Abstract
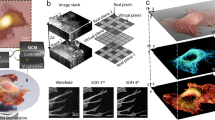
Microbial cells have developed sophisticated multicomponent structures and machineries to govern basic cellular processes, such as chromosome segregation, gene expression, cell division, mechanosensing, cell adhesion and biofilm formation. Because of the small cell sizes, subcellular structures have long been difficult to visualize using diffraction-limited light microscopy. During the last three decades, optical and force nanoscopy techniques have been developed to probe intracellular and extracellular structures with unprecedented resolutions, enabling researchers to study their organization, dynamics and interactions in individual cells, at the single-molecule level, from the inside out, and all the way up to cell–cell interactions in microbial communities. In this Review, we discuss the principles, advantages and limitations of the main optical and force nanoscopy techniques available in microbiology, and we highlight some outstanding questions that these new tools may help to answer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Ghosal, D. & Lowe, J. Collaborative protein filaments. Embo J. 34, 2312–2320 (2015).
Cloud-Hansen, K. A. et al. Breaching the great wall: peptidoglycan and microbial interactions. Nat. Rev. Microbiol. 4, 710–716 (2006).
Kolter, R. & Greenberg, E. P. Microbial sciences: The superficial life of microbes. Nature 441, 300–302 (2006).
Müller, D. J. & Dufrêne, Y. F. Atomic force microscopy as a multifunctional molecular toolbox in nanobiotechnology. Nat. Nanotech. 3, 261–269 (2008).
Dufrêne, Y. F. Towards nanomicrobiology using atomic force microscopy. Nat. Rev. Microbiol. 6, 674–680 (2008).
Engel, A. & Muller, D. J. Observing single biomolecules at work with the atomic force microscope. Nat. Struct. Biol. 7, 715–718 (2000).
Engel, A. & Gaub, H. E. Structure and Mechanics of Membrane Proteins. Annu. Rev. Biochem. 77, 127–148 (2008).
Ando, T., Uchihashi, T. & Scheuring, S. Filming Biomolecular Processes by High-Speed Atomic Force Microscopy. Chem. Rev. 114, 3120–3188 (2014).
Hinterdorfer, P. & Dufrêne, Y. F. Detection and localization of single molecular recognition events using atomic force microscopy. Nat. Methods 3, 347–355 (2006).
Müller, D. J., Helenius, J., Alsteens, D. & Dufrêne, Y. F. Force probing surfaces of living cells to molecular resolution. Nat. Chem. Biol. 5, 383–390 (2009).
Strunz, T., Oroszlan, K., Schafer, R. & Guntherodt, H. J. Dynamic force spectroscopy of single DNA molecules. Proc. Natl Acad. Sci. USA 96, 11277–11282 (1999).
Dupres, V. et al. Nanoscale mapping and functional analysis of individual adhesins on living bacteria. Nat. Methods 2, 515–520 (2005).
Alsteens, D., Garcia, M. C., Lipke, P. N. & Dufrene, Y. F. Force-induced formation and propagation of adhesion nanodomains in living fungal cells. Proc. Natl Acad. Sci. 107, 20744–20749 (2010).
Dupres, V. et al. The yeast Wsc1 cell surface sensor behaves like a nanospring in vivo. Nat. Chem. Biol. 5, 857–862 (2009).
Helenius, J., Heisenberg, C. P., Gaub, H. E. & Muller, D. J. Single-cell force spectroscopy. J. Cell Sci. 121, 1785–1791 (2008).
Beaussart, A. et al. Single-cell force spectroscopy of probiotic bacteria. Biophys. J. 104, 1886–1892 (2013).
Beaussart, A. et al. Nanoscale adhesion forces of Pseudomonas aeruginosa type IV pili. ACS Nano 8, 10723–10733 (2014).
Dufrêne, Y. F., Martínez-Martín, D., Medalsy, I., Alsteens, D. & Müller, D. J. Multiparametric imaging of biological systems by force-distance curve–based AFM. Nat. Methods 10, 847–854 (2013).
Medalsy, I., Hensen, U. & Muller, D. J. Imaging and quantifying chemical and physical properties of native proteins at molecular resolution by force-volume AFM. Angew. Chem. Int. Ed. 50, 12103–12108 (2011).
