dis. 6, e1698 (2012).
Sun, J. et al. Tracing the origin of Treponema pallidum in China using next-generation sequencing. Oncotarget http://dx.doi.org/10.18632/oncotarget.10154 (17 June 2016).
Mitchell, S. J. et al. Azithromycin-resistant syphilis infection: San Francisco, California, 2000–2004. Clin. Infect. Dis. 42, 337–345 (2006).
Nechvátal, L. et al. Syphilis-causing strains belong to separate SS14-like or Nichols-like groups as defined by multilocus analysis of 19 Treponema pallidum strains. Int. J. Med. Microbiol. 304, 645–653 (2014).
Gomes, J. P. et al. Correlating Chlamydia trachomatis infectious load with urogenital ecological success and disease pathogenesis. Microbes Infect. 8, 16–26 (2006).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Zerbino, D. R. & Birney, E. Velvet: Algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008).
Darling, A. E., Mau, B. & Perna, N. T. progressiveMauve: multiple genome alignment with gene gain, loss and rearrangement. PLoS ONE 5, e11147 (2010).
Librado, P. & Rozas, J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452 (2009).
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011).
Felsenstein, J. Confidence-limits on phylogenies—an approach using the bootstrap. Evolution 39, 783–791 (1985).
Saitou, N. & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987).
Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 16, 111–120 (1980).
Acknowledgements
This study was partially supported by grant EXPL/BIA-MIC/0309/2013 from the Fundação para a Ciência e a Tecnologia (FCT).
Author information
Authors and Affiliations
Contributions
J.P.G. conceived the study. M.Pint., V.B. and J.P.G. designed the study. M.Pint., M.A., J.M., D.C. and L.V. performed the wet lab experiments. M.Pint. and V.B. performed bioinformatics and comparative genomics analyses. M.Pint., V.B., M.A., A.N., M.J.B., J.M., D.C., L.V. and J.P.G. performed research and analysed data. J.A. performed sample collection and clinical diagnostics. M.Pinh. provided bioinformatics support. M.Pint., V.B. and J.P.G. wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary Figures 1-6, Supplementary Tables 1 and 2, legends for Supplementary Tables 3 and 4, Supplementary References (PDF 1057 kb)
Supplementary Table 3
Poly(G/C) tracts in T. pallidum pallidum genomes analysed for the presence of in-length genetic heterogeneity within clinical samples (XLSX 79 kb)
Supplementary Table 4
Sequence diversity within the seven variable regions (V1-V7) of tprK captured directly from 24 T. pallidum pallidum clinical samples. (XLSX 38 kb)
Rights and permissions
About this article
Cite this article
Pinto, M., Borges, V., Antelo, M. et al. Genome-scale analysis of the non-cultivable Treponema pallidum reveals extensive within-patient genetic variation. Nat Microbiol 2, 16190 (2017). https://doi.org/10.1038/nmicrobiol.2016.190
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2016.190
This article is cited by
-
Redefining the treponemal history through pre-Columbian genomes from Brazil
Nature (2024)
-
Core genome sequencing and genotyping of Leptospira interrogans in clinical samples by target capture sequencing
BMC Infectious Diseases (2023)
-
Investigation of the immune escape mechanism of Treponema pallidum
Infection (2023)
-
A suite of PCR-LwCas13a assays for detection and genotyping of Treponema pallidum in clinical samples
Nature Communications (2022)
-
Phylogenetic and genetic characterization of Treponema pallidum strains from syphilis patients in Japan by whole-genome sequence analysis from global perspectives
Scientific Reports (2021)