Abstract
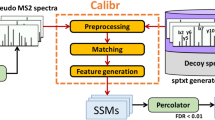
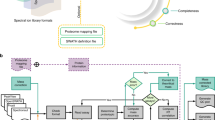
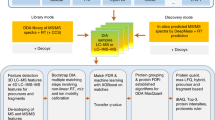
Mass spectrometry with data-independent acquisition (DIA) is a promising method to improve the comprehensiveness and reproducibility of targeted and discovery proteomics, in theory by systematically measuring all peptide precursors in a biological sample. However, the analytical challenges involved in discriminating between peptides with similar sequences in convoluted spectra have limited its applicability in important cases, such as the detection of single-nucleotide polymorphisms (SNPs) and alternative site localizations in phosphoproteomics data. We report Specter (https://github.com/rpeckner-broad/Specter), an open-source software tool that uses linear algebra to deconvolute DIA mixture spectra directly through comparison to a spectral library, thus circumventing the problems associated with typical fragment-correlation-based approaches. We validate the sensitivity of Specter and its performance relative to that of other methods, and show that Specter is able to successfully analyze cases involving highly similar peptides that are typically challenging for DIA analysis methods.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Tabb, D.L. et al. Repeatability and reproducibility in proteomic identifications by liquid chromatography-tandem mass spectrometry. J. Proteome Res. 9, 761–776 (2010).
Picotti, P. & Aebersold, R. Selected reaction monitoring–based proteomics: workflows, potential, pitfalls and future directions. Nat. Methods 9, 555–566 (2012).
Röst, H.L. et al. OpenSWATH enables automated, targeted analysis of data-independent acquisition MS data. Nat. Biotechnol. 32, 219–223 (2014).
Tsou, C.-C. et al. DIA-Umpire: comprehensive computational framework for data-independent acquisition proteomics. Nat. Methods 12, 258–264 (2015).
Wang, J. et al. MSPLIT-DIA: sensitive peptide identification for data-independent acquisition. Nat. Methods 12, 1106–1108 (2015).
Bilbao, A. et al. Ranking fragment ions based on outlier detection for improved label-free quantification in data-independent acquisition LC-MS/MS. J. Proteome Res. 14, 4581–4593 (2015).
Bruderer, R. et al. Extending the limits of quantitative proteome profiling with data-independent acquisition and application to acetaminophen-treated three-dimensional liver microtissues. Mol. Cell. Proteomics 14, 1400–1410 (2015).
Bern, M. et al. Deconvolution of mixture spectra from ion-trap data-independent-acquisition tandem mass spectrometry. Anal. Chem. 82, 833–841 (2010).
Navarro, P. et al. A multicenter study benchmarks software tools for label-free proteome quantification. Nat. Biotechnol. 34, 1130–1136 (2016).
Schulz-Trieglaff, O. et al. Statistical quality assessment and outlier detection for liquid chromatography-mass spectrometry experiments. BioData Min. 2, 4 (2009).
Frewen, B.E., Merrihew, G.E., Wu, C.C., Noble, W.S. & MacCoss, M.J. Analysis of peptide MS/MS spectra from large-scale proteomics experiments using spectrum libraries. Anal. Chem. 78, 5678–5684 (2006).
Ritter, G.L., Lowry, S.R., Isenhour, T.L. & Wilkins, C.L. Factor analysis of the mass spectra of mixtures. Anal. Chem. 48, 591–595 (1976).
Likic´, V.A. Extraction of pure components from overlapped signals in gas chromatography-mass spectrometry (GC-MS). BioData Min. 2, 6 (2009).
Nikolskiy, I., Mahieu, N.G., Chen, Y.-J., Tautenhahn, R. & Patti, G.J. An untargeted metabolomic workflow to improve structural characterization of metabolites. Anal. Chem. 85, 7713–7719 (2013).
Du, X. & Zeisel, S.H. Spectral deconvolution for gas chromatography mass spectrometry-based metabolomics: current status and future perspectives. Comput. Struct. Biotechnol. J. 4, e201301013 (2013).
Lam, H., Deutsch, E.W. & Aebersold, R. Artificial decoy spectral libraries for false discovery rate estimation in spectral library searching in proteomics. J. Proteome Res. 9, 605–610 (2010).
Cheng, C.-Y., Tsai, C.-F., Chen, Y.-J., Sung, T.-Y. & Hsu, W.-L. Spectrum-based method to generate good decoy libraries for spectral library searching in peptide identifications. J. Proteome Res. 12, 2305–2310 (2013).
