Abstract
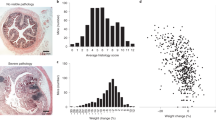
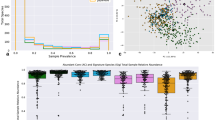
Inflammatory bowel disease (IBD) is an autoimmune condition that is difficult to diagnose, and animal models of this disease have questionable human relevance1. Here, we show that the dysbiosis network underlying IBD in dogs differs from that in humans, with some bacteria such as Fusobacterium switching roles between the two species (as Bacteroides fragilis switches roles between humans and mice)2. For example, a dysbiosis index trained on humans fails when applied to dogs, but a dog-specific dysbiosis index achieves high correlations with the overall dog microbial community diversity patterns. In addition, a random forest classifier trained on dog-specific samples achieves high discriminatory power, even when using stool samples rather than the mucosal biopsies required for high discriminatory power in humans2. These relationships were not detected in previously published dog IBD data sets due to their limited sample size and statistical power3. Taken together, these results reveal the need to train host-specific dysbiosis networks and point the way towards a generalized understanding of IBD across different mammalian models.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Jergens, A. E. & Simpson, K. W. Inflammatory bowel disease in veterinary medicine. Front. Biosci. (Elite Ed.) 4, 1404–1419 (2012).
Gevers, D. et al. The treatment-naive microbiome in new-onset Crohn's disease. Cell Host Microbe 15, 382–392 (2014).
Honneffer, J. B., Minamoto, Y. & Suchodolski, J. S. Microbiota alterations in acute and chronic gastrointestinal inflammation of cats and dogs. World J. Gastroenterol. 20, 16489–16497 (2014).
Hayward, J. J. et al. Complex disease and phenotype mapping in the domestic dog. Nat. Commun. 7, 10460 (2016).
Day, M. J. et al. Histopathological standards for the diagnosis of gastrointestinal inflammation in endoscopic biopsy samples from the dog and cat: a report from the World Small Animal Veterinary Association Gastrointestinal Standardization Group. J. Comp. Pathol. 138(Suppl. 1), S1–S43 (2008).
Simpson, K. W. et al. Adherent and invasive Escherichia coli is associated with granulomatous colitis in boxer dogs. Infect. Immun. 74, 4778–4792 (2006).
Suchodolski, J. S., Dowd, S. E., Wilke, V., Steiner, J. M. & Jergens, A. E. 16S rRNA gene pyrosequencing reveals bacterial dysbiosis in the duodenum of dogs with idiopathic inflammatory bowel disease. PLoS ONE 7, e39333 (2012).
Suchodolski, J. S. et al. The fecal microbiome in dogs with acute diarrhea and idiopathic inflammatory bowel disease. PLoS ONE 7, e51907 (2012).
Frank, D. N. et al. Disease phenotype and genotype are associated with shifts in intestinal-associated microbiota in inflammatory bowel diseases. Inflamm. Bowel Dis. 17, 179–184 (2011).
Minamoto, Y. et al. Alteration of the fecal microbiota and serum metabolite profiles in dogs with idiopathic inflammatory bowel disease. Gut Microbes 6, 33–47 (2015).
Breiman, L. Random forests. Machine Learning 45, 5–32 (2001).
Castellarin, M. et al. Fusobacterium nucleatum infection is prevalent in human colorectal carcinoma. Genome Res. 22, 299–306 (2012).
Suchodolski, J. S., Camacho, J. & Steiner, J. M. Analysis of bacterial diversity in the canine duodenum, jejunum, ileum, and colon by comparative 16S rRNA gene analysis. FEMS Microbiol. Ecol. 66, 567–578 (2008).
Ley, R. E. et al. Evolution of mammals and their gut microbes. Science 320, 1647–1651 (2008).
Muegge, B. D. et al. Diet drives convergence in gut microbiome functions across mammalian phylogeny and within humans. Science 332, 970–974 (2011).
Song, S. J. et al. Cohabiting family members share microbiota with one another and with their dogs. eLife 2, e00458 (2013).
Hedges, S. B., Marin, J., Suleski, M., Paymer, M. & Kumar, S. Tree of life reveals clock-like speciation and diversification. Mol. Biol. Evol. 32, 835–845 (2015).
Price, S. A., Hopkins, S. S., Smith, K. K. & Roth, V. L. Tempo of trophic evolution and its impact on mammalian diversification. Proc. Natl Acad. Sci. USA 109, 7008–7012 (2012).
Wesley-Hunt, G. D. & Flynn, J. J. Phylogeny of the Carnivora: basal relationships among the carnivoramorphans, and assessment of the position of ‘Miacoidea’ relative to Carnivora. J. System. Palaeontol. 3, 1–28 (2005).