Rico, F., Su, C. & Scheuring, S. Mechanical mapping of single membrane proteins at submolecular resolution. Nano Lett. 11, 3983–3986 (2011).
Alsteens, D., Trabelsi, H., Soumillion, P. & Dufrêne, Y. F. Multiparametric atomic force microscopy imaging of single bacteriophages extruding from living bacteria. Nat. Commun. 4, 2926 (2013).
Formosa, C. et al. Multiparametric imaging of adhesive nanodomains at the surface of Candida albicans by atomic force microscopy. Nanomed. Nanotechnol. Biol. Med. 11, 57–65 (2015).
Formosa-Dague, C., Speziale, P., Foster, T. J., Geoghegan, J. A. & Dufrêne, Y. F. Zinc-dependent mechanical properties of Staphylococcus aureus biofilm-forming surface protein SasG. Proc. Natl Acad. Sci. 113, 410–415 (2015).
Formosa-Dague, C. et al. Sticky matrix: adhesion mechanism of the Staphylococcal polysaccharide intercellular adhesin. ACS Nano 10, 3443–3452 (2016).
Hell, S. W. & Wichmann, J. Breaking the diffraction resolution limit by stimulated emission: stimulated-emission-depletion fluorescence microscopy. Opt. Lett. 19, 780–782 (1994).
Klar, T. A., Jakobs, S., Dyba, M., Egner, A. & Hell, S. W. Fluorescence microscopy with diffraction resolution barrier broken by stimulated emission. Proc. Natl Acad. Sci. USA 97, 8206–8210 (2000).
Schneider, J. et al. Ultrafast, temporally stochastic STED nanoscopy of millisecond dynamics. Nat. Methods 12, 827–830 (2015).
Schmidt, R. et al. Mitochondrial cristae revealed with focused light. Nano Lett. 9, 2508–2510 (2009).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Hess, S. T., Girirajan, T. P. & Mason, M. D. Ultra-high resolution imaging by fluorescence photoactivation localization microscopy. Biophys. J. 91, 4258–4272 (2006).
Rust, M. J., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–795 (2006).
Heilemann, M. et al. Subdiffraction-resolution fluorescence imaging with conventional fluorescent probes. Angew. Chem. Int. Ed. Engl. 47, 6172–6176 (2008).
van Oijen, A. M., Köhler, J., Schmidt, J., Müller, M. & Brakenhoff, G. J. 3-Dimensional super-resolution by spectrally selective imaging. Chem. Phys. Lett. 292, 183–187 (1998).
Thompson, R. E., Larson, D. R. & Webb, W. W. Precise nanometer localization analysis for individual fluorescent probes. Biophys. J. 82, 2775–2783 (2002).
Chozinski, T. J., Gagnon, L. A. & Vaughan, J. C. Twinkle, twinkle little star: photoswitchable fluorophores for super-resolution imaging. FEBS Lett. 588, 3603–3612 (2014).
Pavani, S. R. et al. Three-dimensional, single-molecule fluorescence imaging beyond the diffraction limit by using a double-helix point spread function. Proc. Natl Acad. Sci. USA 106, 2995–2999 (2009).
Juette, M. F. et al. Three-dimensional sub-100 nm resolution fluorescence microscopy of thick samples. Nat. Methods 5, 527–529 (2008).
Tang, J., Akerboom, J., Vaziri, A., Looger, L. L. & Shank, C. V. Near-isotropic 3D optical nanoscopy with photon-limited chromophores. Proc. Natl Acad. Sci. USA 107, 10068–10073 (2010).
Huang, B., Wang, W., Bates, M. & Zhuang, X. Three-dimensional super-resolution imaging by stochastic optical reconstruction microscopy. Science 319, 810–813 (2008).
Shtengel, G. et al. Interferometric fluorescent super-resolution microscopy resolves 3D cellular ultrastructure. Proc. Natl Acad. Sci. USA 106, 3125–3130 (2009).
Shroff, H., Galbraith, C. G., Galbraith, J. A. & Betzig, E. Live-cell photoactivated localization microscopy of nanoscale adhesion dynamics. Nat. Methods 5, 417–423 (2008).
