Abstract
Zika virus is a mosquito-borne pathogen that is rapidly spreading across the Americas. Due to associations between Zika virus infection and a range of fetal maladies1,2, the epidemic trajectory of this viral infection poses a significant concern for the nearly 15 million children born in the Americas each year. Ascertaining the portion of this population that is truly at risk is an important priority. One recent estimate3 suggested that 5.42 million childbearing women live in areas of the Americas that are suitable for Zika occurrence. To improve on that estimate, which did not take into account the protective effects of herd immunity, we developed a new approach that combines classic results from epidemiological theory with seroprevalence data and highly spatially resolved data about drivers of transmission to make location-specific projections of epidemic attack rates. Our results suggest that 1.65 (1.45–2.06) million childbearing women and 93.4 (81.6–117.1) million people in total could become infected before the first wave of the epidemic concludes. Based on current estimates of rates of adverse fetal outcomes among infected women2,4,5, these results suggest that tens of thousands of pregnancies could be negatively impacted by the first wave of the epidemic. These projections constitute a revised upper limit of populations at risk in the current Zika epidemic, and our approach offers a new way to make rapid assessments of the threat posed by emerging infectious diseases more generally.
Similar content being viewed by others
On 1 February 2016, the World Health Organization (WHO) designated the ongoing Zika virus epidemic in the Americas as a Public Health Emergency of International Concern (PHEIC), defined as an ‘extraordinary event’ that ‘potentially require[s] a coordinated international response’6. This declaration acknowledges the high potential for Zika to establish across the Americas given that its dominant vector, the Aedes aegypti mosquito, is endophilic and occupies an exceptionally broad geographical range7. Concern underlying this rare WHO declaration also stems from an association between Zika virus infection in pregnant women and a range of adverse fetal outcomes2, most notably congenital microcephaly1. As of 30 June 2016, there were 1,674 confirmed cases of microcephaly associated with Zika virus infection in five countries8, and there is widespread concern that these numbers could increase further as the virus continues to spread across the Americas9.
A number of uncertainties surround the future of the Zika epidemic in the Americas, particularly questions about how many women may be at risk of having children with congenital microcephaly and other adverse outcomes associated with Zika virus infection10. Of women who become infected with Zika virus during a vulnerable stage of their pregnancy, evidence is emerging that 1–13% may go on to develop congenital microcephaly2,4,5. However, the number of women who become infected with Zika virus during that timeframe is difficult to ascertain. One recent study3 estimated that 5.42 million births occurred in 2015 in regions of the Americas with ‘suitability’ for Zika ‘occurrence’. Such estimates come with many caveats though, as they rely on a relatively limited number of reported cases and apply a method based on equilibrium assumptions to a situation involving active range expansion11. Most importantly, the estimate of 5.42 million births3 reflects the total population within a demarcated area and does not take into account that large fractions of the populations in those areas may remain uninfected due to herd immunity generated over the course of the first wave of the epidemic12,13.
To quantify the potential magnitude of the ongoing Zika epidemic in terms of people who realistically might become infected, we formulated and applied a method for projecting location-specific epidemic attack rates on highly spatially resolved human demographic projections14. The central concept behind our approach is that of the ‘first-wave’ epidemic. Zika and other mosquito-borne viruses have been known to exhibit explosive outbreaks, infecting as much as 75% of a population in a single year15. Classical epidemiological theory predicts that some proportion of a population will remain uninfected during an epidemic, because herd immunity eventually causes the epidemic to burn out12. A related prediction of this theory is that the proportion infected before epidemic burnout (that is, the epidemic attack rate) has a one-to-one relationship with the basic reproduction number, R0 (ref. 13). The latter quantity has a well-known mechanistic formulation for mosquito-borne pathogens16 that accommodates the effects of environmental drivers on transmission17,18. For example, the incubation periods of dengue viruses in the Ae. aegypti mosquitoes that transmit Zika virus have an empirically derived relationship with temperature18, which can in turn be used to inform calculations of R0. Together with similar relationships for other transmission parameters, it is possible to characterize R0, a fundamental measure of transmission potential, as a function of local environmental conditions.