Langille, M. G. et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 31, 814–821 (2013).
Mardini, H. E. & Grigorian, A. Y. Probiotic mix VSL#3 is effective adjunctive therapy for mild to moderately active ulcerative colitis: a meta-analysis. Inflamm. Bowel Dis. 20, 1562–1567 (2014).
Rossi, G. et al. Comparison of microbiological, histological, and immunomodulatory parameters in response to treatment with either combination therapy with prednisone and metronidazole or probiotic VSL#3 strains in dogs with idiopathic inflammatory bowel disease. PLoS ONE 9, e94699 (2014).
Uronis, J. M. et al. Gut microbial diversity is reduced by the probiotic VSL#3 and correlates with decreased TNBS-induced colitis. Inflamm. Bowel Dis. 17, 289–297 (2011).
Mawby, D. I. et al. Comparison of various methods for estimating body fat in dogs. J. Am. Anim. Hosp. Assoc. 40, 109–114 (2004).
EMP protocols and standards Earth Microbiome Project (2015); http://www.earthmicrobiome.org/emp-standard-protocols/
McDonald, D. et al. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 6, 610–618 (2012).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Weiss, S. J. et al. Effects of library size variance, sparsity, and compositionality on the analysis of microbiome data. PeerJ. PrePrints 3, e1408 (2015).
Perez, F. & Granger, B. E. IPython: a system for interactive scientific computing. Comput. Sci. Eng. 9, 21–29 (2007).
Lozupone, C. & Knight, R. Unifrac: a new phylogenetic method for comparing microbial communities. Appl. Environ. Microbiol. 71, 8228–8235 (2005).
Kuhn, M. Building predictive models in R using the caret package. J. Stat. Soft. 28, 1–26 (2008).
van der Walt, S., Colbert, S. C. & Varoquaux, G. The numPy array: a structure for efficient numerical computation. Comput. Sci. Eng. 13, 22–30 (2011).
Seaborn: Statistical Data Visualization (Waskom, M., 2012); https://stanford.edu/~mwaskom/software/seaborn/
Vazquez-Baeza, Y., Pirrung, M., Gonzalez, A. & Knight, R. EMPeror: a tool for visualizing high-throughput microbial community data. Gigascience 2, 16 (2013).
Detecting statistically significant associtations between sparse and high dimensional compositional data (Schwager, E. E. A. et al., 2016); http://huttenhower.sph.harvard.edu/ccrepe
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Stone, L. & Roberts, A. The checkerboard score and species distributions. Oecologia 85, 74–79 (1990).
Acknowledgements
The authors acknowledge support provided by the Crohn's and Colitis Foundation of American, the Templeton Foundation and the Keck Foundation (via the Earth Microbiome Project), the National institutes of Health. The authors thank Z. Xu, J. Sanders, A. Amir, G. Ackermann, J. Morton, L. Ursell, J. Metcalf, A. Gonzalez and E. Schwager for their comments and feedback on the manuscript.
Author information
Authors and Affiliations
Contributions
Y.V.-B. wrote the manuscript and managed, interpreted and analysed the data. E.R.H. contributed to the manuscript and analysed the data. J.S.S. contributed to the manuscript and analysed and interpreted the data. R.K. wrote the manuscript and interpreted the data. All authors worked together to finalize and approve this manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary Tables 1–3, Supplementary Figures 1–5. (PDF 2637 kb)
Rights and permissions
About this article
Cite this article
Vázquez-Baeza, Y., Hyde, E., Suchodolski, J. et al. Dog and human inflammatory bowel disease rely on overlapping yet distinct dysbiosis networks. Nat Microbiol 1, 16177 (2016). https://doi.org/10.1038/nmicrobiol.2016.177
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2016.177
This article is cited by
-
Immune-mediated hematological disease in dogs is associated with alterations of the fecal microbiota: a pilot study
Animal Microbiome (2023)
-
A comprehensive analysis of gut and skin microbiota in canine atopic dermatitis in Shiba Inu dogs
Microbiome (2023)
-
Noninvasive sampling of the small intestinal chyme for microbiome, metabolome and antimicrobial resistance genes in dogs, a proof of concept
Animal Microbiome (2023)
-
Gut microbiome signatures of Yorkshire Terrier enteropathy during disease and remission
Scientific Reports (2023)
-
The Effect of Lactobacillus sakei on Growth Performance and Intestinal Health in Dogs: Gut Microbiota and Metabolism Study
Probiotics and Antimicrobial Proteins (2023)