Coltharp, C., Kessler, R. P. & Xiao, J. Accurate construction of photoactivated localization microscopy (PALM) images for quantitative measurements. PLoS ONE 7, e51725 (2012).
Endesfelder, U., Malkusch, S., Fricke, F. & Heilemann, M. A simple method to estimate the average localization precision of a single-molecule localization microscopy experiment. Histochem. Cell Biol. 141, 629–638 (2014).
Huang, F. et al. Video-rate nanoscopy using sCMOS camera-specific single-molecule localization algorithms. Nat. Methods 10, 653–658 (2013).
Gustafsson, M. G. Surpassing the lateral resolution limit by a factor of two using structured illumination microscopy. J. Microsc. 198, 82–87 (2000).
Schermelleh, L. et al. Subdiffraction multicolor imaging of the nuclear periphery with 3D structured illumination microscopy. Science 320, 1332–1336 (2008).
Gustafsson, M. G. Nonlinear structured-illumination microscopy: wide-field fluorescence imaging with theoretically unlimited resolution. Proc. Natl Acad. Sci. USA 102, 13081–13086 (2005).
Li, D. et al. ADVANCED IMAGING. Extended-resolution structured illumination imaging of endocytic and cytoskeletal dynamics. Science 349, aab3500 (2015).
Hell, S. W. Microscopy and its focal switch. Nat. Methods 6, 24–32 (2009).
Hell, S. W. & Kroug, M. Ground-state-depletion fluorscence microscopy: A concept for breaking the diffraction resolution limit. Appl. Phys. B 60, 495–497 (1995).
Hofmann, M., Eggeling, C., Jakobs, S. & Hell, S. W. Breaking the diffraction barrier in fluorescence microscopy at low light intensities by using reversibly photoswitchable proteins. Proc. Natl Acad. Sci. USA 102, 17565–17569 (2005).
Sharonov, A. & Hochstrasser, R. M. Wide-field subdiffraction imaging by accumulated binding of diffusing probes. Proc. Natl Acad. Sci. USA 103, 18911–18916 (2006).
Jungmann, R. et al. Single-molecule kinetics and super-resolution microscopy by fluorescence imaging of transient binding on DNA origami. Nano Lett. 10, 4756–4761 (2010).
Schoen, I., Ries, J., Klotzsch, E., Ewers, H. & Vogel, V. Binding-activated localization microscopy of DNA structures. Nano Lett. 11, 4008–4011 (2011).
Huang, B., Babcock, H. & Zhuang, X. Breaking the diffraction barrier: super-resolution imaging of cells. Cell 143, 1047–1058 (2010).
Dempsey, G. T., Vaughan, J. C., Chen, K. H., Bates, M. & Zhuang, X. Evaluation of fluorophores for optimal performance in localization-based super-resolution imaging. Nat. Methods 8, 1027–1036 (2011).
Durisic, N., Laparra-Cuervo, L., Sandoval-Alvarez, A., Borbely, J. S. & Lakadamyali, M. Single-molecule evaluation of fluorescent protein photoactivation efficiency using an in vivo nanotemplate. Nat. Methods 11, 156–162 (2014).
Gahlmann, A. & Moerner, W. E. Exploring bacterial cell biology with single-molecule tracking and super-resolution imaging. Nat. Rev. Microbiol. 12, 9–22 (2014).
Coltharp, C. & Xiao, J. Superresolution microscopy for microbiology. Cell Microbiol. 14, 1808–1818 (2012).
Coltharp, C., Yang, X. & Xiao, J. Quantitative analysis of single-molecule superresolution images. Curr. Opin. Struct. Biol. 28, 112–121 (2014).
Small, A. R. & Parthasarathy, R. Superresolution localization methods. Annu. Rev. Phys. Chem. 65, 107–125 (2014).
Dame, R. T. & Tark-Dame, M. Bacterial chromatin: converging views at different scales. Curr. Opin. Cell Biol. 40, 60–65 (2016).
Spahn, C., Endesfelder, U. & Heilemann, M. Super-resolution imaging of Escherichia coli nucleoids reveals highly structured and asymmetric segregation during fast growth. J. Struct. Biol. 185, 243–249 (2014).
Spahn, C., Cella-Zannacchi, F., Endesfelder, U. & Heilemann, M. Correlative super-resolution imaging of RNA polymerase distribution and dynamics, bacterial membrane and chromosomal structure in Escherichia coli. Methods Appl. Fluoresc. 3, 014005 (2015).
