Abstract
Single-molecule localization microscopy techniques have proven to be essential tools for quantitatively monitoring biological processes at unprecedented spatial resolution. However, these techniques are very low throughput and are not yet compatible with fully automated, multiparametric cellular assays. This shortcoming is primarily due to the huge amount of data generated during imaging and the lack of software for automation and dedicated data mining. We describe an automated quantitative single-molecule-based super-resolution methodology that operates in standard multiwell plates and uses analysis based on high-content screening and data-mining software. The workflow is compatible with fixed- and live-cell imaging and allows extraction of quantitative data like fluorophore photophysics, protein clustering or dynamic behavior of biomolecules. We demonstrate that the method is compatible with high-content screening using 3D dSTORM and DNA-PAINT based super-resolution microscopy as well as single-particle tracking.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Neumann, B. et al. High-throughput RNAi screening by time-lapse imaging of live human cells. Nat. Methods 3, 385–390 (2006).
Wachsmuth, M. et al. High-throughput fluorescence correlation spectroscopy enables analysis of proteome dynamics in living cells. Nat. Biotechnol. 33, 384–389 (2015).
Bray, M.A. et al. Cell Painting, a high-content image-based assay for morphological profiling using multiplexed fluorescent dyes. Nat. Protoc. 11, 1757–1774 (2016).
Liu, Z., Lavis, L.D. & Betzig, E. Imaging live-cell dynamics and structure at the single-molecule level. Mol. Cell 58, 644–659 (2015).
Rust, M.J., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–795 (2006).
van de Linde, S., Sauer, M. & Heilemann, M. Subdiffraction-resolution fluorescence imaging of proteins in the mitochondrial inner membrane with photoswitchable fluorophores. J. Struct. Biol. 164, 250–254 (2008).
van de Linde, S. et al. Direct stochastic optical reconstruction microscopy with standard fluorescent probes. Nat. Protoc. 6, 991–1009 (2011).
Duwé, S. et al. Expression-enhanced fluorescent proteins based on enhanced green fluorescent protein for super-resolution microscopy. ACS Nano 9, 9528–9541 (2015).
Sharonov, A. & Hochstrasser, R.M. Wide-field subdiffraction imaging by accumulated binding of diffusing probes. Proc. Natl. Acad. Sci. USA 103, 18911–18916 (2006).
Giannone, G. et al. Dynamic superresolution imaging of endogenous proteins on living cells at ultra-high density. Biophys. J. 99, 1303–1310 (2010).
Jungmann, R. et al. Multiplexed 3D cellular super-resolution imaging with DNA-PAINT and Exchange-PAINT. Nat. Methods 11, 313–318 (2014).
Hess, S.T., Girirajan, T.P. & Mason, M.D. Ultra-high resolution imaging by fluorescence photoactivation localization microscopy. Biophys. J. 91, 4258–4272 (2006).
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Manley, S. et al. High-density mapping of single-molecule trajectories with photoactivated localization microscopy. Nat. Methods 5, 155–157 (2008).
Sibarita, J.B. High-density single-particle tracking: quantifying molecule organization and dynamics at the nanoscale. Histochem. Cell Biol. 141, 587–595 (2014).
Franke, C., Sauer, M. & van de Linde, S. Photometry unlocks 3D information from 2D localization microscopy data. Nat. Methods 14, 41–44 (2016).
Min, J. et al. 3D high-density localization microscopy using hybrid astigmatic/ biplane imaging and sparse image reconstruction. Biomed. Opt. Express 5, 3935–3948 (2014).
Winterflood, C.M., Platonova, E., Albrecht, D. & Ewers, H. Dual-color 3D superresolution microscopy by combined spectral-demixing and biplane imaging. Biophys. J. 109, 3–6 (2015).
Ovesný, M., Pavel, K., Švindrych, Z. & Hagen, G. M. High density 3D localization microscopy using sparse support recovery. Opt. Express 22, 31263–31276 (2014).
Babcock, H., Sigal, Y.M. & Zhuang, X. A high-density 3D localization algorithm for stochastic optical reconstruction microscopy. Opt. Nanoscopy 1, 6 (2012).
Bullen, A. Microscopic imaging techniques for drug discovery. Nat. Rev. Drug Discov. 7, 54–67 (2008).
Flottmann, B. et al. Correlative light microscopy for high-content screening. Biotechniques 55, 243–252 (2013).
Gunkel, M., Flottmann, B., Heilemann, M., Reymann, J. & Erfle, H. Integrated and correlative high-throughput and super-resolution microscopy. Histochem. Cell Biol. 141, 597–603 (2014).
