Abstract
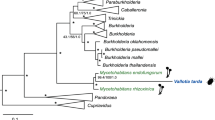
Diverse organisms are associated with obligate microbial mutualists. How such essential symbionts have originated from free-living ancestors is of evolutionary interest. Here we report that, in natural populations of the stinkbug Plautia stali, obligate bacterial mutualists are evolving from environmental bacteria. Of six distinct bacterial lineages associated with insect populations, two are uncultivable with reduced genomes, four are cultivable with non-reduced genomes, one uncultivable symbiont is fixed in temperate populations, and the other uncultivable symbiont coexists with four cultivable symbionts in subtropical populations. Symbiont elimination resulted in host mortality for all symbionts, while re-infection with any of the symbionts restored normal host growth, indicating that all the symbionts are indispensable and almost equivalent functionally. Some aseptic newborns incubated with environmental soils acquired the cultivable symbionts and normal growth was restored, identifying them as environmental Pantoea spp. Our finding uncovers an evolutionary transition from a free-living lifestyle to obligate mutualism that is currently ongoing in nature.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Douglas, A. E. The Symbiotic Habit (Princeton Univ. Press, 2010).
Archibald, J. One Plus One Equals One: Symbiosis and the Evolution of Complex Life (Oxford Univ. Press, 2014).
Buchner, P. Endosymbiosis of Animals with Plant Microorganisms (Interscience Publisher, 1965).
Bourtzis, K. & Miller, T. A. Insect Symbiosis (CRC Press, 2003).
Moran, N. A., McCutcheon, J. P. & Nakabachi, A. Genomics and evolution of heritable bacterial symbionts. Annu. Rev. Genet. 42, 165–190 (2008).
Shigenobu, S., Watanabe, H., Hattori, M., Sakaki, Y. & Ishikawa, H. Genome sequence of the endocellular bacterial symbiont of aphids Buchnera sp. APS. Nature 407, 81–86 (2000).
Akman, L. et al. Genome sequence of the endocellular obligate symbiont of tsetse flies, Wigglesworthia glossinidia. Nature Genet. 32, 402–407 (2002).
Nikoh, N., Hosokawa, T., Oshima, K., Hattori, M. & Fukatsu, T. Reductive evolution of bacterial genome in insect gut environment. Genome Biol. Evol. 3, 702–714 (2011).
Kaiwa, N. et al. Symbiont-supplemented maternal investment underpinning host's ecological adaptation. Curr. Biol. 24, 2465–2470 (2014).
Wernegreen, J. J. Genome evolution in bacterial endosymbionts of insects. Nature Rev. Genet. 3, 850–861 (2002).
McCutcheon, J. P. & Moran, N. A. Extreme genome reduction in symbiotic bacteria. Nature Rev. Microbiol. 10, 13–26 (2011).
Schaefer, C. W. & Panizzi, A. R. Heteroptera of Economic Importance (CRC, 2000).
Abe, Y., Mishiro, K. & Takanashi, M. Symbiont of brown-winged green bug, Plautia stali Scott. Jpn J. Appl. Entomol. Zool. 39, 109–115 (1995).
Prado, S. S. & Almeida, R. P. P. Phylogenetic placement of pentatomid stink bug gut symbionts. Curr. Microbiol. 58, 64–69 (2009).
Kobayashi, H., Kawasaki, K., Takeishi, K. & Noda, H. Symbiont of the stink bug Plautia stali synthesizes rough-type lipopolysaccharide. Microbiol. Res. 167, 48–54 (2011).
Kikuchi, Y., Hosokawa, T. & Fukatsu, T. in Microbial Ecology Research Trends (ed. Dijk, T. V. ) 39–63 (Nova Science, 2008).
Brady, C. et al. Phylogeny and identification of Pantoea species associated with plants, humans and the natural environment based on multilocus sequence analysis (MLSA). Syst. Appl. Microbiol. 31, 447–460 (2008).
Kikuchi, Y., Hosokawa, T. & Fukatsu, T. An ancient but promiscuous host–symbiont association between Burkholderia gut symbionts and their heteropteran hosts. ISME J. 5, 446–460 (2011).
Hosokawa, T., Kikuchi, Y., Nikoh, N. & Fukatsu, T. Polyphyly of gut symbionts in stinkbugs of the family Cydnidae. Appl. Environ. Microbiol. 78, 4758–4761 (2012).
Bansal, R., Michel, A. P. & Sabree, Z. L. The crypt-dwelling primary bacterial symbiont of the polyphagous pentatomid pest Halyomorpha halys (Hemiptera: Pentatomidae). Environ. Entomol. 43, 617–625 (2014).
Bistolas, K. S. I., Sakamoto, R. I., Fernandes, J. A. M. & Goffredi, S. K. Symbiont polyphyly, co-evolution, and necessity in pentatomid stinkbugs from Costa Rica. Front. Microbiol. 5, 349 (2014).