Foo, Y. H., Spahn, C., Zhang, H., Heilemann, M. & Kenney, L. J. Single cell super-resolution imaging of E. coli OmpR during environmental stress. Integr. Biol. 7, 1297–1308 (2015).
Kolb, H. C., Finn, M. G. & Sharpless, K. B. Click chemistry: diverse chemical function from a few good reactions. Angew. Chem. Int. Ed. Engl. 40, 2004–2021 (2001).
Marbouty, M. et al. Condensin- and replication-mediated bacterial chromosome folding and origin condensation revealed by Hi-C and super-resolution imaging. Mol. Cell 59, 588–602 (2015).
Stracy, M. et al. Live-cell superresolution microscopy reveals the organization of RNA polymerase in the bacterial nucleoid. Proc. Natl Acad. Sci. USA 112, E4390–E4399 (2015).
Lee, S. F., Thompson, M. A., Schwartz, M. A., Shapiro, L. & Moerner, W. E. Super-resolution imaging of the nucleoid-associated protein HU in Caulobacter crescentus. Biophys. J. 100, L31–33 (2011).
Wang, W., Li, G. W., Chen, C., Xie, X. S. & Zhuang, X. Chromosome organization by a nucleoid-associated protein in live bacteria. Science 333, 1445–1449 (2011).
Straight, A. F., Belmont, A. S., Robinett, C. C. & Murray, A. W. GFP tagging of budding yeast chromosomes reveals that protein-protein interactions can mediate sister chromatid cohesion. Curr. Biol. 6, 1599–1608 (1996).
Hensel, Z., Weng, X., Lagda, A. C. & Xiao, J. Transcription-factor-mediated DNA looping probed by high-resolution, single-molecule imaging in live E. coli cells. PLoS Biol. 11, e1001591 (2013).
Liao, Y., Schroeder, J. W., Gao, B., Simmons, L. A. & Biteen, J. S. Single-molecule motions and interactions in live cells reveal target search dynamics in mismatch repair. Proc. Natl Acad. Sci. USA 112, E6898–E6906 (2015).
Uphoff, S., Reyes-Lamothe, R., Garza de Leon, F., Sherratt, D. J. & Kapanidis, A. N. Single-molecule DNA repair in live bacteria. Proc. Natl Acad. Sci. USA 110, 8063–8068 (2013).
Tang, M. et al. UmuD'(2)C is an error-prone DNA polymerase, Escherichia coli pol V. Proc. Natl Acad. Sci. USA 96, 8919–8924 (1999).
Robinson, A. et al. Regulation of mutagenic DNA polymerase V activation in space and time. PLoS Genet. 11, e1005482 (2015).
Lesterlin, C., Ball, G., Schermelleh, L. & Sherratt, D. J. RecA bundles mediate homology pairing between distant sisters during DNA break repair. Nature 506, 249–253 (2014).
Rajendram, M. et al. Anionic phospholipids stabilize RecA filament bundles in Escherichia coli. Mol Cell 60, 374–384 (2015).
Bakshi, S., Dalrymple, R. M., Li, W., Choi, H. & Weisshaar, J. C. Partitioning of RNA polymerase activity in live Escherichia coli from analysis of single-molecule diffusive trajectories. Biophys. J. 105, 2676–2686 (2013).
Bakshi, S., Siryaporn, A., Goulian, M. & Weisshaar, J. C. Superresolution imaging of ribosomes and RNA polymerase in live Escherichia coli cells. Mol. Microbiol. 85, 21–38 (2012).
Endesfelder, U. et al. Multiscale spatial organization of RNA polymerase in Escherichia coli. Biophys. J. 105, 172–181 (2013).
Cagliero, C., Zhou, Y. N. & Jin, D. J. Spatial organization of transcription machinery and its segregation from the replisome in fast-growing bacterial cells. Nucleic Acids Res. 42, 13696–13705 (2014).
Jin, D. J., Cagliero, C., Martin, C. M., Izard, J. & Zhou, Y. N. The dynamic nature and territory of transcriptional machinery in the bacterial chromosome. Front. Microbiol. 6, 497 (2015).
