Abstract
Top-down proteomics, the analysis of intact proteins in their endogenous form, preserves valuable information about post-translation modifications, isoforms and proteolytic processing. The quality of top-down liquid chromatography–tandem MS (LC-MS/MS) data sets is rapidly increasing on account of advances in instrumentation and sample-processing protocols. However, top-down mass spectra are substantially more complex than conventional bottom-up data. New algorithms and software tools for confident proteoform identification and quantification are needed. Here we present Informed-Proteomics, an open-source software suite for top-down proteomics analysis that consists of an LC-MS feature-finding algorithm, a database search algorithm, and an interactive results viewer. We compare our tool with several other popular tools using human-in-mouse xenograft luminal and basal breast tumor samples that are known to have significant differences in protein abundance based on bottom-up analysis.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
13 June 2018
In the version of this article initially published, the authors erroneously reported the search mode that was used for ProSightPC 3.0 in the Online Methods and in Supplementary Table 3. The results presented in Fig. 5 were obtained with 'absolute mass' search mode, not 'biomarker discovery' search mode. The 'biomarker discovery' search mode of ProSightPC 3.0 looks for subsequences of those contained in the annotated proteoform database (e.g., truncated forms from degradation and/or cleavage). This search mode is expected to generate similar numbers of identifications as Informed-Proteomics, but is also expected to take dramatically longer (~480 CPU hours). Unfortunately, because of these heavy computational requirements, the authors were unable to complete an analysis using this search mode. They chose to use 'absolute mass' mode to illustrate the effect of search mode and database choice on the results. 'Absolute mass' mode is the most restrictive of the search modes illustrated in Fig. 5, as it searches only for proteoforms explicitly listed in the proteoform database within a user-defined mass tolerance. In addition, in the supplementary information originally published online, Supplementary Table 3 incorrectly stated that ProSightPC v3.0 was used in 'biomarker discovery' mode. 'Absolute mass' mode was the mode actually used in this comparison. These errors have been corrected in the HTML and PDF versions of this article and in the associated supplementary information.
References
Garcia, B.A. What does the future hold for top down mass spectrometry? J. Am. Soc. Mass Spectrom. 21, 193–202 (2010).
Siuti, N. & Kelleher, N.L. Decoding protein modifications using top-down mass spectrometry. Nat. Methods 4, 817–821 (2007).
Smith, L.M. & Kelleher, N.L. Proteoform: a single term describing protein complexity. Nat. Methods 10, 186–187 (2013).
Zhang, Z., Wu, S., Stenoien, D.L. & Paša-Tolic´, L. High-throughput proteomics. Annu. Rev. Anal. Chem. (Palo Alto Calif.) 7, 427–454 (2014).
Tran, J.C. et al. Mapping intact protein isoforms in discovery mode using top-down proteomics. Nature 480, 254–258 (2011).
Chait, B.T. Chemistry. Mass spectrometry: bottom-up or top-down? Science 314, 65–66 (2006).
Lanucara, F. & Eyers, C.E. Top-down mass spectrometry for the analysis of combinatorial post-translational modifications. Mass Spectrom. Rev. 32, 27–42 (2013).
Horn, D.M., Zubarev, R.A. & McLafferty, F.W. Automated reduction and interpretation of high resolution electrospray mass spectra of large molecules. J. Am. Soc. Mass Spectrom. 11, 320–332 (2000).
Zabrouskov, V., Senko, M.W., Du, Y., Leduc, R.D. & Kelleher, N.L. New and automated MSn approaches for top-down identification of modified proteins. J. Am. Soc. Mass Spectrom. 16, 2027–2038 (2005).
Liu, X. et al. Deconvolution and database search of complex tandem mass spectra of intact proteins: a combinatorial approach. Mol. Cell. Proteomics 9, 2772–2782 (2010).
Kou, Q., Wu, S. & Liu, X. A new scoring function for top-down spectral deconvolution. BMC Genomics 15, 1140 (2014).
LeDuc, R.D. et al. ProSight PTM: an integrated environment for protein identification and characterization by top-down mass spectrometry. Nucleic Acids Res. 32, W340–W345 (2004).
Zamdborg, L. et al. ProSight PTM 2.0: improved protein identification and characterization for top down mass spectrometry. Nucleic Acids Res. 35, W701–W706 (2007).
Liu, X. et al. Protein identification using top-down. Mol. Cell. Proteomics 11, M111.008524 (2012).
Kou, Q., Xun, L. & Liu, X. TopPIC: a software tool for top-down mass spectrometry-based proteoform identification and characterization. Bioinformatics 32, 3495–3497 (2016).
Sun, R.-X. et al. pTop 1.0: a high-accuracy and high-efficiency search engine for intact protein identification. Anal. Chem. 88, 3082–3090 (2016).
Cai, W. et al. MASH Suite Pro: a comprehensive software tool for top-down proteomics. Mol. Cell. Proteomics 15, 703–714 (2016).
Guner, H. et al. MASH Suite: a user-friendly and versatile software interface for high-resolution mass spectrometry data interpretation and visualization. J. Am. Soc. Mass Spectrom. 25, 464–470 (2014).
