Abstract
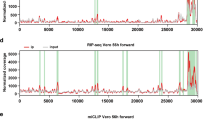
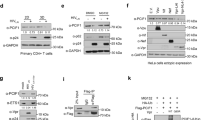
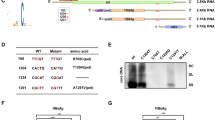
N6-methyladenosine (m6A) is the most prevalent internal modification of eukaryotic mRNA. Very little is known of the function of m6A in the immune system or its role in host–pathogen interactions. Here, we investigate the topology, dynamics and bidirectional influences of the viral–host RNA methylomes during HIV-1 infection of human CD4 T cells. We show that viral infection triggers a massive increase in m6A in both host and viral mRNAs. In HIV-1 mRNA, we identified 14 methylation peaks in coding and noncoding regions, splicing junctions and splicing regulatory sequences. We also identified a set of 56 human gene transcripts that were uniquely methylated in HIV-1-infected T cells and were enriched for functions in viral gene expression. The functional relevance of m6A for viral replication was demonstrated by silencing of the m6A writer or the eraser enzymes, which decreased or increased HIV-1 replication, respectively. Furthermore, methylation of two conserved adenosines in the stem loop II region of HIV-1 Rev response element (RRE) RNA enhanced binding of HIV-1 Rev protein to the RRE in vivo and influenced nuclear export of RNA. Our results identify a new mechanism for the control of HIV-1 replication and its interaction with the host immune system.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Sharp, P. A. The centrality of RNA. Cell 136, 577–580 (2009).
Squires, J. E. et al. Widespread occurrence of 5-methylcytosine in human coding and non-coding RNA. Nucleic Acids Res. 40, 5023–5033 (2012).
Yi, C. & Pan, T. Cellular dynamics of RNA modification. Acc. Chem. Res. 44, 1380–1388 (2011).
Desrosiers, R. C., Friderici, K. H. & Rottman, F. M. Characterization of Novikoff hepatoma mRNA methylation and heterogeneity in the methylated 5′ terminus. Biochemistry 14, 4367–4374 (1975).
Perry, R. P., Kelley, D. E., Friderici, K. & Rottman, F. The methylated constituents of L cell messenger RNA: evidence for an unusual cluster at the 5′ terminus. Cell 4, 387–394 (1975).
Schibler, U., Kelley, D. E. & Perry, R. P. Comparison of methylated sequences in messenger RNA and heterogeneous nuclear RNA from mouse L cells. J. Mol. Biol. 115, 695–714 (1977).
Wei, C. M., Gershowitz, A. & Moss, B. Methylated nucleotides block 5′ terminus of HeLa cell messenger RNA. Cell 4, 379–386 (1975).
Meyer, K. D. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3′ UTRs and near stop codons. Cell 149, 1635–1646 (2012).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012).
Schwartz, S. et al. Perturbation of m6A writers reveals two distinct classes of mRNA methylation at internal and 5′ sites. Cell Rep. 8, 284–296 (2014).
Frayling, T. M. et al. A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science 316, 889–894 (2007).
Jia, G. et al. N6-methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nature Chem. Biol. 7, 885–887 (2011).
Zheng, G. et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol. Cell 49, 18–29 (2013).
Bokar, J. A., Shambaugh, M. E., Polayes, D., Matera, A. G. & Rottman, F. M. Purification and cDNA cloning of the AdoMet-binding subunit of the human mRNA (N6-adenosine)-methyltransferase. RNA 3, 1233–1247 (1997).
Ping, X. L. et al. Mammalian WTAP is a regulatory subunit of the RNA N6-methyladenosine methyltransferase. Cell Res. 24, 177–189 (2014).
Liu, J. et al. A METTL3–METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nature Chem. Biol. 10, 93–95 (2014).
Jia, G., Fu, Y. & He, C. Reversible RNA adenosine methylation in biological regulation. Trends Genet. 29, 108–115 (2013).
Lee, M., Kim, B. & Kim, V. N. Emerging roles of RNA modification: m6A and U-tail. Cell 158, 980–987 (2014).
Wang, X. et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505, 117–120 (2014).
Xu, C. et al. Structural basis for selective binding of m6A RNA by the YTHDC1 YTH domain. Nature Chem. Biol. 10, 927–929 (2014).
Geula, S. et al. m6A mRNA methylation facilitates resolution of naive pluripotency toward differentiation. Science 347, 1002–1006 (2015).
Liu, N. et al. N6-methyladenosine-dependent RNA structural switches regulate RNA-protein interactions. Nature 518, 560–564 (2015).
Zhou, J. et al. Dynamic m6A mRNA methylation directs translational control of heat shock response. Nature 526, 591–594 (2015).
Zhao, X. et al. FTO-dependent demethylation of N6-methyladenosine regulates mRNA splicing and is required for adipogenesis. Cell Res. 24, 1403–1419 (2014).
Fustin, J. M. et al. RNA-methylation-dependent RNA processing controls the speed of the circadian clock. Cell 155, 793–806 (2013).
Batista, P. J. et al. m6A RNA modification controls cell fate transition in mammalian embryonic stem cells. Cell Stem Cell 15, 707–719 (2014).
Wang, Y. et al. N6-methyladenosine modification destabilizes developmental regulators in embryonic stem cells. Nature Cell Biol. 16, 191–198 (2014).
Kane, S. E. & Beemon, K. Precise localization of m6A in Rous sarcoma virus RNA reveals clustering of methylation sites: implications for RNA processing. Mol. Cell. Biol. 5, 2298–2306 (1985).
