Abstract
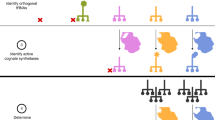
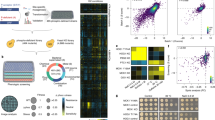
The phosphorylation of threonine residues in proteins regulates diverse processes in eukaryotic cells, and thousands of threonine phosphorylations have been identified. An understanding of how threonine phosphorylation regulates biological function will be accelerated by general methods to biosynthesize defined phosphoproteins. Here we describe a rapid approach for directly discovering aminoacyl-tRNA synthetase–tRNA pairs that selectively incorporate non-natural amino acids into proteins; our method uses parallel positive selections combined with deep sequencing and statistical analysis and enables the direct, scalable discovery of aminoacyl-tRNA synthetase–tRNA pairs with mutually orthogonal substrate specificity. By combining a method to biosynthesize phosphothreonine in cells with this selection approach, we discover a phosphothreonyl-tRNA synthetase–tRNACUA pair and create an entirely biosynthetic route to incorporating phosphothreonine in proteins. We biosynthesize several phosphoproteins and demonstrate phosphoprotein structure determination and synthetic protein kinase activation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Oakhill, J.S., Scott, J.W. & Kemp, B.E. AMPK functions as an adenylate charge-regulated protein kinase. Trends Endocrinol. Metab. 23, 125–132 (2012).
Kelly, A.E. et al. Survivin reads phosphorylated histone H3 threonine 3 to activate the mitotic kinase Aurora B. Science 330, 235–239 (2010).
Ho, D.H. et al. Leucine-rich repeat kinase 2 (LRRK2) phosphorylates p53 and induces p21WAF1/CIP1 expression. Mol. Brain 8, 54 (2015).
Li, J. et al. EYA1′s conformation-specificity in dephosphorylating phosphothreonine in Myc and its activity on Myc stabilization in breast cancer. Mol. Cell. Biol. 37, e00499–16 (2017).
Reinardy, J.L. et al. Phosphorylation of threonine 794 on Tie1 by Rac1/PAK1 reveals a novel angiogenesis regulatory pathway. PLoS One 10, e0139614 (2015).
Mahajan, A. et al. Structure and function of the phosphothreonine-specific FHA domain. Sci. Signal. 1, re12 (2008).
Olsen, J.V. et al. Global, in vivo, and site-specific phosphorylation dynamics in signaling networks. Cell 127, 635–648 (2006).
Zhao, Y.-W., Lai, H.-Y., Tang, H., Chen, W. & Lin, H. Prediction of phosphothreonine sites in human proteins by fusing different features. Sci. Rep. 6, 34817 (2016).
Liu, C.C. & Schultz, P.G. Adding new chemistries to the genetic code. Annu. Rev. Biochem. 79, 413–444 (2010).
Chin, J.W. Expanding and reprogramming the genetic code of cells and animals. Annu. Rev. Biochem. 83, 379–408 (2014).
Davis, L. & Chin, J.W. Designer proteins: applications of genetic code expansion in cell biology. Nat. Rev. Mol. Cell Biol. 13, 168–182 (2012).
Masania, J., Li, J., Smerdon, S.J. & Macmillan, D. Access to phosphoproteins and glycoproteins through semi-synthesis, native chemical ligation and N→S acyl transfer. Org. Biomol. Chem. 8, 5113–5119 (2010).
Rogerson, D.T. et al. Efficient genetic encoding of phosphoserine and its nonhydrolyzable analog. Nat. Chem. Biol. 11, 496–503 (2015).
Park, H.-S. et al. Expanding the genetic code of Escherichia coli with phosphoserine. Science 333, 1151–1154 (2011).
Sauerwald, A. et al. RNA-dependent cysteine biosynthesis in archaea. Science 307, 1969–1972 (2005).
