Abstract
Single-cell RNA sequencing and high-dimensional cytometry can be used to generate detailed trajectories of dynamic biological processes such as differentiation or development. Here we present cellAlign, a quantitative framework for comparing expression dynamics within and between single-cell trajectories. By applying cellAlign to mouse and human embryonic developmental trajectories, we systematically delineate differences in the temporal regulation of gene expression programs that would otherwise be masked.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Trapnell, C. et al. Nat. Biotechnol. 32, 381–386 (2014).
Shin, J. et al. Cell Stem Cell 17, 360–372 (2015).
Haghverdi, L., Büttner, M., Wolf, F.A., Buettner, F. & Theis, F.J. Nat. Methods 13, 845–848 (2016).
Bendall, S.C. et al. Cell 157, 714–725 (2014).
Llorens-Bobadilla, E. et al. Cell Stem Cell 17, 329–340 (2015).
Shalek, A.K. et al. Nature 510, 363–369 (2014).
Yan, L. et al. Nat. Struct. Mol. Biol. 20, 1131–1139 (2013).
Deng, Q. et al. Science 343, 193–196 (2014).
Bouniol, C., Nguyen, E. & Debey, P. Exp. Cell Res. 218, 57–62 (1995).
Braude, P., Bolton, V. & Moore, S. Nature 332, 459–461 (1988).
Giorgino, T. J. Stat. Softw. 31, 1–24 (2009).
Yuan, Y. et al. BMC Bioinformatics 12, 347 (2011).
Kamburov, A., Stelzl, U., Lehrach, H. & Herwig, R. Nucleic Acids Res. 41, 793–800 (2013).
Acknowledgements
This study was supported by the Israel Science Foundation (ISF) grants 1365/12 and the Applebaum Foundation. We thank Y. Ofran, M. Drukker, N. Kaplan, K. Brennand, and members of the Shen-Orr lab for fruitful discussions, and D. Alpert for assistance in algorithmic design and technical support.
Author information
Authors and Affiliations
Contributions
A.A., L.S.M. and S.S.S.-O. designed the algorithm and simulations; A.A. and L.S.M. implemented the algorithm and analyzed data; T.D. generated and helped analyze mouse B cell CyTOF data; and all authors wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
S.S.S.-O. is a scientific adviser and holds equity in CytoReason. All other authors declare no competing interests.
Integrated supplementary information
Supplementary Figure 1 Rescaling of pseudo-time reduces technical impact of sampling density and trajectory building algorithm.
(a) Points distributed along a linear curve in the 200-dimensional space were sampled either uniformly or with bias to increase the density at the trajectory midpoint. Trajectories were built per sampling by Monocle2 algorithm and the resulting pseudo-time scores distributions are shown with color denoting magnitude of sampling bias. (b) Absolute pseudo-time shift elements (n=200 elements) resulting from alignment of the uniformly with the biased-sampled trajectories with (light blue) or without (pink) pseudo-time rescaling. (c) Absolute pseudo-time shift elements (n=200 elements) resulting from alignment of the uniformly sampled trajectory generated by different trajectory building algorithms with (light blue) or without (pink) pseudo-time rescaling. Per boxplot, the center, minimum, maximum, and range denote median, lowest value within 1.5 IQR of lower quartile, highest value within 1.5 IQR of upper quartile, and inter-quartile range, respectively. Abbreviations: DPT: diffusion pseudo time.
Supplementary Figure 2 Alignment-based distance calculated by cellAlign reflects known biological similarities.
Hierarchical clustering of the eight single cell RNA sequencing trajectories analyzed throughout the manuscript using alignment-based distances as calculated by cellAlign. Abbreviations: SVZ: sub-ventricular zone; DC: dendritic cells.
Supplementary Figure 3 Assembly of single-cell trajectories describing gene-expression dynamics following dendritic cells stimulation by LPS and PAM.
Diffusion maps algorithm applied to single cell RNA sequencing data of dendritic cells stimulated with either LPS or PAM. Dots represent single cells after either LPS (a, c) or PAM (b, d) stimulation. Colors denote either the actual collection time post-stimulation (a, b) or scaled pseudo-time scores (c, d) calculated by the diffusion pseudo-time algorithm. Abbreviations: DC: diffusion component.
Supplementary Figure 4 Expression dynamics of different gene-modules under LPS and PAM.
Scaled imputed expression (z-scores) of genes included in the modules used for either global (a, b) or local (c, d) alignment under either LPS (a, c) or PAM (b, d) stimulation. Genes are ordered along the rows, while single cells are ordered by their pseudo-time scores on columns. Collection times and pseudo-time scores per single cell are displayed at the top.
Supplementary Figure 5 Local alignment by cellAlign.
Dissimilarity matrix and local alignment between expression dynamics of a single gene module along LPS (left to right) and PAM (top to bottom) trajectories under similarity threshold of 0.25 of the range of values in the dissimilarity matrix.
Supplementary Figure 6 cellAlign performance under noise and pre-processing methodologies.
(a) The effect of increasing levels of noise in the form of additional dropout events on local alignment. Assessment was performed by calculating the fraction of the originally identified, locally-conserved region along the PAM trajectory that was still identified in the noised data. Error bars and centers denote standard deviations and mean values across n=500 simulations. (b) Performance of cellAlign for data processed by different preprocessing techniques, including: cells subsampling (purple), normalization method (turquoise), trajectory building algorithms (red) and application of data-imputation (green). Spearman correlation of alignment-based distances calculated across gene modules between original and the modified expression data are shown. For cells subsampling, the average and standard deviation across ten simulations are shown as error bars. (c) Spearman correlations between alignment based distances calculated across modules of co-expressed genes using expression data obtained either without batch correction or corrected by Combat or Limma.