We leveraged these classic results from epidemiological theory to first perform highly spatially resolved calculations of R0 and then to translate those calculations into location-specific projections of first-wave epidemic attack rates (Fig. 1). Because Zika-specific values of transmission parameters are largely unknown at present but may be well approximated by dengue-specific values19, we used some parameter values for dengue virus in our R0 calculations. We also calibrated our attack rate projections to match empirically estimated attack rates from 12 chikungunya epidemics and one Zika epidemic in naive populations (Supplementary Table 1). This step afforded us the flexibility to enhance the realism of the model with respect to firmly established but poorly quantified associations between human–mosquito contact and economic prosperity20. In doing so, one departure from the classic relationship between R0 and attack rate that we made was to rescale R0 by an exponent α ∈ (0, 1] to allow for better correspondence with observed attack rates. Although there is no theoretical justification for this or any other particular scaling relationship, it is consistent with theoretical expectations21 that attack rates should be lower in populations with equal R0 values but more heterogeneous contact patterns, which are typical for transmission by Ae. aegypti22. To provide a point of reference for our model-based approach, we also fitted a statistical description of the 13 seroprevalence estimates as a function of the environmental drivers that we considered. For both approaches, we applied their respective location-specific attack rate projections to demographic projections on a 5 km × 5 km grid across Latin America and the Caribbean to obtain the expected numbers of infections in the overall population and among childbearing women in particular (Fig. 2a). All such calculations were performed for 1,000 Monte Carlo samples of model parameters.
In total, our median projection suggests that as many as 93.4 (range: 81.6–117.1) million people in Latin America and the Caribbean could become infected during the first wave of the epidemic (Table 1). To place this number into context, we refer to an estimate23 that 53.8 (40.0–71.8) million dengue infections occurred in this region in 2010 alone. Our projections of nearly double this number for Zika are not surprising, given that there is extensive immunity to dengue but not Zika in this region and given that it would probably take longer than a year for the first wave of the epidemic to conclude in all locations within this region. At the country level, we project that Brazil will have the largest total number of infections by more than double that of any other country, due to a combination of its size and suitability for transmission. Island countries in the Caribbean are projected to experience the highest nationally averaged attack rates, with seven of the highest ten values projected for countries including Aruba, Haiti and Cuba. This projection is consistent with a frequent history of arbovirus outbreaks on islands24 and may be due to the uniformity of environmental conditions on the portions of islands where people tend to live. In more heterogeneous regions, the 5 km × 5 km spatial resolution of our maps allows for nuanced projections for areas of interest to local stakeholders (Fig. 2b,c). To facilitate the use of these local projections, we have made 5 km × 5 km minimum, median and maximum projections of attack rates, total infections and infections among childbearing women publically available online (http://github.com/TAlexPerkins/Zika_nmicrobiol_2016).
Among childbearing women, our median projection suggests that there could be as many as 1.65 (range: 1.45–2.06) million infections in Latin America and the Caribbean before the first wave of the epidemic concludes (Table 1). Assuming that birth rates are temporally constant, our projections are robust to uncertainty about the timing of local epidemics and the timeframe of the first wave of the epidemic, because they are based on cumulative proportions infected. These projections can also be used to postulate numbers at risk of microcephaly by multiplying them by the fraction of a year in which a pregnant woman is susceptible to developing microcephaly (for example, multiply by 1/4 in the case of first-trimester susceptibility). We also note that there were some discrepancies in our projections in terms of the rank order of countries experiencing the most infections among childbearing women versus the population as a whole. In particular, Cuba was fifth in terms of projected infections in the overall population but twelfth in terms of infections among childbearing women due to its low birth rate compared to other countries in the Americas25. Such discrepancies are also likely to exist subnationally26, and their elucidation should be a priority for future work.
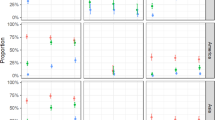
By accounting for uncertainty distributions for each of the key drivers of our model (Fig. 3a–e), we found that uncertainty distributions for infections across the region as a whole and by country were often multimodal (Fig. 3f–o) due to uncertainty in the shape of the relationship between mosquito–human contact and the local economic index that we considered (Fig. 3d). Summing our projections across Latin America and the Caribbean revealed variation that was modest, in the sense that none of our 1,000 Monte Carlo samples resulted in fewer than 81 million infections overall and 1.4 million among childbearing women (Fig. 3f,k). There are many reasons that even these numbers could be overestimates though. Our projections are conditioned on a local epidemic taking place in each 5 km × 5 km grid cell in the region, which is unlikely to happen given dispersal limitation, stochastic fadeout, geographic mismatches in seasonality and other factors. Therefore, it is most appropriate to interpret our projections as either a plausible worst-case scenario or an expectation of local epidemic size conditional on there being a local epidemic in the first place.
a–o, Monte Carlo samples from the uncertainty distributions surrounding each of the key drivers in the model (a–e) and uncertainty distributions for projected numbers of infections among all individuals (f–j) and among childbearing women (k–o) in different areas. All panels reflect the full range of uncertainty considered in 1,000 Monte Carlo samples. a, Posterior distributions of mosquito occurrence probabilities for two example 5 km × 5 km grid cells.