Fukunaga, R. & Yokoyama, S. Structural insights into the first step of RNA-dependent cysteine biosynthesis in archaea. Nat. Struct. Mol. Biol. 14, 272–279 (2007).
Huguenin-Dezot, N. et al. Synthesis of isomeric phosphoubiquitin chains reveals that phosphorylation controls deubiquitinase activity and specificity. Cell Rep. 16, 1180–1193 (2016).
Hauenstein, S.I., Hou, Y.-M. & Perona, J.J. The homotetrameric phosphoseryl-tRNA synthetase from Methanosarcina mazei exhibits half-of-the-sites activity. J. Biol. Chem. 283, 21997–22006 (2008).
Xie, J. & Schultz, P.G. An expanding genetic code. Methods 36, 227–238 (2005).
Fan, C., Fromm, H.J. & Bobik, T.A. Kinetic and functional analysis of L-threonine kinase, the PduX enzyme of Salmonella enterica. J. Biol. Chem. 284, 20240–20248 (2009).
Fan, C. & Bobik, T.A. The PduX enzyme of Salmonella enterica is an L-threonine kinase used for coenzyme B12 synthesis. J. Biol. Chem. 283, 11322–11329 (2008).
Giegé, R., Sissler, M. & Florentz, C. Universal rules and idiosyncratic features in tRNA identity. Nucleic Acids Res. 26, 5017–5035 (1998).
Hohn, M.J., Park, H.-S., O'Donoghue, P., Schnitzbauer, M. & Söll, D. Emergence of the universal genetic code imprinted in an RNA record. Proc. Natl. Acad. Sci. USA 103, 18095–18100 (2006).
Cooley, R.B. et al. Structural basis of improved second-generation 3-nitro-tyrosine tRNA synthetases. Biochemistry 53, 1916–1924 (2014).
Gautier, A. et al. Genetically encoded photocontrol of protein localization in mammalian cells. J. Am. Chem. Soc. 132, 4086–4088 (2010).
Gautier, A., Deiters, A. & Chin, J.W. Light-activated kinases enable temporal dissection of signaling networks in living cells. J. Am. Chem. Soc. 133, 2124–2127 (2011).
Hemphill, J., Borchardt, E.K., Brown, K., Asokan, A. & Deiters, A. Optical control of CRISPR/Cas9 gene editing. J. Am. Chem. Soc. 137, 5642–5645 (2015).
Walker, O.S. et al. Photoactivation of mutant isocitrate dehydrogenase 2 reveals rapid cancer-associated metabolic and epigenetic changes. J. Am. Chem. Soc. 138, 718–721 (2016).
Elliott, T.S. et al. Proteome labeling and protein identification in specific tissues and at specific developmental stages in an animal. Nat. Biotechnol. 32, 465–472 (2014).
Nikić, I. et al. Minimal tags for rapid dual-color live-cell labeling and super-resolution microscopy. Angew. Chem. Int. Ed. Engl. 53, 2245–2249 (2014).
Tang, L. et al. Construction of “small-intelligent” focused mutagenesis libraries using well-designed combinatorial degenerate primers. Biotechniques 52, 149–158 (2012).
Herhaus, L. & Dikic, I. Expanding the ubiquitin code through post-translational modification. EMBO Rep. 16, 1071–1083 (2015).
Wang, K. et al. Optimized orthogonal translation of unnatural amino acids enables spontaneous protein double-labelling and FRET. Nat. Chem. 6, 393–403 (2014).
Neumann, H., Wang, K., Davis, L., Garcia-Alai, M. & Chin, J.W. Encoding multiple unnatural amino acids via evolution of a quadruplet-decoding ribosome. Nature 464, 441–444 (2010).
Wang, K. et al. Defining synonymous codon compression schemes by genome recoding. Nature 539, 59–64 (2016).
Faircloth, B.C. & Glenn, T.C. Not all sequence tags are created equal: designing and validating sequence identification tags robust to indels. PLoS One 7, e42543 (2012).