Bath, J., Wu, L. J., Errington, J. & Wang, J. C. Role of Bacillus subtilis SpoIIIE in DNA transport across the mother cell-prespore division septum. Science 290, 995–997 (2000).
Yen Shin, J. et al. Visualization and functional dissection of coaxial paired SpoIIIE channels across the sporulation septum. eLife 4, e06474 (2015).
Fiche, J. B. et al. Recruitment, assembly, and molecular architecture of the SpoIIIE DNA pump revealed by superresolution microscopy. PLoS Biol 11, e1001557 (2013).
Ptacin, J. L. et al. Bacterial scaffold directs pole-specific centromere segregation. Proc. Natl Acad. Sci. USA 111, E2046–E2055 (2014).
de Boer, P., Crossley, R. & Rothfield, L. The essential bacterial cell-division protein FtsZ is a GTPase. Nature 359, 254–256 (1992).
RayChaudhuri, D. & Park, J. T. Escherichia coli cell-division gene ftsZ encodes a novel GTP-binding protein. Nature 359, 251–254 (1992).
Fu, G. et al. In vivo structure of the E. coli FtsZ-ring revealed by photoactivated localization microscopy (PALM). PLoS ONE 5, e12682 (2010).
Strauss, M. P. et al. 3D-SIM super resolution microscopy reveals a bead-like arrangement for FtsZ and the division machinery: implications for triggering cytokinesis. PLoS Biol 10, e1001389 (2012).
Holden, S. J. et al. High throughput 3D super-resolution microscopy reveals Caulobacter crescentus in vivo Z-ring organization. Proc. Natl Acad. Sci. USA 111, 4566–4571 (2014).
Fleurie, A. et al. MapZ marks the division sites and positions FtsZ rings in Streptococcus pneumoniae. Nature 516, 259–262 (2014).
Jacq, M. et al. Remodeling of the Z-ring nanostructure during the Streptococcus pneumoniae cell cycle revealed by photoactivated localization microscopy. mBio 6, e01108–15 (2015).
Leisch, N. et al. Growth in width and FtsZ ring longitudinal positioning in a gammaproteobacterial symbiont. Curr. Biol. 22, R831–R832 (2012).
Grangeon, R., Zupan, J. R., Anderson-Furgeson, J. & Zambryski, P. C. PopZ identifies the new pole, and PodJ identifies the old pole during polar growth in Agrobacterium tumefaciens. Proc. Natl Acad. Sci. USA 112, 11666–11671 (2015).
Coltharp, C., Buss, J., Plumer, T. M. & Xiao, J. Defining the rate-limiting processes of bacterial cytokinesis. Proc. Natl Acad. Sci. USA 113, E1044–E1053 (2016).
Rowlett, V. W. & Margolin, W. 3D-SIM super-resolution of FtsZ and its membrane tethers in Escherichia coli cells. Biophys. J. 107, L17–L20 (2014).
Biteen, J. S., Goley, E. D., Shapiro, L. & Moerner, W. E. Three-dimensional super-resolution imaging of the midplane protein FtsZ in live Caulobacter crescentus cells using astigmatism. ChemPhysChem 13, 1007–1012 (2012).
Lyu, Z., Carla Coltharp, C., Yang, X. & Xiao, J. Influence of FtsZ GTPase activity and concentration on nanoscale Z-ring structure in vivo revealed by three-dimensional Superresolution imaging. Biopolymers 105, 725–734 (2016).
Eswaramoorthy, P. et al. Cellular architecture mediates DivIVA ultrastructure and regulates min activity in Bacillus subtilis. mBio 2, e00257–11 (2011).
Buss, J. et al. In vivo organization of the FtsZ-ring by ZapA and ZapB revealed by quantitative super-resolution microscopy. Mol. Microbiol. 89, 1099–1120 (2013).
Buss, J. et al. A multi-layered protein network stabilizes the Escherichia coli FtsZ-ring and modulates constriction dynamics. PLoS Genet. 11, e1005128 (2015).
Olshausen, P. V. et al. Superresolution imaging of dynamic MreB filaments in B. subtilis—a multiple-motor-driven transport? Biophys. J. 105, 1171–1181 (2013).