Although our approach was very much rooted in mechanistic models from epidemiological theory, two critical steps in our method involved fitting curves to describe theoretically motivated but heretofore unknown relationships: an association between mosquito–human contact and economic prosperity (Fig. 3d) and a scaling relationship between R0 and attack rates (Fig. 3e). Allowing these relationships to be informed by local seroprevalence estimates (Supplementary Table 1) left open the question of the extent to which our projections were informed by the mechanistic assumptions of the model versus statistical fits to the seroprevalence estimates that we used. On the one hand, an alternative statistical approach accounted for much more variation in seroprevalence estimates (R2 = 0.89) than did the model-based approach (R2 = 0.32). On the other hand, the statistical approach offered a dichotomous set of projections about numbers of infections outside the context of the data to which it was fitted: either everyone will become infected or very few people will (Fig. 4). Relationships between attack rates and predictor variables inferred by the statistical approach (Fig. 5d–i) were also implausible: a narrow temperature range in which attack rates increase sharply towards 100% (Fig. 5d–h) and a reversal of economic effects whereby wealthy populations experience higher attack rates than poor populations when mosquito occurrence probabilities are high (Fig. 5f,i). By contrast, the model-based approach yielded more moderate attack rate projections overall (Figs 3f versus 4a) in which temperature, economic prosperity and mosquito occurrence probability all had plausible relationships with attack rates (Fig. 5a–c).
a–j, Projected numbers of infections among all individuals (a–e) and among childbearing women (f–j) for 1,000 Monte Carlo samples from the uncertainty distribution around parameters of the statistical model that was fitted to seroprevalence estimates from Supplementary Table 1.
In conclusion, our model-based approach offers a unique way to leverage a variety of spatially detailed data products7,14,27,28 to make a priori projections of attack rates and infections that could be experienced in the first wave of the ongoing Zika epidemic. Projections such as these have an important role to play in the early stages of an epidemic, when planning for surveillance and outbreak response is actively under way both internationally and locally9. At the same time, it is important for consumers of this information to be aware of uncertainties in these and other projections, which often exceed the amount of uncertainty that can be identified a priori29. Similarly, following up on these projections in the aftermath of the epidemic—by comparing against projections made with alternative models and additional serological surveys30—will provide an exceptional opportunity to enhance capabilities to anticipate the severity of future epidemic threats.
Methods
Data sources and processing
Human demography. To estimate the annual numbers of pregnancies per 1 km × 1 km grid cell in 2015, methods developed by the WorldPop project (www.worldpop.org)25,31 were adapted for the Americas region. High-resolution estimates of population counts per 100 m × 100 m grid cell for 2015 were recently constructed for Latin American, Asian and African countries14,32. With consistent subnational data on sex and age structures, as well as subnational age-specific fertility rate data across the Americas currently unavailable for fully replicating the approaches of Tatem and colleagues31, national-level adjustments were made to construct pregnancy and birth counts. Data on estimated total numbers of births33 and pregnancies31 occurring annually in 2012 were assembled for all Latin American study countries, as well as births in 2015 (ref. 33). As no 2015 pregnancy estimates existed at the time of writing, the ratios of births to pregnancies for each country in the Americas were calculated using 2011 and 2012 estimates, and these were then applied to the 2015 birth numbers to obtain 2015 estimates of annual pregnancy numbers per country. This made the assumption that per-country births-to-pregnancies ratios remained the same in 2015 as they were in 2011 and 2012. The 100 m × 100 m gridded population totals were aggregated to 1 km × 1 km spatial resolution and the per-country totals were linearly adjusted to match the 2015 pregnancy estimates.
Temperature
We used interpolated meteorological station temperature data from the 1950–2000 period at 5 km × 5 km spatial resolution, processed to create climatological monthly averages that represent ‘typical’ conditions (www.worldclim.org)27.