Holden, S.J. et al. High throughput 3D super-resolution microscopy reveals Caulobacter crescentus in vivo Z-ring organization. Proc. Natl. Acad. Sci. USA 111, 4566–4571 (2014).
Pereira, P.M., Almada, P. & Henriques, R. High-content 3D multicolor super-resolution localization microscopy. In Methods in Cell Biology Vol 125 (ed. Paluch, E.K.) Ch. 7 (Elsevier, 2015).
Jones, T.R. et al. CellProfiler Analyst: data exploration and analysis software for complex image-based screens. BMC Bioinformatics 9, 482 (2008).
Ovesný, M., Křížek, P., Borkovec, J., Švindrych, Z. & Hagen, G.M. ThunderSTORM: a comprehensive ImageJ plug-in for PALM and STORM data analysis and super-resolution imaging. Bioinformatics 30, 2389–2390 (2014).
Huang, B., Wang, W., Bates, M. & Zhuang, X. Three-dimensional super-resolution imaging by stochastic optical reconstruction microscopy. Science 319, 810–813 (2008).
Heilemann, M., van de Linde, S., Mukherjee, A. & Sauer, M. Super-resolution imaging with small organic fluorophores. Angew. Chem. Int. Edn Engl. 48, 6903–6908 (2009).
Endesfelder, U. et al. Chemically induced photoswitching of fluorescent probes—a general concept for super-resolution microscopy. Molecules 16, 3106–3118 (2011).
Swoboda, M. et al. Enzymatic oxygen scavenging for photostability without pH drop in single-molecule experiments. ACS Nano 6, 6364–6369 (2012).
Olivier, N., Keller, D., Gönczy, P. & Manley, S. Resolution doubling in 3D-STORM imaging through improved buffers. PLoS One 8, e69004 (2013).
Jungmann, R. et al. Single-molecule kinetics and super-resolution microscopy by fluorescence imaging of transient binding on DNA origami. Nano Lett. 10, 4756–4761 (2010).
Thompson, R.E., Larson, D.R. & Webb, W.W. Precise nanometer localization analysis for individual fluorescent probes. Biophys. J. 82, 2775–2783 (2002).
Zhang, X. et al. Highly photostable, reversibly photoswitchable fluorescent protein with high contrast ratio for live-cell superresolution microscopy. Proc. Natl. Acad. Sci. USA 113, 10364–10369 (2016).
Legant, W.R. et al. High-density three-dimensional localization microscopy across large volumes. Nat. Methods 13, 359–365 (2016).
Dempsey, G.T., Vaughan, J.C., Chen, K.H., Bates, M. & Zhuang, X. Evaluation of fluorophores for optimal performance in localization-based super-resolution imaging. Nat. Methods 8, 1027–1036 (2011).
Vogelsang, J. et al. Make them blink: probes for super-resolution microscopy. Chemphyschem 11, 2475–2490 (2010).
Chazeau, A. et al. Nanoscale segregation of actin nucleation and elongation factors determines dendritic spine protrusion. EMBO J. 33, 2745–2764 (2014).
Zhang, M. et al. Rational design of true monomeric and bright photoactivatable fluorescent proteins. Nat. Methods 9, 727–729 (2012).
Triller, A. & Choquet, D. New concepts in synaptic biology derived from single-molecule imaging. Neuron 59, 359–374 (2008).
Penn, A.C. et al. Hippocampal LTP and contextual learning require surface diffusion of AMPA receptors. Nature 549, 384–388 (2017).
Heine, M. et al. Surface mobility of postsynaptic AMPARs tunes synaptic transmission. Science 320, 201–205 (2008).
Zhang, H. et al. Regulation of AMPA receptor surface trafficking and synaptic plasticity by a cognitive enhancer and antidepressant molecule. Mol. Psychiatry 18, 471–484 (2013).
Levet, F. et al. SR-Tesseler: a method to segment and quantify localization-based super-resolution microscopy data. Nat. Methods 12, 1065–1071 (2015).
Jungmann, R. et al. Quantitative super-resolution imaging with qPAINT. Nat. Methods 13, 439–442 (2016).
Min, J. et al. FALCON: fast and unbiased reconstruction of high-density super-resolution microscopy data. Sci. Rep. 4, 4577 (2014).
Douglass, K.M., Sieben, C., Archetti, A., Lambert, A. & Manley, S. Super-resolution imaging of multiple cells by optimised flat-field epi-illumination. Nat. Photonics 10, 705–708 (2016).
Jones, T.R. et al. Scoring diverse cellular morphologies in image-based screens with iterative feedback and machine learning. Proc. Natl. Acad. Sci. USA 106, 1826–1831 (2009).