Reimold, C., Defeu Soufo, H. J., Dempwolff, F. & Graumann, P. L. Motion of variable-length MreB filaments at the bacterial cell membrane influences cell morphology. Mol. Biol. Cell 24, 2340–2349 (2013).
Kim, S. Y., Gitai, Z., Kinkhabwala, A., Shapiro, L. & Moerner, W. E. Single molecules of the bacterial actin MreB undergo directed treadmilling motion in Caulobacter crescentus. Proc. Natl Acad. Sci. USA 103, 10929–10934 (2006).
Biteen, J. S. et al. Super-resolution imaging in live Caulobacter crescentus cells using photoswitchable EYFP. Nat. Methods 5, 947–949 (2008).
Ptacin, J. L. et al. A spindle-like apparatus guides bacterial chromosome segregation. Nat. Cell Biol. 12, 791–798 (2010).
Ausmees, N., Kuhn, J. R. & Jacobs-Wagner, C. The bacterial cytoskeleton: an intermediate filament-like function in cell shape. Cell 115, 705–713 (2003).
Lew, M. D. et al. Three-dimensional superresolution colocalization of intracellular protein superstructures and the cell surface in live Caulobacter crescentus. Proc. Natl Acad. Sci. USA 108, E1102–E1110 (2011).
Greenfield, D. et al. Self-organization of the Escherichia coli chemotaxis network imaged with super-resolution light microscopy. PLoS Biol 7, e1000137 (2009).
Fredlund, J., Broder, D., Fleming, T., Claussin, C. & Pogliano, K. The SpoIIQ landmark protein has different requirements for septal localization and immobilization. Mol. Microbiol. 89, 1053–1068 (2013).
Haas, B. L., Matson, J. S., DiRita, V. J. & Biteen, J. S. Single-molecule tracking in live Vibrio cholerae reveals that ToxR recruits the membrane-bound virulence regulator TcpP to the toxT promoter. Mol Microbiol 96, 4–13 (2015).
Llorente-Garcia, I. et al. Single-molecule in vivo imaging of bacterial respiratory complexes indicates delocalized oxidative phosphorylation. Biochim. Biophys. Acta 1837, 811–824 (2014).
Karunatilaka, K. S., Cameron, E. A., Martens, E. C., Koropatkin, N. M. & Biteen, J. S. Superresolution imaging captures carbohydrate utilization dynamics in human gut symbionts. mBio 5, e02172 (2014).
Dufrene, Y. F., Boonaert, C. J., Gerin, P. A., Asther, M. & Rouxhet, P. G. Direct probing of the surface ultrastructure and molecular interactions of dormant and germinating spores of Phanerochaete chrysosporium. J. Bacteriol. 181, 5350–5354 (1999).
Schabert, F., Henn, C. & Engel, A. Native Escherichia coli OmpF porin surfaces probed by atomic force microscopy. Science 268, 92–94 (1995).
Kailas, L. et al. Surface architecture of endospores of the Bacillus cereus/anthracis/thuringiensis family at the subnanometer scale. Proc. Natl Acad. Sci. 108, 16014–16019 (2011).
Casuso, I., Kodera, N., Le Grimellec, C., Ando, T. & Scheuring, S. Contact-mode high-resolution high-speed atomic force microscopy movies of the purple membrane. Biophys. J. 97, 1354–1361 (2009).
Shibata, M., Yamashita, H., Uchihashi, T., Kandori, H. & Ando, T. High-speed atomic force microscopy shows dynamic molecular processes in photoactivated bacteriorhodopsin. Nat. Nanotech. 5, 208–212 (2010).
Yamashita, H. et al. Role of trimer–trimer interaction of bacteriorhodopsin studied by optical spectroscopy and high-speed atomic force microscopy. J. Struct. Biol. 184, 2–11 (2013).
Casuso, I. et al. Characterization of the motion of membrane proteins using high-speed atomic force microscopy. Nat. Nanotech. 7, 525–529 (2012).
Turner, R. D., Hurd, A. F., Cadby, A., Hobbs, J. K. & Foster, S. J. Cell wall elongation mode in Gram-negative bacteria is determined by peptidoglycan architecture. Nat. Commun. 4, 1496 (2013).
Turner, R. D., Vollmer, W. & Foster, S. J. Different walls for rods and balls: the diversity of peptidoglycan. Mol. Microbiol. 91, 862–874 (2014).