Ae. aegypti occurrence probability
To predict the likely distribution of Ae. aegypti mosquitoes, Kraemer et al.7 generated high-resolution occurrence probability surfaces based on a species distribution modelling approach11. More specifically, a boosted regression tree model was applied using a comprehensive set of known occurrences (n = 19,930) of Ae. aegypti and a set of environmental predictors known to influence the distribution of the species7. Covariates included a temperature suitability index17, contemporary mean and range maps of the Enhanced Vegetation Index and precipitation34, and an urbanization index from the Global Rural Urban Mapping Project. We used a set of 100 spatial layers sampled from the posterior distribution estimated by Kraemer et al.7
Economic index
To account for socio-economic differences among populations residing in different regions, we used one-degree-resolution gridded estimates of purchasing power parity (PPP) in US$ from 2005 adjusted for inflation (G-Econ)28. When we encountered missing values, we imputed values in one of two ways. Grid cells in small island countries with data missing for the entire country were uniformly filled with population-adjusted PPP figures obtained from the US CIA World Factbook35. Missing values in continental grid cells were imputed with the mean of the surrounding eight grid cell values. Once we obtained a complete PPP grid layer at one-degree resolution, we resampled the layer to a resolution of 5 km × 5 km to match the resolution of gridded layers for human demography, temperature and Ae. aegypti occurrence probability.
Seroprevalence estimates
To calibrate our model, we identified published estimates of seroprevlance that were relevant to the context of our study (Supplementary Table 1). Specifically, we sought estimates of seroprevalence to either Zika or chikungunya viruses in populations that were presumably naive before an outbreak. Thus, we excluded some seroprevalence estimates that were obtained from endemic populations. We also excluded estimates from small islands—namely, Reunion and Grande Comore—for which it was clear that gridded temperature data were unrealistically low due to steep elevational gradients and other features of island geography. Although the focus of our analysis was on Latin America and the Caribbean, we were not able to exclude locations on the basis of location given that only 2 of 13 came from the focal region. Appropriately, however, a number of the seroprevalence estimates we obtained pertained specifically to pregnant women, although there did not appear to be differences in the seroprevalence of pregnant women and the population at large, at least in the context of a naive population following an outbreak36.
Calculation of derived quantities
Mosquito abundance. Occurrence probabilities can be translated into proxies for abundance provided that an assumption is made about how abundance is distributed as a random variable37. Assuming that mosquito abundance is distributed as a Poisson random variable, the probability that there is at least one mosquito present in a given location is 1 – exp(–λ), where λ is the expected abundance of mosquitoes. Inverting this relationship, we obtained an estimate λ = –ln(1 – occurrence probability) of expected mosquito abundance under the Poisson model and used this as a proxy for mosquito abundance in our calculations.
Mosquito–human ratios
The estimates of mosquito occurrence probability that we used incorporated a number of environmental variables7. They did not account for factors that modulate contact between mosquitoes and humans, however. Due in part to economic differences, factors such as air conditioning and piped water can drastically limit mosquito–human contact and virus transmission, even when mosquitoes are abundant20. We accounted for the effect of economic differences between locations by multiplying our proxy for mosquito abundance λ by a multiplication factor, which we specified as a function of the aforementioned economic index. We specified the relationship between the economic index and the multiplication factor by using a shape constrained additive model (SCAM38). This allowed for flexibility in the shape of this relationship but constrained it such that the multiplication factor (and thus, presumed mosquito–human contact) could only have a monotonically decreasing relationship with increasing values of the economic index. The specific shape of this relationship was determined by fitting it to values of the multiplication factors that would be necessary for modelled attack rates to perfectly match published seroprevalence estimates.
Basic reproduction number R0
We calculated the basic reproduction number R0 according to its classic Ross–Macdonald formulation and as a function of temperature T,
with adult mosquito mortality µ and extrinsic incubation period n specified as functions of temperature. Because temperature values were available for each location on a monthly basis, we computed monthly values of R0 for each location and then used the mean of the highest six monthly values of R0 as a singular estimate of R0 for each location. This approach was broadly consistent with the way in which a temperature suitability index was used to inform mosquito occurrence probabilities by Kraemer and co-authors7.
For mosquito mortality, we used the temperature- and age-dependent model of Brady and colleagues39, to which we added an additional force of extrinsic mortality (0.025 d−1) to match an overall daily mortality value of 0.115 estimated in a mark–release–recapture experiment carried out under temperatures ranging from 20 to 34 °C (ref. 40). We then computed the mean of the age- and temperature-dependent lifespan distribution as a function of temperature to inform μ(T). For the relationship between temperature and mean duration of the extrinsic incubation period, we used the temperature-dependent exponential rate estimated by Chan and Johansson18. The ratio of mosquitoes to humans, m, was quantified using a combination of occurrence probabilities and the gross cell product economic index, as described in the previous two sections. Parameters that did not depend on temperature were set at the following values according to published estimates for Ae. aegypti and dengue virus: mosquito-to-human transmission probability, b = 0.4 (ref. 41); human-to-mosquito transmission probability times number of days of human infectiousness, c/r = 3.5 (ref. 42); mosquito biting rate, a = 0.67 (ref. 43). Although there is uncertainty around these parameter values, any such uncertainty was effectively subsumed by fitting m to seroprevalence data given that bca2/r entered R0 as a constant.