Kim, S., Gupta, N. & Pevzner, P.A. Spectral probabilities and generating functions of tandem mass spectra: a strike against decoy databases. J. Proteome Res. 7, 3354–3363 (2008).
Kim, S. & Pevzner, P.A.M.S.-G.F. MS-GF+ makes progress towards a universal database search tool for proteomics. Nat. Commun. 5, 5277 (2014).
Elias, J.E. & Gygi, S.P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Pevzner, P.A., Dancík, V. & Tang, C.L. Mutation-tolerant protein identification by mass spectrometry. J. Comput. Biol. 7, 777–787 (2000).
Liu, X. et al. Identification of ultramodified proteins using top-down tandem mass spectra. J. Proteome Res. 12, 5830–5838 (2013).
Frank, A.M., Pesavento, J.J., Mizzen, C.A., Kelleher, N.L. & Pevzner, P.A. Interpreting top-down mass spectra using spectral alignment. Anal. Chem. 80, 2499–2505 (2008).
Tsur, D., Tanner, S., Zandi, E., Bafna, V. & Pevzner, P.A. Identification of post-translational modifications by blind search of mass spectra. Nat. Biotechnol. 23, 1562–1567 (2005).
Frank, A., Tanner, S., Bafna, V. & Pevzner, P. Peptide sequence tags for fast database search in mass-spectrometry. J. Proteome Res. 4, 1287–1295 (2005).
Domon, B. & Aebersold, R. Options and considerations when selecting a quantitative proteomics strategy. Nat. Biotechnol. 28, 710–721 (2010).
Nagaraj, N. & Mann, M. Quantitative analysis of the intra- and inter-individual variability of the normal urinary proteome. J. Proteome Res. 10, 637–645 (2011).
Zhu, W., Smith, J.W. & Huang, C.-M. Mass spectrometry-based label-free quantitative proteomics. J. Biomed. Biotechnol. 2010, 840518 (2010).
Qian, W.-J., Jacobs, J.M., Liu, T., Camp, D.G. II & Smith, R.D. Advances and challenges in liquid chromatography-mass spectrometry-based proteomics profiling for clinical applications. Mol. Cell. Proteomics 5, 1727–1744 (2006).
Li, S. et al. Endocrine-therapy-resistant ESR1 variants revealed by genomic characterization of breast-cancer-derived xenografts. Cell Rep. 4, 1116–1130 (2013).
Tabb, D.L. et al. Reproducibility of differential proteomic technologies in CPTAC fractionated xenografts. J. Proteome Res. 15, 691–706 (2016).
Ntai, I. et al. Integrated bottom-up and top-down proteomics of patient-derived breast tumor xenografts. Mol. Cell. Proteomics 15, 45–56 (2016).
Senko, M.W., Beu, S.C. & McLaffertycor, F.W. Determination of monoisotopic masses and ion populations for large biomolecules from resolved isotopic distributions. J. Am. Soc. Mass Spectrom. 6, 229–233 (1995).
Wang, X. et al. JUMP: a tag-based database search tool for peptide identification with high sensitivity and accuracy. Mol. Cell. Proteomics 13, 3663–3673 (2014).
Martens, L. et al. mzML--a community standard for mass spectrometry data. Mol. Cell. Proteomics 10, R110.000133 (2011).
Jones, A.R. et al. The mzIdentML data standard for mass spectrometry-based proteomics results. Mol. Cell. Proteomics 11, M111.014381 (2012).
Chambers, M.C. et al. A cross-platform toolkit for mass spectrometry and proteomics. Nat. Biotechnol. 30, 918–920 (2012).
Acknowledgements
Portions of this work were supported by the NIH National Institute of General Medical Sciences grant GM103493 (R.D.S.), the National Cancer Institute Clinical Proteomic Tumor Analysis Consortium (CPTAC) grant U24CA160019 (R.D.S.), the National Institute of Allergy and Infectious Diseases NIH/DHHS through interagency agreement Y1-A1-8401-01 (J. Adkins, PNNL), and the U.S. Department of Energy (DOE) Office of Science and Office of Biological and Environmental Research, under the Pan-omics program (R.D.S.). L.P.T., N.T., M.Z., and J.B.S. were supported as part of the “High Resolution and Mass Accuracy Capability” development project at the Environmental Molecular Science Laboratory (EMSL), a U.S. DOE national scientific user facility at Pacific Northwest National Laboratory (PNNL) in Richland, Washington. Battelle operates PNNL for the DOE under contract DE-AC05-76RLO01830.
Author information
Authors and Affiliations
Contributions
J.P., P.D.P., S.H.P., and S.K. designed and executed the study. J.P., C.W., J.M., G.M.F., B.C.G., and S.K. implemented algorithms in software. T.L. contributed samples. P.D.P., Y.S., A.K.S., R.J.M. performed LC-MS/MS experiments. J.P., P.D.P., J.B.S., V.A.P., M.Z., T.L., and N.T. analyzed data. L.P.-T. and R.D.S. provided technical leadership and oversight. J.P., P.D.P., and S.K. contributed to writing the manuscript with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
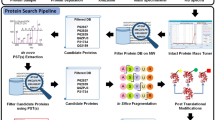
Supplementary Figure 1 Top-down proteomics data analysis workflow in Informed-Proteomics.