Hayhurst, E. J., Kailas, L., Hobbs, J. K. & Foster, S. J. Cell wall peptidoglycan architecture in Bacillus subtilis. Proc. Natl Acad. Sci. 105, 14603–14608 (2008).
Turner, R. D. et al. Peptidoglycan architecture can specify division planes in Staphylococcus aureus. Nat. Commun. 1, 26 (2010).
Wheeler, R., Mesnage, S., Boneca, I. G., Hobbs, J. K. & Foster, S. J. Super-resolution microscopy reveals cell wall dynamics and peptidoglycan architecture in ovococcal bacteria. Mol. Microbiol. 82, 1096–1109 (2011).
Plomp, M., Leighton, T. J., Wheeler, K. E., Hill, H. D. & Malkin, A. J. In vitro high-resolution structural dynamics of single germinating bacterial spores. Proc. Natl Acad. Sci. 104, 9644–9649 (2007).
Andre, G. et al. Imaging the nanoscale organization of peptidoglycan in living Lactococcus lactis cells. Nat. Commun. 1, 27 (2010).
Lotan, R., Sharon, N. & Mirelman, D. Interaction of wheat-germ agglutinin with bacterial cells and cell-wall polymers. Eur. J. Biochem. 55, 257–262 (1975).
Kuru, E. et al. In Situ probing of newly synthesized peptidoglycan in live bacteria with fluorescent D-amino acids. Angew. Chem. Int. Ed. Engl. 51, 12519–12523 (2012).
Liechti, G. W. et al. A new metabolic cell-wall labelling method reveals peptidoglycan in Chlamydia trachomatis. Nature 506, 507–510 (2014).
Sheldrick, G. M., Jones, P. G., Kennard, O., Williams, D. H. & Smith, G. A. Structure of vancomycin and its complex with acetyl-D-alanyl-D-alanine. Nature 271, 223–225 (1978).
Monteiro, J. M. et al. Cell shape dynamics during the staphylococcal cell cycle. Nat. Commun. 6, 8055 (2015).
van Teeseling, M. C. et al. Anammox Planctomycetes have a peptidoglycan cell wall. Nat. Commun. 6, 6878 (2015).
Tsui, H. C. et al. Pbp2x localizes separately from Pbp2b and other peptidoglycan synthesis proteins during later stages of cell division of Streptococcus pneumoniae D39. Mol. Microbiol. 94, 21–40 (2014).
Lee, T. K. et al. A dynamically assembled cell wall synthesis machinery buffers cell growth. Proc. Natl Acad. Sci. USA 111, 4554–4559 (2014).
Land, A. D. et al. Requirement of essential Pbp2x and GpsB for septal ring closure in Streptococcus pneumoniae D39. Mol. Microbiol. 90, 939–955 (2013).
Kocaoglu, O. et al. Selective penicillin-binding protein imaging probes reveal substructure in bacterial cell division. ACS Chem. Biol. 7, 1746–1753 (2012).
Bustanji, Y. et al. Dynamics of the interaction between a fibronectin molecule and a living bacterium under mechanical force. Proc. Natl Acad. Sci. 100, 13292–13297 (2003).
Lower, S. K. et al. Polymorphisms in fibronectin binding protein A of Staphylococcus aureus are associated with infection of cardiovascular devices. Proc. Natl Acad. Sci. USA 108, 18372–18377 (2011).
Herman, P. et al. The binding force of the staphylococcal adhesin SdrG is remarkably strong. Mol. Microbiol. 93, 356–368 (2014).
Herman-Bausier, P. & Dufrêne, Y. F. Atomic force microscopy reveals a dual collagen-binding activity for the staphylococcal surface protein SdrF. Mol. Microbiol. 99, 611–621 (2015).
Herman-Bausier, P., El-Kirat-Chatel, S., Foster, T. J., Geoghegan, J. A. & Dufrêne, Y. F. Staphylococcus aureus fibronectin-binding protein A mediates cell-cell adhesion through low-affinity homophilic bonds. mBio 6, e00413–15 (2015).
Gruszka, D. T. et al. Cooperative folding of intrinsically disordered domains drives assembly of a strong elongated protein. Nat. Commun. 6, 7271 (2015).