Attack rates under a model-based formulation
Under a susceptible–infected–recovered (SIR) transmission model, there is a one-to-one relationship between R0 and final epidemic size, which is equivalent to the attack rate over the course of an epidemic13. Intuitively, the final epidemic size is reached once herd immunity is sufficient to limit contacts between infectious and susceptible individuals to the extent necessary to reduce the pathogen's force of infection to zero. There is no explicit solution for final epidemic size as a function of model parameters, but it can be calculated numerically by obtaining an implicit solution of for S∞, which is the proportion remaining susceptible after the epidemic has burned out13. Under the assumptions of the SIR model, the attack rate over the course of an epidemic is AR = 1 – S∞.
To apply this theoretical insight to Zika or other mosquito-borne pathogens, several limiting assumptions of the SIR model must first be reconciled. One such assumption is that individuals become infectious immediately upon becoming infected and remain infectious for an exponentially distributed period of time44; mosquito-borne pathogens such as Zika virus are instead characterized by a distinct lag between human and mosquito infection45. Despite this discrepancy between assumptions of the SIR model and the reality of many pathogen systems, mathematical analyses46 have shown that final epidemic size is insensitive to details about the shape of the distribution that characterizes the time period between successive cases (that is, the generation interval).
Another limiting assumption of the SIR model is that of homogeneous encounters between people and mosquitoes44, which are understood to be extensive for mosquito-borne diseases22. Mathematical analyses21 in this case show that a seemingly infinite complexity of relationships between R0 and final epidemic size are possible in a heterogeneous system. As a general rule, however, final epidemic size in a system with contact heterogeneity and proportional mixing is expected to be strictly less than the final epidemic size in an otherwise equivalent system with homogeneous contacts21. How the ratio of these final epidemic sizes scales as a function of R0 depends entirely on the details of a given system and would therefore be extremely difficult to generalize without copious data on mosquito–human contact and further investigation, which is beyond the scope of our study.
To capture the potentially very strong effects of heterogeneity in reducing final epidemic size in populations subject to Zika epidemics, we scaled the final epidemic size by substituting for R0 in the SIR-based final epidemic size formula given some constant α ∈ (0, 1]. Although there is no theoretical justification for this or any other choice of how to scale R0 and AR in the presence of contact heterogeneity, the choice we made has the following desirable properties: (1) it implies that AR → 1 as R0 → ∞; (2) it leads to the function AR(R0) having a more gradual slope and thereby allows for intermediate attack rates to be more common than they would be otherwise; (3) it preserves the property that AR = 0 for R0 < 1. At the same time, this and possible alternative formulations are limited by a general lack of understanding about the relationship between R0 and AR in heterogeneous systems, relationships that may furthermore be heterogeneous themselves across different areas47.
To estimate α, we performed the following procedure for candidate values of α between 0.01 and 1 in increments of 0.01: (1) calculate R0 according to equation (1) and assume m = λ for each of the 13 sites from which seroprevalence estimates were derived; (2) use those R0 values to calculate AR values for each of those sites based on the classic SIR formulation; (3) calculate what multiplication factor of R0 would be necessary for AR to match the empirical seroprevalence estimate; (4) fit a SCAM model of the economic index to the multiplication factors; and (5) use the fitted SCAM values to recalculate R0 and then AR for each site. Next, we calculated the sum of squares between the final predicted AR values associated with each α and the empirical seroprevalence estimates and we then selected the value of α that minimized the sum of squares. Supplementary Fig. 1 illustrates this process given mean estimates of λ from Ae. aegypti occurrence probabilities, μ(T) and n(T).
Attack rates under a statistical formulation
As an alternative to our model-based characterization of attack rates, we also considered a purely statistical approach that modelled probit-transformed seroprevalence observations as functions of averaged monthly temperatures, Ae. aegypti occurrence probabilities and the economic index. We considered all combinations of linear, quadratic and pairwise interaction terms of these variables, comparing them on the basis of the Akaike Information Criterion using the lm and step functions in R (ref. 48). Although additional functional forms would have been of interest, this suite of models was as complex as the limited set of 13 seroprevalence observations would support.