Du, X. et al. Linear discriminant analysis-based estimation of the false discovery rate for phosphopeptide identifications. J. Proteome Res. 7, 2195–2203 (2008).
Shastry, B.S. SNPs in disease gene mapping, medicinal drug development and evolution. J. Hum. Genet. 52, 871–880 (2007).
Lawrence, R.T., Searle, B.C., Llovet, A. & Villén, J. Plug-and-play analysis of the human phosphoproteome by targeted high-resolution mass spectrometry. Nat. Methods 13, 431–434 (2016).
Rosenberger, G. et al. Inference and quantification of peptidoforms in large sample cohorts by SWATH-MS. Nat. Biotechnol. 35, 781–788 (2017).
Malecz, N., Foisner, R., Stadler, C. & Wiche, G. Identification of plectin as a substrate of p34cdc2 kinase and mapping of a single phosphorylation site. J. Biol. Chem. 271, 8203–8208 (1996).
Bouameur, J.-E. et al. Phosphorylation of serine 4,642 in the C-terminus of plectin by MNK2 and PKA modulates its interaction with intermediate filaments. J. Cell Sci. 126, 4195–4207 (2013).
McQueen, P. et al. Whole cell, label free protein quantitation with data independent acquisition: quantitation at the MS2 level. Proteomics 15, 16–24 (2015).
MacLean, B. et al. Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 26, 966–968 (2010).
Gillet, L.C. et al. Targeted data extraction of the MS/MS spectra generated by data-independent acquisition: a new concept for consistent and accurate proteome analysis. Mol. Cell. Proteomics 11, O111.016717 (2012).
Yang, W. & Zurbenko, I. Kolmogorov-Zurbenko filters. Wiley Interdiscip. Rev. Comput. Stat. 2, 340–351 (2010).
Deutsch, E.W. File formats commonly used in mass spectrometry proteomics. Mol. Cell. Proteomics 11, 1612–1621 (2012).
Frewen, B. & MacCoss, M.J. Using BiblioSpec for creating and searching tandem MS peptide libraries. Curr. Protoc. Bioinformatics 20, 13.7.1–13.7.12 (2007).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906 (2007).
Abelin, J.G. et al. Reduced-representation phosphosignatures measured by quantitative targeted MS capture cellular states and enable large-scale comparison of drug-induced phenotypes. Mol. Cell. Proteomics 15, 1622–1641 (2016).
Vizcaíno, J.A. et al. 2016 update of the PRIDE database and its related tools. Nucleic Acids Res. 44, D447–D456 (2016).
Acknowledgements
This work was supported by the NIH (grant U54 HG008097 to J.D.J.). The authors thank S. Tenzer and L. Gillet for providing details of the data sets for the LFQbench study, and K. Clauser for many insightful discussions.
Author information
Authors and Affiliations
Contributions
R.P. conceived of the methodology, created the software, performed the analyses, and wrote the manuscript. S.A.M. conceived and carried out the experiments involving similar synthetic peptides and DIA-DDA comparison, and helped revise the manuscript. A.S.V.J. carried out the experiments and manual analysis of the synthetic phosphopeptide spike-in experiments. J.D.E. and J.G.A. developed the DIA acquisition method. M.J.M., S.A.C., and J.D.J. provided laboratory resources and guidance on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4, Supplementary Notes 1–6 and Supplementary Tables 1–6 (PDF 4934 kb)
Rights and permissions
About this article
Cite this article
Peckner, R., Myers, S., Jacome, A. et al. Specter: linear deconvolution for targeted analysis of data-independent acquisition mass spectrometry proteomics. Nat Methods 15, 371–378 (2018). https://doi.org/10.1038/nmeth.4643
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4643
This article is cited by
-
Nanoparticle enrichment mass-spectrometry proteomics identifies protein-altering variants for precise pQTL mapping
Nature Communications (2024)
-
SeFilter-DIA: Squeeze-and-Excitation Network for Filtering High-Confidence Peptides of Data-Independent Acquisition Proteomics
Interdisciplinary Sciences: Computational Life Sciences (2024)
-
Prediction of peptide mass spectral libraries with machine learning
Nature Biotechnology (2023)
-
A comprehensive LFQ benchmark dataset on modern day acquisition strategies in proteomics
Scientific Data (2022)
-
CHICKN: extraction of peptide chromatographic elution profiles from large scale mass spectrometry data by means of Wasserstein compressive hierarchical cluster analysis
BMC Bioinformatics (2021)