McGuire, H., Aurousseau, M.R.P., Bowie, D. & Blunck, R. Automating single subunit counting of membrane proteins in mammalian cells. J. Biol. Chem. 287, 35912–35921 (2012).
Kechkar, A., Nair, D., Heilemann, M., Choquet, D. & Sibarita, J.B. Real-time analysis and visualization for single-molecule based super-resolution microscopy. PLoS One 8, e62918 (2013).
Schnitzbauer, J., Strauss, M.T., Schlichthaerle, T., Schueder, F. & Jungmann, R. Super-resolution microscopy with DNA-PAINT. Nat. Protoc. 12, 1198–1228 (2017).
Nair, D. et al. Super-resolution imaging reveals that AMPA receptors inside synapses are dynamically organized in nanodomains regulated by PSD95. J. Neurosci. 33, 13204–13224 (2013).
Acknowledgements
This work was supported by the advanced ERC grant ADOS to D.C., the regional council of Aquitaine grant Dynascreen to D.C. and J.-B.S., the FranceBioImaging infrastructure ANR-10-INBS-04 to J.-B.S., the LabEx BRAIN and the IdEx Bordeaux to J.-B.S., and the Fondation pour la Recherche Médicale DEI20151234402 and ANR-Integractome to G.G.
Author information
Authors and Affiliations
Contributions
A.B. performed all the experiments with the help of R.G., C.B. and M.C. A.K. developed the software for online processing and Single Molecule Profiler. F.L. and C.B. performed quantitative data analysis. M.C. developed the HCS-SMLM acquisition interface. O.R. and G.G. designed some biological control experiments. All the authors contributed to the manuscript. J.-B.S. and D.C. came up with the original idea. J.-B.S. supervised this work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 File Hierarchy, Single Molecule Profiler software interface and datamining using Cell Profiler Analyst.
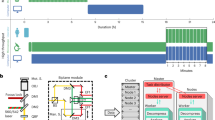
The Single Molecule Profiler (SMP) software automatically generates a database from the single molecule localization files that have been organized in a plate layout format, as illustrated by the tree folder hierarchy. ASCII metadata text files (i.e. DetectionsFile.txt, TracksFile.txt, ClusterFile.txt, etc.) must contain the same number of header lines, identical column separators and descriptors for all wells and all positions, all of which can be specified by the user. A single metadata text file is used as a model for the data format and is parsed to generate descriptors for each column in the file. The user may select which descriptors, as well as the maximum number of objects (detections, tracks or clusters...), to be included in the database to reduce the final database size. The Single Molecule Profiler automatically generates the all files (red rectangle) necessary for importing into Cell Profiler Analyst (http://cellprofiler.org/cp-analyst/).
Supplementary Figure 2 Acquisition template and 3D calibration for astigmatic-based 3D dSTORM of a complete 96-well plate.
(a) Snake-like acquisition template corresponding to a travel length of 85 cm. Dark-grey wells containing adsorbed nanodiamonds have been used to compute the astigmatic-based 3D calibration (Z distance of 1 μm with 50 nm step size). (b) Mean of SigmaY-SigmaX of the astigmatic PSF as a function of Z position computed from 15 nanodiamonds Z-stacks (left) and representative images of nanodiamonds as a function of Z position in 4 extreme wells of a plate (right). Error bars represent S.D. We can notice the excellent reproducibility of the astigmatic calibration across entire plate.
Supplementary Figure 3 Single-molecule signal quality control across entire p96-well plate for 3D dSTORM acquisition.
(a) Heat maps of the total number of localizations (top), mean of Gaussian fit Chi² (middle), and median of the centroid Z coordinate for all the localizations (bottom). We can notice that there is no correlation between the position inside the plate and these quantitative metadata, illustrating that this p96-well plate is suitable to achieve reproducible 3D super-resolution imaging. (b) Normalized histoplots of integrated intensity per detection (I0) grouped by well. A similar distribution can be observed from the first well (A1) until the last well (H3). (c) Comparison between extreme positions on a plate (A1 and H12) acquired automatically using HCS-SMLM with a manual acquisition done in the middle of the plate (E6). These quality control experiments are coherent with the 3D super-resolution images acquired in Figure 3, demonstrating the possibility to automatically acquire hundreds of images in 3D SMLM with the same quality compared to manual acquisitions.
Supplementary Figure 4 dSTORM buffer efficiency over time.