Quantifying uncertainty around attack rate projections
To quantify uncertainty associated with our projections, we generated 1,000 Monte Carlo samples from the uncertainty distributions of each model parameter as described in each of the references7,17,18 in which those parameters were originally described. For μ(T) and n(T), we took random draws of their parameters consistent with published descriptions of uncertainty in the parameters of those functions from their original sources17,18. For Ae. aegypti occurrence probabilities, we drew randomly with replacement from 100 sample layers from the posterior distribution7. For the relationship involving the economic index and the R0 scaling factor α, we used best-fit SCAM models and α values corresponding to each set of random draws of the parameters of μ(T), n(T) and λ from the Ae. aegypti layers. For each of the 1,000 Monte Carlo samples of the statistical model, we performed resampling with replacement among the 13 seroprevalence values, performed the same model fitting and model selection procedure described in the previous section, and took a multivariate normal random sample of the parameter values of the best-fit model based on the model's best-fit parameters and variance–covariance matrix.
Projecting attack rates and numbers of infections
To obtain estimates of the numbers of infections in total and among childbearing women for the model-based and statistical approaches, we multiplied their respective attack rate projections applied to 5 km × 5 km grids across Latin America and the Caribbean by human demographic layers for total population and births in 2015. For both the model-based and statistical approaches, we performed these calculations and summed at the country level once for each of the 1,000 Monte Carlo samples that we produced. High-resolution spatial projections of attack rates and numbers of infected childbearing women under the model-based approach are presented in Supplementary Figs 2–10. Most projections based on the statistical approach resulted in attack rates of 100% in nearly all locations throughout Latin America and the Caribbean.
Code availability
Code in the R language for reproducing all analyses is available at http://github.com/TAlexPerkins/Zika_nmicrobiol_2016.
References
Mlakar, J. et al. Zika virus associated with microcephaly. New Engl. J. Med. 374, 951–958 (2016).
Brasil, P. et al. Zika virus infection in pregnant women in Rio de Janeiro—preliminary report. New Engl. J. Med. http://dx.doi.org/10.1056/NEJMoa1602412 (2016).
Messina, J. P. et al. Mapping global environmental suitability for Zika virus. eLife 5, 15272 (2016).
Cauchemez, S. et al. Association between Zika virus infection and microcephaly in French Polynesia, 2013–2015: a retrospective study. Lancet 387, 2125–2132 (2016).
Johansson, M. A. et al. Risk estimates for microcephaly related to Zika virus infection—from French Polynesia to Bahia, Brazil. Preprint at http://dx.doi.org/10.1101/051060 (2016).
WHO Statement on the First Meeting of the International Health Regulations (2005) (IHR 2005) Emergency Committee on Zika Virus and Observed Increase in Neurological Disorders and Neonatal Malformations (WHO, 2016); http://www.who.int/mediacentre/news/statements/2016/1st-emergency-committee-zika/en/
Kraemer, M. U. G. et al. The global distribution of the arbovirus vectors Aedes aegypti and Ae. albopictus. eLife 4, e08347 (2015).
Zika—Epidemiological Update (Pan American Health Organization, 2016).
Lucey, D. R. & Gostin, L. O. The emerging Zika pandemic: enhancing preparedness. J. Am. Med. Assoc. 315, 865–866 (2016).
The next steps on Zika. Nature 530, 5 (2016).
Elith, J. & Leathwick, J. Species distribution models: ecological explanation and prediction across space and time. Annu. Rev. Ecol. Evol. System. 40, 677–697 (2009).
Kermack, W. & McKendrick, A. A contribution to the mathematical theory of epidemics. Proc. R. Soc. A 115, 700–721 (1927).
Keeling, M. & Rohani, P. Modeling Infectious Diseases in Humans and Animals (Princeton Univ. Press, 2007).
Sorichetta, A. et al. High-resolution gridded population datasets for Latin America and the Caribbean in 2010, 2015, and 2020. Sci. Data 2, 150045 (2015).
Sergon, K. et al. Seroprevalence of chikungunya virus (CHIKV) infection on Lamu Island, Kenya, October 2004. Am. J. Trop. Med. Hyg. 78, 333–337 (2008).
Smith, D. L. et al. Ross, Macdonald, and a theory for the dynamics and control of mosquito-transmitted pathogens. PLOS Pathogens 8, e1002588 (2012).