Grant, P. R. & Grant, B. R. Unpredictable evolution in a 30-year study of Darwin's finches. Science 296, 707–711 (2002).
Salem, H. et al. Vitamin supplementation by gut symbionts ensures metabolic homeostasis in an insect host. Proc. R. Soc. Lond. B 282, 1803 (2014).
Kado, C. I. Erwinia and related genera. Prokaryotes 6, 443–450 (2006).
Russell, J. A. & Moran, N. A. Costs and benefits of symbiont infection in aphids: variation among symbionts and across temperatures. Proc. R. Soc. Lond. B 273, 603–610 (2006).
Dunbar, H. E., Wilson, A. C. C., Ferguson, N. R. & Moran, N. A. Aphid thermal tolerance is governed by a point mutation in bacterial symbionts. PLoS Biol. 5, e96 (2007).
Tsuchida, T., Koga, R. & Fukatsu, T. Host plant specialization governed by facultative symbiont. Science 303, 1989 (2004).
Hosokawa, T., Kikuchi, Y., Shimada, M. & Fukatsu, T. Obligate symbiont involved in pest status of host insect. Proc. R. Soc. Lond. B 274, 1979–1984 (2007).
Clayton, A. L. et al. A novel human-infection-derived bacterium provides insights into the evolutionary origins of mutualistic insect–bacterial symbioses. PLoS Pathog. 8, e1002990 (2012).
Sachs, J. L., Skophammer, R. G., Bansal, N. & Stajich, J. E. Evolutionary origins and diversification of proteobacterial mutualists. Proc. R. Soc. Lond. B 281, 20132146 (2014).
Mau, R. F. L. & Mitchell, W. C. Development and reproduction of the oriental stink bug, Plautia stali (Hemiptera: Pentatomidae). Ann. Entomol. Soc. Am. 71, 756–757 (1978).
Hosokawa, T., Kikuchi, Y., Nikoh, N., Shimada, M. & Fukatsu, T. Strict host–symbiont cospeciation and reductive genome evolution in insect gut bacteria. PLoS Biol. 4, e337 (2006).
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013).
Delcher, A. L., Bratke, K. A., Edwin, C., Powers, E. C. & Salzberg, S. L. Identifying bacterial genes and endosymbiont DNA with Glimmer. Bioinformatics 23, 673–679 (2007).
Acknowledgements
The authors thank M. Baba, N. Baba, Y. G. Baba, D. Haraguchi, H. Hirayama, M. Hironaka, S. Kada, N. Kaiwa, Y. Kikuchi, K. Kizaki, S. Kudo, T. Makino, M. Moriyama, N. Nagata, S. Ohno, T. Ohtani, M. Ono, M. Sakakibara, H. Toju, K. Tsuji, N. Tsurusaki, T. Uesato, R. Ukuda, H. Watanabe and T. Yamaguchi for insect samples; J. Makino, N. Tanifuji, U. Asaga and W. Kikuchi for technical assistance; J. P. McCutcheon for comments on the manuscript; and Y. Nakajima and Y. Kikuchi for logistic and technical support. This study was supported by a JSPS KAKENHI grant (25221107) to T.F. and by the University of the Ryukyus Foundation to T.H., and Y.I. was supported by a JSPS Fellowship for Young Scientists.
Author information
Authors and Affiliations
Contributions
T.H. and T.F. designed the study. T.H. performed most experiments and data analyses. Y.I. conducted some symbiont-replacing experiments. N.N. analysed the symbiont genome data. M.F. and N.S. determined the draft genome sequences of the symbionts. T.H. and T.F. wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1–8 and Tables 1–5. (PDF 7326 kb)
Rights and permissions
About this article
Cite this article
Hosokawa, T., Ishii, Y., Nikoh, N. et al. Obligate bacterial mutualists evolving from environmental bacteria in natural insect populations. Nat Microbiol 1, 15011 (2016). https://doi.org/10.1038/nmicrobiol.2015.11
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2015.11
This article is cited by
-
Mixta mediterraneensis as a novel and abundant gut symbiont of the allergen-producing domestic mite Blomia tropicalis
Experimental and Applied Acarology (2024)
-
Obligate Gut Symbiotic Association with Caballeronia in the Mulberry Seed Bug Paradieuches dissimilis (Lygaeoidea: Rhyparochromidae)
Microbial Ecology (2023)
-
Structural remodeling of midgut symbiotic organ and altered food flow upon metamorphosis of the stinkbug Plautia stali
Applied Entomology and Zoology (2023)
-
Comparative seasonal analysis of Eri silkworm (Samia ricini Donovan) gut composition: implications for lignocellulose degradation
Environmental Science and Pollution Research (2023)
-
Host’s genetic background determines the outcome of reciprocal faecal transplantation on life-history traits and microbiome composition
Animal Microbiome (2022)