(a) (Top) Heatmap representation of frame_50, the median frame at which 50% of the localized have been detected. In a perfect dSTORM regime, the median frame should be close to the middle frame (4,000 for an acquisition of 8,000 frames) as for well A1. (Bottom) Boxplots displaying the median values and IQR of 8 groups of 12 wells grouped per hour after buffer incubation (corresponding to an entire line of the plate, as color box next to the heatmap). (b) Normalized histoplots of the number of localizations per frame in wells separated by 1 hour. Even if the total number of detections and the quality of the single molecule detections are suitable during the complete 96-well plate acquisition (8 hours, see Fig. 3 and Fig. 4), we can observe that the buffer efficiency for Alexa 647 starts being affected already after 1 hour of incubation (well A10 as compared to well A1). A continuous decrease of the frame_50 metadata correlates with the buffer incubation time. (c) 3D dSTORM images taken 6 hours, 10 hours and 14 hours after the first acquisition. Images have been acquired automatically in the same plate in different wells. We can observe that, after 14 hours of buffer incubation, the total number of localizations became too small (less than 1 million in a FOV of 20.5X20.5 μm2) to properly reconstruct continuous microtubules with the buffer 200 mM of thiols at pH7.2.
Supplementary Figure 5 3D DNA-PAINT acquisitions of microtubules in a p96-well plate.
(a) Left: Normalized histoplots of the number of detections per frame (for the first 8,000 frames) in 3 different wells. In contrast to the dSTORM acquisition, we can observe a constant number of localizations over the course of the 9-hour acquisition. Right: Normalized histoplots of integrated intensity per detection (I0) grouped by well showing a similar distribution from the first well (A1) until the last well (B7). (b) 18 astigmatism-based 3D DNA-PAINT SMLM images (FOV 20.5 x 20.5 μm, 40 nm/px) of microtubules in Cos-7 cells automatically gathered using our HCS-SMLM approach. It corresponds to an acquisition time of 9 hours (30 min per cell), illustrating (1) the capability to efficiently acquire DNA-PAINT data using our HCS-SMLM pipeline, and (2) the time-consuming process of DNA-PAINT as compared to dSTORM. In 9 hours, the HCS-dSTORM pipeline acquired 5-fold more positions with similar microtubule reconstruction quality compared to HCS-DNA-PAINT (only 18 cells in 9 hours). The DNA-PAINT acquisition generated also a larger database as more molecules were detected (2 to 5 fold) than in a dSTORM acquisition due to a higher duty cycle of DNA probes than organic dyes. (c) Examples of astigmatism-based 3D DNA-PAINT images in well A1, after 4 hours (well A8), and after 9 hours (well B7). Color codes for the Z-position. Scale bar: 5 μm.
Supplementary Figure 6 Stepbleaching analysis of isolated Alexa 647 fluorophores inside a P96-well plate and acquisition scheme for fluorophore photophysics study.
(a) Isolated single fluorophore conditions of the coated 96-well plate were controlled using a stepbleaching experiment using the PIF software 1. For stepbleaching analysis, fluorophores were imaged at 5 different positions per well in Tris-HCl pH7.5 buffer (10 kW/cm², 20ms exposure time for 20 sec). Left: Maximum intensity projection of a stack of Alexa 647 fluorophores (AF 647). Top right: representative intensity time trace for one detected fluorophore showing one single bleaching step. Bottom right: stepbleaching distribution for all detected molecules (n = 214). (b) dSTORM acquisition template for organic dyes well plate: a “pumping” phase with 100% of laser power was applied (640 nm: 40 kW/cm², 532 nm: 20 kW/cm² both for 10 sec) in order to excite a maximum of molecules to the triplet state. Immediately after the pumping phase, the single-molecule sequence (“STORM regime”) was automatically launched for a duration of 2 minutes at 50% laser power. For mEOS3.2 isolated proteins, no “pumping” phase was needed and the 405 nm and 561 nm lasers were used at constant power during the acquisition phase. 16 buffers on 3 different fluorophores (48 conditions) were tested.
Supplementary Figure 7 2 color dSTORM acquisition.
Microtubules labelled with Alexa 532 and Vimentin labelled with Alexa 647 in Cos-7 cells using the same dSTORM buffer (200mM bMEA at pH7.2) selected from the characterization of isolated fluorophores (same as Figure 2c). Super-resolution (SR) reconstructions were compared to high resolution (HR) images. Focus position was set at the bottom of the cell (approx. 300 nm above the coverslip surface in TIRF illumination at 50fps).
Supplementary Figure 8 Comparison of buffers for 2 color dSTORM acquisition.
