Abstract
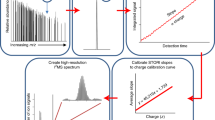
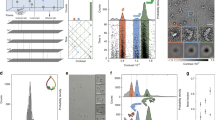
We have used native mass spectrometry to analyze macromolecular complexes involved in the chaperonin-assisted refolding of gp23, the major capsid protein of bacteriophage T4. Adapting the instrumental methods allowed us to monitor all intermediate complexes involved in the chaperonin folding cycle. We found that GroEL can bind up to two unfolded gp23 substrate molecules. Notably, when GroEL is in complex with the cochaperonin gp31, it binds exclusively one gp23. We also demonstrated that the folding and assembly of gp23 into 336-kDa hexamers by GroEL-gp31 can be monitored directly by electrospray ionization mass spectrometry (ESI-MS). These data reinforce the great potential of ESI-MS as a technique to investigate structure-function relationships of protein assemblies in general and the chaperonin-protein folding machinery in particular. A major advantage of native mass spectrometry is that, given sufficient resolution, it allows the analysis at the picomole level of sensitivity of heterogeneous protein complexes with molecular masses up to several million daltons.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Houry, W.A., Frishman, D., Eckerskorn, C., Lottspeich, F. & Hartl, F.U. Identification of in vivo substrates of the chaperonin GroEL. Nature 402, 147–154 (1999).
Hartl, F.U. & Hayer-Hartl, M. Molecular chaperones in the cytosol: from nascent chain to folded protein. Science 295, 1852–1858 (2002).
Ellis, R.J. Molecular chaperones: inside and outside the Anfinsen cage. Curr. Biol. 11, R1038–R1040 (2001).
Sigler, P.B. et al. Structure and function in GroEL-mediated protein folding. Annu. Rev. Biochem. 67, 581–608 (1998).
Walters, C., Errington, N., Rowe, A.J. & Harding, S.E. Hydrolysable ATP is a requirement for the correct interaction of molecular chaperonins cpn60 and cpn10. Biochem. J. 364, 849–855 (2002).
van der Vies, S.M., Viitanen, P.V., Gatenby, A.A., Lorimer, G.H. & Jaenicke, R. Conformational states of ribulosebisphosphate carboxylase and their interaction with chaperonin 60. Biochemistry 31, 3635–3644 (1992).
Hunt, J.F., van der Vies, S.M., Henry, L. & Deisenhofer, J. Structural adaptations in the specialized bacteriophage T4 cochaperonin Gp31 expand the size of the Anfinsen cage. Cell 90, 361–371 (1997).
Langer, T., Pfeifer, G., Martin, J., Baumeister, W. & Hartl, F.U. Chaperonin-mediated protein folding: GroES binds to one end of the GroEL cylinder, which accommodates the protein substrate within its central cavity. EMBO J. 11, 4757–4765 (1992).
Braig, K., Simon, M., Furuya, F., Hainfeld, J.F. & Horwich, A.L. A polypeptide bound by the chaperonin GroEL is localized within a central cavity. Proc. Natl. Acad. Sci. USA 90, 3978–3982 (1993).
Chen, S. et al. Location of a folding protein and shape changes in GroEL-GroES complexes imaged by cryo-electron microscopy. Nature 371, 261–264 (1994).
Braig, K. et al. The crystal structure of the bacterial chaperonin GroEL at 2.8 Å. Nature 371, 578–586 (1994).
Fiaux, J., Bertelsen, E.B., Horwich, A.L. & Wuthrich, K. NMR analysis of a 900K GroEL GroES complex. Nature 418, 207–211 (2002).
Riek, R., Fiaux, J., Bertelsen, E.B., Horwich, A.L. & Wuthrich, K. Solution NMR techniques for large molecular and supramolecular structures. J. Am. Chem. Soc. 124, 12144–12153 (2002).
Griswold, I.J. & Dahlquist, F.W. Bigger is better: megadalton protein NMR in solution. Nat. Struct. Biol. 9, 567–568 (2002).
van den Heuvel, R.H. & Heck, A.J. Native protein mass spectrometry: from intact oligomers to functional machineries. Curr. Opin. Chem. Biol. 8, 519–526 (2004).
Verentchikov, A.N., Ens, W. & Standing, K.G. Reflecting time-of-flight mass spectrometer with an electrospray ion source and orthogonal extraction. Anal. Chem. 66, 126–133 (1994).
Loo, J.A. Studying noncovalent protein complexes by electrospray ionization mass spectrometry. Mass Spectrom. Rev. 16, 1–23 (1997).
Heck, A.J. & Van Den Heuvel, R.H. Investigation of intact protein complexes by mass spectrometry. Mass Spectrom. Rev. 23, 368–389 (2004).
Robinson, C.V. Protein complexes take flight. Nat. Struct. Biol. 9, 505–506 (2002).
Heuvel, R.H. & Heck, A.J. Native protein mass spectrometry: from intact oligomers to functional machineries. Curr. Opin. Chem. Biol. 8, 519–526 (2004).