Given a LC-MS data, ProMex detects LC-MS features, each of which represents a group of isotopomer envelopes corresponding to the same putative proteoform ion. Detected LC-MS features are fed into a database search tool, MSPathFinder to characterize proteoforms from MS/MS spectra. In LcMsSpectator, users can visualize top-down proteomics data and further refine analysis results reported by ProMex and MSPathFinder. LC-MS features or identified proteoforms from multiple datasets can be aligned and grouped to find differentially expressed features or proteoforms.
Supplementary Figure 2 Averaged Pearson correlation coefficients between observed and theoretical isotopomer envelopes of LC-MS features in Shewanella oneidensis samples.
Due to ions being dispersed widely across LC time, various charge states and isotope species, single isotope envelopes typically have a poor shape and correlation to expected profiles. Aggregating isotopomer envelopes across adjacent charge states and elution times increases the similarity to expected profiles. This improvement is more significant in high molecular weight proteins (e.g. > 15,000 Da).
Supplementary Figure 3 Sequence tag based search for multiply cleaved protein sequences.
Once a protein sequence matches to a sequence tag, two sequence graphs originating from the both ends of the sequence tag are generated and explored toward opposite directions. The flanking mass of sequence tags and the mass of LC-MS features are used to constrain candidate proteoforms to be searched.
Supplementary Figure 4 Examples of data plots in LcMsSpectator.
(a) Matched fragment ion peaks in MS2 spectrum view (left), and precursor ion peaks in previous and next MS1 spectra (right), (b) Peak error heat map for the fragment ions on the MS2 spectrum plot, and (c) Extracted Ion Chromatogram (XIC) view showing neighboring charge states of the precursor ion and the total area of all XIC points displayed.
Supplementary Figure 5 Example of PTM refining from acetylation to tri-methylation at K36 in histone H3 protein.
Peak error heat maps for the fragment ions. The degree of mass errors for fragment ions is represented by the color of grid cells. High mass errors are represented by either red or green cells as indicated by a color bar on right side. (a) Initially identified proteoform having acetylation at 27th Lys residue, and (b) Refined proteoform after changing the acetylation to tri-methylation. LcMsSpectator allows users to visually check and refine MSPathFinder results.
Supplementary Figure 6 LC-MS feature-based analysis on CompRef Sample.
(a) PCA plot and (b) volcano plot. 3604 features are up-regulated in P32(WHIM2) while 3696 features are up-regulated in P33(WHIM16) at 1% FDR and fold change > 2.
Supplementary Figure 7 Comparison of top-down data analysis pipeline.
(a) Differentially expressed proteoforms reported in Ntail et al., Mol Cell Proteomics (2016), and (b) those found by Informed-Proteomics software suite. The same dataset created for Study 3 in the article were used. Informed-Proteomics analysis pipeline found 2.7 and 2.4 times more differentially expressed proteoforms and proteins, respectively.
Supplementary Figure 8 Bayesian network modelling LC-MS features to determine the probability of observing aggregated isotopomer envelopes Ei given mass M .
The Ei is specified by four parameters: ratio abundance (Ai), isotopomer envelope similarity score (Si), normalized intensity (Ii), and elution profile score (Xi). These four parameters are assumed as independent of each other at the given mass (M) and charge state (Ci).
Supplementary Figure 9 Multiple interpretations of an observed isotopomer envelope.
(a) a cluster of observed peaks, and (b-e) different theoretical isotopomer envelopes matched to the envelope in (a). Besides the true match in (b), theoretical envelopes involving ±1 Da monoisotopic mass errors in (c) or different monoisotopic masses with multiples of true charge in (d) or (e) have good matches with the observed peaks.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Tables 1–3. (PDF 1529 kb)
Reporting Summary
Life Sciences Reporting Summary. (PDF 158 kb)
Supplementary Protocol
MSPathFinder Tutorial. (PDF 408 kb)
Supplementary Software
Informed-Proteomics software suite. (ZIP 8537 kb)
Rights and permissions
About this article
Cite this article
Park, J., Piehowski, P., Wilkins, C. et al. Informed-Proteomics: open-source software package for top-down proteomics. Nat Methods 14, 909–914 (2017). https://doi.org/10.1038/nmeth.4388
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4388
This article is cited by
-
Online protein unfolding characterized by ion mobility electron capture dissociation mass spectrometry: cytochrome C from neutral and acidic solutions
Analytical and Bioanalytical Chemistry (2023)
-
Proteogenomics 101: a primer on database search strategies
Journal of Proteins and Proteomics (2023)
-
O-Pair Search with MetaMorpheus for O-glycopeptide characterization
Nature Methods (2020)
-
Getting more for less: new software solutions for glycoproteomics
Nature Methods (2020)