Sachdeva, A., Wang, K., Elliott, T. & Chin, J.W. Concerted, rapid, quantitative, and site-specific dual labeling of proteins. J. Am. Chem. Soc. 136, 7785–7788 (2014).
Steinfeld, J.B., Aerni, H.R., Rogulina, S., Liu, Y. & Rinehart, J. Expanded cellular amino acid pools containing phosphoserine, phosphothreonine, and phosphotyrosine. ACS Chem. Biol. 9, 1104–1112 (2014).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Vijay-Kumar, S., Bugg, C.E. & Cook, W.J. Structure of ubiquitin refined at 1.8 A resolution. J. Mol. Biol. 194, 531–544 (1987).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Adams, P.D. et al. The Phenix software for automated determination of macromolecular structures. Methods 55, 94–106 (2011).
Murshudov, G.N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D Biol. Crystallogr. 67, 355–367 (2011).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B Stat. Methodol. 57, 289–300 (1995).
Zhang, M.S. et al. Parallel positive selections for the discovery of selective aminoacyl-tRNA synthetase. Protoc. Exch. http://dx.doi.org/10.1038/protex.2017.046 (2017).
Acknowledgements
This work was supported by the Medical Research Council, UK (MC_U105181009 and MC_UP_A024_1008), BBSRC (BB/M000842/1, for automation) and an ERC Advanced Grant (SGCR, 669351), all to J.W.C. M.S.Z. was supported by an EMBO Fellowship (ALTF 297-2015), S.F.B. was supported by a Boehringer Fonds PhD fellowship, and A.D.L. was supported by an NSF fellowship (1523390). We are grateful to D. Cervettini, M. Mahesh, and Y.-H. Tsai for contributions; and to T. Elliott (MRC-LMB) for compound 4. We are grateful to the MRC-LMB mass spectrometry facility for performing ESI-MS/MS. We are grateful to M. Yu, P. Emsley, G. Murshudov, O. Perisic, and R. Williams for help in crystallization and structural data analysis.
Author information
Authors and Affiliations
Contributions
J.W.C. defined the direction of research. D.T.R. suggested using PduX. M.S.Z. demonstrated pThr biosynthesis in E. coli and optimized the tRNA with the help of W.H.S. A.D.L. developed the assay for determining intracellular amino acid concentration. S.F.B. developed the parallel selection and sequencing approach. M.S.Z. and S.F.B. applied the approach to pThrRS discovery. M.S.Z. performed protein expression and biochemical experiments. N.H.-D. carried out protein expression, purification and structural studies with assistance from M.S.Z. M.S.Z., S.F.B., and J.W.C. wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–21 and Supplementary Tables 1–4. (PDF 8259 kb)
Supplementary Data Set 1
Sequences of all plasmids used in this study. Genbank file with all plasmid sequences. (TXT 146 kb)
Supplementary Protocol
Parallel positive selections for the discovery of selective aminoacyl-tRNA synthetase. (PDF 302 kb)
Rights and permissions
About this article
Cite this article
Zhang, M., Brunner, S., Huguenin-Dezot, N. et al. Biosynthesis and genetic encoding of phosphothreonine through parallel selection and deep sequencing. Nat Methods 14, 729–736 (2017). https://doi.org/10.1038/nmeth.4302
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4302
This article is cited by
-
Essential factors, advanced strategies, challenges, and approaches involved for efficient expression of recombinant proteins in Escherichia coli
Archives of Microbiology (2024)
-
Strategies for efficient production of recombinant proteins in Escherichia coli: alleviating the host burden and enhancing protein activity
Microbial Cell Factories (2022)
-
Unleashing the potential of noncanonical amino acid biosynthesis to create cells with precision tyrosine sulfation
Nature Communications (2022)
-
Enhanced access to the human phosphoproteome with genetically encoded phosphothreonine
Nature Communications (2022)
-
Reprogramming the genetic code
Nature Reviews Genetics (2021)