LaminB1 labelled with Alexa 532 and microtubules labelled with Alexa 647 in Cos-7 cells using 2 different buffers (Top panel: 25mM bMEA at pH7.2; Bottom panel: 25 mM bMEA at pH5.2) displaying poor photophysics characteristics for both fluorophores (see Fig. 2b). Focus was at the top of the cell, approximately 5 μm above the coverslip surface. Acquisitions were achieved in the same conditions as for Figure 2c. As described by the photophysics parameters in Figure 2b, Alexa 647 detection was clearly affected by the decrease in thiols concentration (Top Panel) and of the pH (Bottom Panel). Also and as previously described, Alexa 532 detection is slightly affected with an apparent smaller number of localized single molecules, especially for the buffer with 25 mM bMEA, pH 5.2 (Bottom Panel).
Supplementary Figure 9 Cell viability and culture medium.
(a) Plate layout with 3 types of culture medium tested on a p96-well plate. (b) Example of cells imaged by widefield time-lapse microscopy for at least 8 hours. After more than 4 hours of acquisition, HeLa cells clearly required serum to stay alive and spread correctly on the support. After 8 hours of acquisition on the setup in the Fluorobrite+Glutamax+serum medium, a similar number of cells and identical shapes were obtained as compared to classical medium (DMEM+Glutamax+SVF), validating the use of Fluorobrite medium as an imaging medium for our living cells experiments. (c) No difference in background fluorescence (yellow boxes, quantification on right) or single molecule intensity (green boxes) was noticed between Fluorobrite+Glutamax+serum medium and PBS.
Supplementary Figure 10 Single molecule signal quality control across an entire 96-well plate for PALM imaging.
(a) Example of single molecule localization quality control. Here, acquired positions whose photon distributions demonstrate a strong (above 10%) population of multi-emitter detections are automatically excluded. The limit between single and multi-emitter is defined on 10 distributions of isolated proteins. White arrows indicate multi-emitter signals due to a too high density of proteins inside the cell. (b) Plate layout of a control assay on p96-well plate with purified mEos3.2 protein adsorbed to the glass (4 dark grey wells on plate corners). Ten random positions per well were acquired and the histogram represents the cumulative diffusion coefficient metadata. The threshold between mobile and immobile molecules was set to 10-2 um²/sec (see Online Methods). Only a weak fraction of mobile tracks (10%) were measured, indicating good stability of the plate on the stage (no drift, no deformation) during the screening process. (c) Left: Plate layout of a control assay with living HeLa cells expressing SEP::GluA1::mEos2 across 60 wells of a p96-well plate. Three positions per well were acquired and the diffusion coefficients pooled in one histogram per well. Right: Histogram of 6 representative wells (corresponding to dark grey wells), showing similar DCoef distributions and fractions of immobile tracks (N=180 cells).
Supplementary Figure 11 Different modes of acquisition sequences.
4 different concentrations of Ab crosslinker (Polyclonal Ab anti-SEP) were tested (0, 1/100, 1/300, 1/1000) on HeLa cells expressing SEP::GluA1::mEos2.1, and 3 types of acquisition sequences were performed: I/ Sequential acquisition: antibodies were incubated well by well, requiring manual intervention between each well. II/ Serial acquisition: all antibody concentrations were loaded at the same time at the beginning of the acquisition (no user intervention during acquisition). The acquisition order was the well without Ab first, then the wells with 1/100, 1/300 and 1/1000 of Ab concentration. III/ Random acquisition: same as II/ except that the positions were randomly acquired between all wells.
Supplementary Figure 12 Percentage of immobile tracks and variability depending of the acquisition sequence.
Cumulative histograms of DCoefs per well, (N=10 cells per well), and corresponding percentage of immobile molecules. The dose response of Ab crosslinkers were similar for the 3 acquisition modes.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12, Supplementary Table 1 and Supplementary Notes 1–4 (PDF 3977 kb)
Life Sciences Reporting Summary
Life Sciences Reporting Summary (PDF 133 kb)
Supplementary Software 1
Single Molecule Profiler (ZIP 1678 kb)
Source data
Rights and permissions
About this article
Cite this article
Beghin, A., Kechkar, A., Butler, C. et al. Localization-based super-resolution imaging meets high-content screening. Nat Methods 14, 1184–1190 (2017). https://doi.org/10.1038/nmeth.4486
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4486
This article is cited by
-
Enhanced detection of fluorescence fluctuations for high-throughput super-resolution imaging
Nature Photonics (2023)
-
Field-dependent deep learning enables high-throughput whole-cell 3D super-resolution imaging
Nature Methods (2023)
-
An integrated platform for high-throughput nanoscopy
Nature Biotechnology (2023)
-
Super-resolution microscopy: a closer look at synaptic dysfunction in Alzheimer disease
Nature Reviews Neuroscience (2021)
-
Single-molecule localization microscopy
Nature Reviews Methods Primers (2021)