Hernandez, H. & Robinson, C.V. Dynamic protein complexes: insights from mass spectrometry. J. Biol. Chem. 276, 46685–46688 (2001).
Robinson, C.V. et al. Conformation of GroEL-bound α-lactalbumin probed by mass spectrometry. Nature 372, 646–651 (1994).
Coyle, J.E., Jaeger, J., Gross, M., Robinson, C.V. & Radford, S.E. Structural and mechanistic consequences of polypeptide binding by GroEL. Fold. Des. 2, R93–104 (1997).
Coyle, J.E. et al. GroEL accelerates the refolding of hen lysozyme without changing its folding mechanism. Nat. Struct. Biol. 6, 683–690 (1999).
Rostom, A.A. & Robinson, C.V. Detection of the intact GroEL chaperonin assembly by mass spectrometry. J. Am. Chem. Soc. 121, 4718–4719 (1999).
Krutchinsky, A.N., Chernushevich, I.V., Spicer, V.L., Ens, W. & Standing, K.G. Collisional damping interface for an electrospray ionization time-of-flight mass spectrometer. J. Am. Soc. Mass Spectrom. 9, 569–579 (1998).
Sobott, F., Hernandez, H., McCammon, M.G., Tito, M.A. & Robinson, C.V. A tandem mass spectrometer for improved transmission and analysis of large macromolecular assemblies. Anal. Chem. 74, 1402–1407 (2002).
Tahallah, N., Pinkse, M., Maier, C.S. & Heck, A.J. The effect of the source pressure on the abundance of ions of noncovalent protein assemblies in an electrospray ionization orthogonal time-of-flight instrument. Rapid Commun. Mass Spectrom. 15, 596–601 (2001).
Chernushevich, I.V. & Thomson, B.A. Collisional cooling of large ions in electrospray mass spectrometry. Anal. Chem. 76, 1754–1760 (2004).
Jackson, G.S. et al. Binding and hydrolysis of nucleotides in the chaperonin catalytic cycle: implications for the mechanism of assisted protein folding. Biochemistry 32, 2554–2563 (1993).
Burston, S.G., Ranson, N.A. & Clarke, A.R. The origins and consequences of asymmetry in the chaperonin reaction cycle. J. Mol. Biol. 249, 138–152 (1995).
Laemmli, U.K., Beguin, F. & Gujer-Kellenberger, G. A factor preventing the major head protein of bacteriophage T4 from random aggregation. J. Mol. Biol. 47, 69–85 (1970).
van der Vies, S.M., Gatenby, A.A. & Georgopoulos, C. Bacteriophage T4 encodes a cochaperonin that can substitute for Escherichia coli GroES in protein folding. Nature 368, 654–656 (1994).
Hartl, F.U. Molecular chaperones in cellular protein folding. Nature 381, 571–579 (1996).
Bukau, B. & Horwich, A.L. The Hsp70 and Hsp60 chaperone machines. Cell 92, 351–366 (1998).
Bakkes, P.J., Faber, B.W., van Heerikhuizen, H. & van der Vies, S.M. The T4-encoded cochaperonin, gp31, has unique properties that explain its requirement for the folding of the T4 major capsid protein. Proc. Natl. Acad. Sci. USA (in the press).
Xu, Z., Horwich, A.L. & Sigler, P.B. The crystal structure of the asymmetric GroEL-GroES-(ADP)7 chaperonin complex. Nature 388, 741–750 (1997).
Koonin, E.V. & van der Vies, S.M. Conserved sequence motifs in bacterial and bacteriophage chaperonins. Trends Biochem. Sci. 20, 14–15 (1995).
Acknowledgements
We thank E. Kroezinga and J. Hendriks, who contributed to some of the work reported in this article. The research was supported by Fundamenteel Onderzoek der Materie, project number 01FB12.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Various ratios of GroEL-gp23 complexes analyzed by gel filtration (PDF 156 kb)
Supplementary Methods
Protein purification (PDF 75 kb)
Rights and permissions
About this article
Cite this article
van Duijn, E., Bakkes, P., Heeren, R. et al. Monitoring macromolecular complexes involved in the chaperonin-assisted protein folding cycle by mass spectrometry. Nat Methods 2, 371–376 (2005). https://doi.org/10.1038/nmeth753
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth753
This article is cited by
-
Mass spectrometry using electrospray ionization
Nature Reviews Methods Primers (2023)
-
Distinct Stabilities of the Structurally Homologous Heptameric Co-Chaperonins GroES and gp31
Journal of the American Society for Mass Spectrometry (2019)
-
Ion concentration in micro and nanoscale electrospray emitters
Analytical and Bioanalytical Chemistry (2018)
-
Elucidation of salt-tolerance metabolic pathways in contrasting rice genotypes and their segregating progenies
Plant Cell Reports (2016)
-
Sizing up large protein complexes by electrospray ionisation-based electrophoretic mobility and native mass spectrometry: morphology selective binding of Fabs to hepatitis B virus capsids
Analytical and Bioanalytical Chemistry (2014)