Brady, O. J. et al. Global temperature constraints on Aedes aegypti and Ae. albopictus persistence and competence for dengue virus transmission. Parasites Vector. 7, 338 (2014).
Chan, M. & Johansson, M. A. The incubation periods of dengue viruses. PLoS ONE 7, e50972 (2012).
Funk, S. et al. Comparative analysis of dengue and Zika outbreaks reveals differences by setting and virus. Preprint at http://dx.doi.org/10.1101/043265 (2016).
Reiter, P. et al. Texas lifestyle limits transmission of dengue virus. Emerg. Infect. Dis. 9, 86–89 (2003).
Andreasen, V. The final size of an epidemic and its relation to the basic reproduction number. Bull. Math. Biol. 73, 2305–2321 (2011).
Liebman, K. A. et al. Determinants of heterogeneous blood feeding patterns by Aedes aegypti in Iquitos, Peru. PLoS Negl. Trop. Dis. 8, e2702 (2014).
Bhatt, S. et al. The global distribution and burden of dengue. Nature 496, 504–507 (2013).
Feldstein, L., Brownstein, J., Brady, O. J., Hay, S. I. & Johansson, M. A. Dengue on islands: a Bayesian approach to understanding the global ecology of dengue viruses. Trans. R. Soc. Trop. Med. Hyg. 109, 303–312 (2015).
The State of the World's Midwifery 2014 (UNFPA, 2014).
Tatem, A. J. et al. Millenium development health metrics: where do Africa's children and women of childbearing age live? Popul. Health Metrics 11, 11 (2013).
Hijmans, R. et al. Very high resolution interpolated climate surfaces for global land areas. Int. J. Climatol. 25, 1965–1978 (2005).
Nordhaus, W. Geography and macroeconomics: new data and new findings. Proc. Natl Acad. Sci. USA 103, 3510–3517 (2006).
Taylor, B. P. et al. Stochasticity and the limits to confidence when estimating R0 of Ebola and other emerging infectious diseases. Preprint at http://arxiv.org/abs/1601.06829 (2016).
Metcalf, C. J. E. et al. Use of serological surveys to generate key insights into the changing global landscape of infectious disease. Lancet http://dx.doi.org/10.1016/S0140-6736(16)30164-7 (2016).
Tatem, A. J. et al. Mapping for maternal and newborn health: the distributions of women of childbearing age, pregnancies and births. Int. J. Health Geogr. 13, 2 (2013).
Stevens, F. R. et al. Disaggregating census data for population mapping using random forests with remotely-sensed and ancillary data. PLoS ONE 10, e0107042 (2015).
World Population Prospects: The 2015 Revision (UN, 2015).
Weiss, D. et al. An effective approach for gap-filling continental scale remotely sensed time-series. ISPRS J. Photogr. Remote Sens. 98, 106–118 (2014).
The World Factbook (US Central Intelligence Agency, 2016); https://www.cia.gov/library/publications/the-world-factbook/geos/xx.html
Gerardin, P. et al. Estimating chikungunya prevalence in Reunion Island outbreak by serosurveys: two methods for two critical times of the epidemic. BMC Infect. Dis. 8, 99 (2008).
Wright, D. Correlations between incidence and abundance are expected by chance. J. Biogeogr. 18, 463–466 (1991).
Pya, N. & Wood, S. Shape constrained additive models. Statist. Comput. 25, 543–559 (2015).
Brady, O. J. et al. Modelling adult Aedes aegypti and Aedes albopictus survival at different temperatures in laboratory and field settings. Parasites Vector. 6, 351–362 (2013).
Muir, L. & Kay, B. Aedes aegypti survival and dispersal estimated by mark–release–recapture in northern Australia. Am. J. Trop. Med. Hyg. 58, 277–282 (1998).
Nishiura, H. & Halstead, S. Natural history of dengue virus (DENV)-1 and DENV-4 infections: reanalysis of classic studies. J. Infect. Dis. 195, 1007–1013 (2007).
Scott, T. W. et al. Longitudinal studies of Aedes aegypti (Diptera: Culicidae) in Thailand and Puerto Rico: blood feeding frequency. J. Med. Entomol. 37, 89–101 (2000).
Reiner, R. C. et al. A systematic review of mathematical models of mosquito-borne pathogen transmission: 1970–2010. J. R. Soc. Interface 10, 20120921 (2013).