Supplementary Figure 7 Validation and performance of cellAlign on non-imputed single-cell RNA sequencing data.
(a-b) Correlation between pseudo-time scores obtained from the normalized data (y-axes) versus those obtained from the imputed data (x-axes) for LPS (a) and PAM (b) trajectories. Scaled interpolated expression of the different 12 gene-modules calculated either using the normalized or imputed data are displayed beside and below the axes, respectively. Names of the different gene-modules, as appear in the original publication, are placed along a diagonal path to the left and below expression dynamics. Spearman correlation values between the imputed and non-imputed expression values appear on top. (c-f) Non-imputed, scaled expression (z-scores) of genes partitioned in the modules used for either global (c, d) or local (e, f) alignment under either LPS (c, e) or PAM (d, f) stimulations. Genes are ordered along the rows, while single cells are ordered by their pseudo-time scores on columns. Collection times and pseudo-time scores per single cell are displayed at the top. White matrix elements correspond to missing values (zeros). (g) Overall, non-imputed, scaled expression of a gene module by single cells (colors denote collection time) and interpolated points (black) along PAM (top) and LPS (bottom) trajectories. Values correspond to the 0.6 quantile scaled expression across the genes of the module. Dashed lines are point alignments from cellAlign global alignment. (h) Dissimilarity matrix and cellAlign global alignment of the non-imputed expression dynamics of one gene module along LPS (left to right) and PAM (top to bottom) trajectories. Histograms depict the distributions of collection time-points along either trajectory. Absolute values and signs of the pseudo-time shift elements, as derived from the alignment, are displayed as color bars at bottom. (i) Dissimilarity matrix and local alignment between non-imputed expression dynamics of a single gene module along LPS (left to right) and PAM (top to bottom) trajectories under similarity threshold of 0.25 of the range of values in the dissimilarity matrix. (j) Overall, non-imputed, expression of a gene-module whose expression pattern is conserved early following both LPS and PAM stimuli and deviates afterwards. Values correspond to the median interpolated scaled expression across module genes. Locally aligned points are red, whereas unaligned, non-conserved points in the trajectory are gray. (k) Distributions of the single-gene correlation difference between the aligned and un-aligned trajectories for global (pink, two-sided t-test p-value = 3*10−9, n = 67 genes) and local (top, two sided t-test p-value=0.027, n = 60 genes) alignment. Only those genes with less than overall 200 missing values in both LPS and PAM stimulated single cells participated in this analysis.
Supplementary Figure 8 Assembly of single-cell trajectories describing gene-expression dynamics during human and mouse preimplantation embryogenesis.
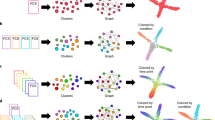
PCA applied on single cell RNA sequencing data using n=80 single cells from human origin (a,b) and n=280 single cells from mouse origin (c,d) preimplantation embryogenesis. Dots represent single cells. Colors denote either different developmental stages (a,c) or scaled pseudo-time scores (b,d) as calculated by Waterfall algorithm. Abbreviations: PC: principal component.
Supplementary Figure 9 Application of cellAlign on mouse and human preimplantation trajectories can be used to discriminate between genes with distinct dynamics rates in both species.
(a-b) Scaled expression (z-scores) of genes assigned to embryonic (a) and maternal (b) clusters across human developmental stages. (c) Scatterplot of the ordering of genes by their upregulation timing along human (x-axis) and mouse (y-axis) trajectories, showing high conservation of temporal regulation (n=288 genes, r = 0.18, linear regression p-value = 0.00173), yet marked differential regulation for a subset of genes. Color denotes point density.
Supplementary Figure 10 Global alignment of B-cell related proteins expression dynamics along B-cell lymphopoiesis in human and mouse mass cytometry data.
(a) A scheme of the comparison between expression dynamics during B-cell lymphopoiesis between human and mouse. (b) A dissimilarity matrix and global alignment of markers expression dynamics between human and mouse B-cell lymphopoiesis. (c, d) Expression dynamics of B-cell related markers along human (c) and mouse (d) B-cells lymphopoiesis. (e) Gating strategy for mouse B-cells mass-cytometry dataset.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 and Supplementary Notes 1–5 (PDF 2638 kb)
Supplementary Table 1
Functional enrichment of embryonic and maternal gene-clusters (XLSX 19 kb)
Supplementary Table 2
Antibodies used for mass cytometry experiment (XLSX 11 kb)
Supplementary Software
cellAlign package (ZIP 735 kb)
Rights and permissions
About this article
Cite this article
Alpert, A., Moore, L., Dubovik, T. et al. Alignment of single-cell trajectories to compare cellular expression dynamics. Nat Methods 15, 267–270 (2018). https://doi.org/10.1038/nmeth.4628
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4628
This article is cited by
-
DELVE: feature selection for preserving biological trajectories in single-cell data
Nature Communications (2024)
-
Baf-mediated transcriptional regulation of teashirt is essential for the development of neural progenitor cell lineages
Experimental & Molecular Medicine (2024)
-
Detection of continuous hierarchical heterogeneity by single-cell surface antigen analysis in the prognosis evaluation of acute myeloid leukaemia
BMC Bioinformatics (2023)
-
A single-cell trajectory atlas of striatal development
Scientific Reports (2023)
-
Reconstructing disease dynamics for mechanistic insights and clinical benefit
Nature Communications (2023)