Abstract
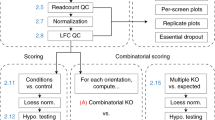
We describe a statistical analysis methodology designed to minimize the impact of off-target activities upon large-scale RNA interference (RNAi) screens in mammalian cells. Application of this approach enhances reconfirmation rates and facilitates the experimental validation of new gene activities through the probability-based identification of multiple distinct and active small interfering RNAs (siRNAs) targeting the same gene. We further extend this approach to establish that the optimal redundancy for efficacious RNAi collections is between 4–6 siRNAs per gene.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Berns, K. et al. Nature 428, 431–437 (2004).
Boutros, M. et al. Science 303, 832–835 (2004).
Brummelkamp, T.R. et al. Nat. Chem. Biol. 2, 202–206 (2006).
Moffat, J. et al. Cell 124, 1283–1298 (2006).
Paddison, P.J. et al. Nature 428, 427–431 (2004).
Sonnichsen, B. et al. Nature 434, 462–469 (2005).
Westbrook, T.F. et al. Cell 121, 837–848 (2005).
Birmingham, A. et al. Nat. Methods 3, 199–204 (2006).
Jackson, A.L. et al. Nat. Biotechnol. 21, 635–637 (2003).
Echeverri, C.J. et al. Nat. Methods 3, 777–779 (2006).
Kittler, R. et al. Nature 432, 1036–1040 (2004).
Yan, S.F., Asatryan, H., Li, J. & Zhou, Y. J. Chem. Inf. Model. 45, 1784–1790 (2005).
Fay, N. & Ullmann, D. Drug Discov. Today 7, S181–S186 (2002).
Bajorath, J. Nat. Rev. Drug Discov. 1, 882–894 (2002).
Shedden, K. et al. BMC Bioinformatics 6, 26 (2005).
Acknowledgements
We thank L. Miraglia for helpful discussions and oversight of screens, J. Zhang for excellent technical assistance, S. Batalov (Genomics Institute of the Novartis Research Foundation) and P. Aza-Blanc (Burnham Institute) for the identification of negative control siRNA sequences, E. Lader (Qiagen) for facilitating collaboration, D. Elleder (Salk Institute) for providing the MLV supernatant, N.R. Landau (New York University, School of Medicine) for providing pNL43-luc-r+e−, and N. Somia (University of Minnesota) for the gift of pCMVgp. R-language implementation of the RSA algorithm was provided by B. Zhou (Genomics Institute of the Novartis Research Foundation). This work was supported by the Novartis Research Foundation and a grant from the US National Institutes of Health (1 R01 AI072645-01).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
T.B. and U.K. are employees of Qiagen GmbH.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–2, Supplementary Methods (PDF 3262 kb)
Supplementary Table 1
Original screen data for screen A. (XLS 12915 kb)
Supplementary Table 2
Original screen data for screen B. (XLS 12910 kb)
Supplementary Table 3
Scrambled Control Sequences. (XLS 17 kb)
Supplementary Table 4
Reconfirmation screen data for screen A. (XLS 53 kb)
Supplementary Table 5
Reconfirmation screen data for screen B. (XLS 60 kb)
Rights and permissions
About this article
Cite this article
König, R., Chiang, Cy., Tu, B. et al. A probability-based approach for the analysis of large-scale RNAi screens. Nat Methods 4, 847–849 (2007). https://doi.org/10.1038/nmeth1089
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth1089
This article is cited by
-
Uncovering hidden cancer self-dependencies through analysis of shRNA-level dependency scores
Scientific Reports (2024)
-
INPP5A phosphatase is a synthetic lethal target in GNAQ and GNA11-mutant melanomas
Nature Cancer (2024)
-
Mitochondrial E3 ubiquitin ligase MARCHF5 controls BAK apoptotic activity independently of BH3-only proteins
Cell Death & Differentiation (2023)
-
Targeting Menin disrupts the KMT2A/B and polycomb balance to paradoxically activate bivalent genes
Nature Cell Biology (2023)
-
Combining CRISPRi and metabolomics for functional annotation of compound libraries
Nature Chemical Biology (2022)