El- Kirat-Chatel, S. et al. Forces in yeast flocculation. Nanoscale 7, 1760–1767 (2015).
El- Kirat-Chatel, S. & Dufrêne, Y. F. Nanoscale adhesion forces between the fungal pathogen Candida albicans and macrophages. Nanoscale Horiz. 1, 69–74 (2016).
Formosa-Dague, C. et al. Forces between Staphylococcus aureus and human skin. Nanoscale Horiz. 1, 298–303 (2016).
Berk, V. et al. Molecular architecture and assembly principles of Vibrio cholerae biofilms. Science 337, 236–239 (2012).
Turnbull, L. et al. Explosive cell lysis as a mechanism for the biogenesis of bacterial membrane vesicles and biofilms. Nature Commun. 7, 11220 (2016).
Ivanov, I. E. et al. Atomic force and super-resolution microscopy support a role for LapA as a cell-surface biofilm adhesin of Pseudomonas fluorescens. Res. Microbiol. 163, 685–691 (2012).
Fritz, J. et al. Translating biomolecular recognition into nanomechanics. Science 288, 316–318 (2000).
Ndieyira, J. W. et al. Nanomechanical detection of antibiotic–mucopeptide binding in a model for superbug drug resistance. Nat. Nanotech. 3, 691–696 (2008).
Longo, G. et al. Rapid detection of bacterial resistance to antibiotics using AFM cantilevers as nanomechanical sensors. Nat. Nanotech. 8, 522–526 (2013).
Kasas, S. et al. Detecting nanoscale vibrations as signature of life. Proc. Natl Acad. Sci. 112, 378–381 (2014).
Beaussart, A., Abellán-Flos, M., El-Kirat-Chatel, S., Vincent, S. P. & Dufrêne, Y. F. Force nanoscopy as a versatile platform for quantifying the activity of antiadhesion compounds targeting bacterial pathogens. Nano Lett. 16, 1299–1307 (2016).
El- Kirat-Chatel, S. & Dufrêne, Y. F. Nanoscale imaging of the Candida-macrophage interaction using correlated fluorescence-atomic force microscopy. ACS Nano 2012, 10792–10799 (2012).
Andre, G. et al. Fluorescence and atomic force microscopy imaging of wall teichoic acids in Lactobacillus plantarum. ACS Chem. Biol. 6, 366–376 (2011).
Odermatt, P. D. et al. High-resolution correlative microscopy: bridging the gap between single molecule localization microscopy and atomic force microscopy. Nano Lett. 15, 4896–4904 (2015).
Acknowledgements
Work in the Dufrêne team was supported by the European Research Council (ERC) under the European Union's Horizon 2020 research and innovation programme (grant agreement no. 693630), the National Fund for Scientific Research (FNRS), the FNRS-WELBIO (grant no. WELBIO-CR-2015A-05), the Federal Office for Scientific, Technical and Cultural Affairs (Interuniversity Poles of Attraction Programme), and the Research Department of the Communauté française de Belgique (Concerted research action). Y.F.D. is Research Director at the FNRS. Work in the Xiao lab is supported by National Institute of Health General Medicines 1R01GM086447-06, 1R01GM112008-01 (Multi-PI), National Science Foundation grant EAGER MCB1019000. Y.F.D. and J.X. thank Carla Coltharp and Xinxing Yang for their critical reading and suggestions of the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interest.
Rights and permissions
About this article
Cite this article
Xiao, J., Dufrêne, Y. Optical and force nanoscopy in microbiology. Nat Microbiol 1, 16186 (2016). https://doi.org/10.1038/nmicrobiol.2016.186
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2016.186
This article is cited by
-
Application of nanotags and nanobodies for live cell single-molecule imaging of the Z-ring in Escherichia coli
Current Genetics (2023)
-
Seeing the unseen: High-resolution AFM imaging captures antibiotic action in bacterial membranes
Nature Communications (2022)
-
Force spectroscopy of single cells using atomic force microscopy
Nature Reviews Methods Primers (2021)
-
Mechanomicrobiology: how bacteria sense and respond to forces
Nature Reviews Microbiology (2020)
-
A review on anti-adhesion therapies of bacterial diseases
Infection (2019)