Krug, R. M., Morgan, M. A. & Shatkin, A. J. Influenza viral mRNA contains internal N6-methyladenosine and 5′-terminal 7-methylguanosine in cap structures. J. Virol. 20, 45–53 (1976).
Finkel, D. & Groner, Y. Methylations of adenosine residues (m6A) in pre-mRNA are important for formation of late simian virus 40 mRNAs. Virology 131, 409–425 (1983).
Fu, L. et al. Simultaneous quantification of methylated cytidine and adenosine in cellular and tissue RNA by nano-flow liquid chromatography–tandem mass spectrometry coupled with the stable isotope-dilution method. Anal. Chem. 87, 7653–7659 (2015).
Heaphy, S. et al. HIV-1 regulator of virion expression (Rev) protein binds to an RNA stem-loop structure located within the Rev response element region. Cell 60, 685–693 (1990).
Kjems, J., Brown, M., Chang, D. D. & Sharp, P. A. Structural analysis of the interaction between the human immunodeficiency virus Rev protein and the Rev response element. Proc. Natl Acad. Sci. USA 88, 683–687 (1991).
Malim, M. H. et al. HIV-1 structural gene expression requires binding of the Rev trans-activator to its RNA target sequence. Cell 60, 675–683 (1990).
Battiste, J. L. et al. Alpha helix-RNA major groove recognition in an HIV-1 rev peptide–RRE RNA complex. Science 273, 1547–1551 (1996).
Harcourt, E. M., Ehrenschwender, T., Batista, P. J., Chang, H. Y. & Kool, E. T. Identification of a selective polymerase enables detection of N6-methyladenosine in RNA. J. Am. Chem. Soc. 135, 19079–19082 (2013).
Heaphy, S., Finch, J. T., Gait, M. J., Karn, J. & Singh, M. Human immunodeficiency virus type 1 regulator of virion expression, rev, forms nucleoprotein filaments after binding to a purine-rich ‘bubble’ located within the rev-responsive region of viral mRNAs. Proc. Natl Acad. Sci. USA 88, 7366–7370 (1991).
Tiley, L. S., Malim, M. H., Tewary, H. K., Stockley, P. G. & Cullen, B. R. Identification of a high-affinity RNA-binding site for the human immunodeficiency virus type 1 Rev protein. Proc. Natl Acad. Sci. USA 89, 758–762 (1992).
Kjems, J., Calnan, B. J., Frankel, A. D. & Sharp, P. A. Specific binding of a basic peptide from HIV-1 Rev. EMBO J. 11, 1119–1129 (1992).
Hammerschmid, M. et al. Scanning mutagenesis of the arginine-rich region of the human immunodeficiency virus type 1 Rev trans activator. J. Virol. 68, 7329–7335 (1994).
Alarcon, C. R. et al. HNRNPA2B1 is a mediator of m6A-dependent nuclear RNA processing events. Cell 162, 1299–1308 (2015).
Roost, C. et al. Structure and thermodynamics of N-methyladenosine in RNA: a spring-loaded base modification. J. Am. Chem. Soc. 137, 2107–2115 (2015).
Spitale, R. C. et al. Structural imprints in vivo decode RNA regulatory mechanisms. Nature 519, 486–490 (2015).
Saletore, Y. et al. The birth of the Epitranscriptome: deciphering the function of RNA modifications. Genome Biol. 13, 175 (2012).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Kalland, K. H., Szilvay, A. M., Langhoff, E. & Haukenes, G. Subcellular distribution of human immunodeficiency virus type 1 Rev and colocalization of Rev with RNA splicing factors in a speckled pattern in the nucleoplasm. J. Virol. 68, 1475–1485 (1994).
Acknowledgements
The authors thank S. Head and the staff of the Next Generation Sequencing core facility at the Scripps Research Institute for help with the early phase of the project and HT-seq. The authors thank J. Klabis for help with the preparation of figures, and members of the Rana laboratory for discussions and advice, especially T.-C. Chao and K. Chang. The authors thank the Sloan Foundation (2015-13964), the Bert L. and N. Kuggie Vallee Foundation, the Irma T. Hirschl and Monique Weill-Caulier Charitable Trusts, the WorldQuant Foundation and the STARR Consortium (I7-A765, I9-A9-071) and acknowledge support from the National Institutes of Health (EB020393, NS076465, C.E.M.; NIAID and NIDA AI43198, DA039562 and DA030199, T.M.R.).
Author information
Authors and Affiliations
Contributions
G.L. designed and performed experiments, analysed data and wrote the paper. S.G. performed experiments and analysed data. Y.S. performed experiments and analysed data. G.M.G. performed experiments and analysed data. V.B., Y.W. and C.M. analysed data. T.M.R. contributed to the concept and design, data analysis and manuscript writing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Methods, Figures 1–6 and Tables 1–4. (PDF 693 kb)
Rights and permissions
About this article
Cite this article
Lichinchi, G., Gao, S., Saletore, Y. et al. Dynamics of the human and viral m6A RNA methylomes during HIV-1 infection of T cells. Nat Microbiol 1, 16011 (2016). https://doi.org/10.1038/nmicrobiol.2016.11
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2016.11
This article is cited by
-
A bibliometric analysis of m6A methylation in viral infection from 2000 to 2022
Virology Journal (2024)
-
N6-methyladenosine-induced miR-182-5p promotes multiple myeloma tumorigenesis by regulating CAMK2N1
Molecular and Cellular Biochemistry (2024)
-
N6-methyladenosine modification—a key player in viral infection
Cellular & Molecular Biology Letters (2023)
-
Identification and validation of m6A RNA regulatory network in pulpitis
BMC Oral Health (2023)
-
RNA modification: mechanisms and therapeutic targets
Molecular Biomedicine (2023)