Lessler, J. et al. Times to key events in the course of Zika infection and their implications for surveillance: a systematic review and pooled analysis. Preprint at http://dx.doi.org/10.1101/041913 (2016).
Ma, J. & Earn, D. J. Generality of the final size formula for an epidemic of a newly invading infectious disease. Bull. Math. Biol. 68, 679–702 (2006).
Kraemer, M. U. G. et al. Big city, small world: density, contact rates, and transmission of dengue across Pakistan. J. R. Soc. Interface 12, 20150468 (2015).
R Core Team R: A Language for Statistical Computing (R Foundation for Statistical Computing, 2014); http://www.R-project.org/
Duffy, M. et al. Zika virus outbreak on Yap Island, Federated States of Micronesia. New Engl. J. Med. 360, 2536–2543 (2009).
Schwarz, N. et al. Seroprevalence of antibodies against chikungunya, dengue, and Rift Valley fever viruses after febrile illness outbreak, Madagascar. Emerg. Infect. Dis. 18, 1780–1786 (2012).
Sissoko, D. et al. Outbreak of chikungunya fever in Mayotte, Comoros archipelago, 2005–2006. Trans. R. Soc. Trop. Med. Hyg. 102, 780–786 (2008).
Dwibedi, B. et al. Rapid spread of chikungunya virus infection in Orissa, India. Indian J. Med. Res. 133, 316–321 (2011).
Gay, N. et al. Seroprevalence and asymptomatic rates of Asian lineage chikungunya virus infection on Saint Martin, Caribbean. Am. J. Trop. Med. Hyg. 94, 393–396 (2015).
Moro, M. et al. Chikungunya virus in north-eastern Italy: a seroprevalence study. Am. J. Trop. Med. Hyg. 82, 508–511 (2010).
Balmaseda, A. et al. Clinical attack rate of chikungunya in a cohort of Nicaraguan children. Am. J. Trop. Med. Hyg. 94, 397–399 (2016).
Acknowledgements
The authors thank three anonymous reviewers, as well as J. Ashander, C.M. Barker, M.A. Johansson, R.C. Reiner, S.T. Stoddard, J.C. Miller and members of the Perkins Lab for discussions. The authors thank O.J. Brady for sharing code for calculating mosquito mortality as a function of temperature. T.A.P., A.S.S. and A.J.T. are supported by funding from the National Science Foundation (DEB 1641130). C.W.R. is supported by funding through the University of Southampton's Economic and Social Research Council's Doctoral Training Centre. M.U.G.K. receives funding from the International Research Consortium on Dengue Risk Assessment Management and Surveillance (IDAMS; European Commission 7th Framework Programme, 21893). A.J.T. is supported by funding from NIH/NIAID (U19AI089674), the BMGF (OPP1106427, 1032350), NORAD and a Wellcome Trust Sustaining Health Grant (106866/Z/15/Z). A.J.T. and C.W.R. acknowledge the support of the WorldPop (www.worldpop.org) and Flowminder Foundation (www.flowminder.org) teams in demographic data set production, and T.A.P. and A.S.S. acknowledge support from the Notre Dame Center for Research Computing.
Author information
Authors and Affiliations
Contributions
T.A.P. conceived the research, designed the analysis and wrote the first draft of the manuscript. A.S.S. assembled data, performed calculations and contributed to writing. C.W.R. assembled data, produced map visuals and contributed to writing. M.U.G.K. assembled data and contributed to writing. A.J.T. contributed to the analysis, map visuals and writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary Table 1, Supplementary Figures 1–10 (PDF 816 kb)
Rights and permissions
About this article
Cite this article
Perkins, T., Siraj, A., Ruktanonchai, C. et al. Model-based projections of Zika virus infections in childbearing women in the Americas. Nat Microbiol 1, 16126 (2016). https://doi.org/10.1038/nmicrobiol.2016.126
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2016.126
This article is cited by
-
Antibody seropositivity and endemicity of chikungunya and Zika viruses in Nigeria
Animal Diseases (2023)
-
Zika virus M protein latches and locks the E protein from transitioning to an immature state after prM cleavage
npj Viruses (2023)
-
Zika, chikungunya and co-occurrence in Brazil: space-time clusters and associated environmental–socioeconomic factors
Scientific Reports (2023)
-
Trade-offs between individual and ensemble forecasts of an emerging infectious disease
Nature Communications (2021)
-
Climate change implications for the distribution of the babesiosis and anaplasmosis tick vector, Rhipicephalus (Boophilus) microplus
Veterinary Research (2020)