Abstract
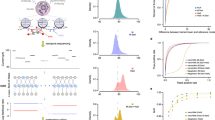
In nanopore sequencing devices, electrolytic current signals are sensitive to base modifications, such as 5-methylcytosine (5-mC). Here we quantified the strength of this effect for the Oxford Nanopore Technologies MinION sequencer. By using synthetically methylated DNA, we were able to train a hidden Markov model to distinguish 5-mC from unmethylated cytosine. We applied our method to sequence the methylome of human DNA, without requiring special steps for library preparation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Miura, F., Enomoto, Y., Dairiki, R. & Ito, T. Nucleic Acids Res. 40, e136 (2012).
Houseman, E.A. et al. BMC Bioinformatics 13, 86 (2012).
Landan, G. et al. Nat. Genet. 44, 1207–1214 (2012).
Flusberg, B.A. et al. Nat. Methods 7, 461–465 (2010).
Beaulaurier, J. et al. Nat. Commun. 6, 7438 (2015).
Clark, T.A. et al. BMC Biol. 11, 4 (2013).
Hahn, M.A., Li, A.X., Wu, X. & Pfeifer, G.P. Methods Mol. Biol. 1238, 273–287 (2015).
Laszlo, A.H. et al. Proc. Natl. Acad. Sci. USA 110, 18904–18909 (2013).
Schreiber, J. et al. Proc. Natl. Acad. Sci. USA 110, 18910–18915 (2013).
Loman, N.J., Quick, J. & Simpson, J.T. Nat. Methods 12, 733–735 (2015).
Szalay, T. & Golovchenko, J.A. Nat. Biotechnol. 33, 1087–1091 (2015).
Schreiber, J. & Karplus, K. Bioinformatics 31, 1897–1903 (2015).
Timp, W., Comer, J. & Aksimentiev, A. Biophys. J. 102, L37–L39 (2012).
Viner, C. et al. bioRxiv http://dx.doi.org/10.1101/043794 (2016).
Ngo, T.T.M. et al. Nat. Commun. 7, 10813 (2016).
Wescoe, Z.L., Schreiber, J. & Akeson, M. J. Am. Chem. Soc. 136, 16582–16587 (2014).
Meyer, K.D. & Jaffrey, S.R. Genome Biol. 17, 5 (2016).
Vogel, M.J., Peric-Hupkes, D. & van Steensel, B. Nat. Protoc. 2, 1467–1478 (2007).
Kelly, T.K. et al. Genome Res. 22, 2497–2506 (2012).
Meissner, A. et al. Nucleic Acids Res. 33, 5868–5877 (2005).
Lee, E.-J. et al. Nucleic Acids Res. 39, e127 (2011).
Krueger, F. & Andrews, S.R. Bioinformatics 27, 1571–1572 (2011).
Hansen, K.D., Langmead, B. & Irizarry, R.A. Genome Biol. 13, R83 (2012).
Quick, J. et al. Nature 530, 228–232 (2016).
Acknowledgements
We thank N. Loman and J. Quick for making the E. coli K12 data set publicly available, and A. Feinberg, K. Hansen and J. McPherson for helpful discussions. J.T.S., P.C.Z., M.D. and L.J.D. are supported by the Ontario Institute for Cancer Research through funds provided by the Government of Ontario. W.T. is supported in part by a Johns Hopkins University Catalyst award.
Author information
Authors and Affiliations
Contributions
J.T.S. and W.T. designed the study; W.T., R.E.W. and P.C.Z. performed the experiments; J.T.S., M.D. and L.J.D. designed and implemented the methylation-training and inference framework; and J.T.S., W.T., R.E.W. and P.C.Z. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
J.T.S. receives research funding from Oxford Nanopore Technologies, and W.T. has two patents licensed to Oxford Nanopore Technologies (US20110226623 A1 and US20120040342 A1). J.T.S. and W.T. have received travel funds to speak at symposia organized by Oxford Nanopore Technologies.
Integrated supplementary information
Supplementary Figure 1 Position dependence of methylation parameter shift.
A histogram of the difference between the trained mean for the methylated R7.3 (left column; PCR+M.SssI-R7.3-timp-021216) and R9 (right column; PCR+M.SssI-R9-timp-061716) datasets and the ONT reference model. Each row is the subset of k-mers with the methylated base in the first position (Mbcdef), the second position (aMcdef), and so on. We only included k-mers that contained a single methylation position. We restrict the plotting range to differences in the range -8 to 8 so some outliers are not shown.
Supplementary Figure 2 Methylation classifier error rate vs threshold.
Panel A shows the error rate of the methylated/unmethylated classifier as a function of the log likelihood ratio threshold required to make a call. In this analysis calls are only made at sites where the absolute value of the log likelihood ratio is greater than the threshold shown. Panel B shows the number of calls made as a function of this threshold.
Supplementary Figure 3 NA12878 CpG island methylation.
In the main panel each point is an annotated CpG island in the human genome that was covered by both bisulfite sequencing data and nanopore reads from the merged natural NA12878 DNA R7.3 data set. The x-coordinate of the point is the percentage of CpGs in the island that were predicted to be methylated from bisulfite data. The y-coordinate is the percentage of CpGs predicted to be methylated by our model using the nanopore data. The points are colored by whether the CGI is in a promoter(blue) or not(red). The histograms on the top and right of the figure are the marginal distributions of methylation percentages for the bisulfite and nanopore calls respectively.
Supplementary Figure 4 Cancer/Normal per CpG methylation analysis.
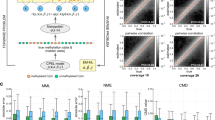
Comparison of bisulfite sequencing and nanopore R7.3 data in a cancer/normal reduced representation dataset. A) Correlation plot of per CpG methylation percentage from MCF10A bisulfite data (x-axis) versus nanopore data (y-axis), Pearson Correlation r=0.91. B) As in A) but for MDA-MB-231 samples, Pearson Correlation r=0.91.
Supplementary Figure 5 Log Likelihood Ratio comparison for human samples.
A histogram of the log likelihood ratios for unmethylated NA12878 DNA (top pane), methylated NA12878 DNA (middle pane) and natural NA12878 DNA (bottom pane). In these figures we only include singleton sites.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Note and Supplementary Tables 1–7 (PDF 1386 kb)
Supplementary Data 1
TSS by chromosomes (PDF 1931 kb)
Supplementary Data 3
Cancer Normal Regions (PDF 1128 kb)
Supplementary Data 4
Cancer-Normal Strand Data (PDF 984 kb)
Supplementary Data 2
CGI bisulfite vs nanopore (PDF 585 kb)
Rights and permissions
About this article
Cite this article
Simpson, J., Workman, R., Zuzarte, P. et al. Detecting DNA cytosine methylation using nanopore sequencing. Nat Methods 14, 407–410 (2017). https://doi.org/10.1038/nmeth.4184
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4184
This article is cited by
-
Prediction of methylation status using WGS data of plasma cfDNA for multi-cancer early detection (MCED)
Clinical Epigenetics (2024)
-
A comparison of methods for detecting DNA methylation from long-read sequencing of human genomes
Genome Biology (2024)
-
The transition from genomics to phenomics in personalized population health
Nature Reviews Genetics (2024)
-
Genome assembly in the telomere-to-telomere era
Nature Reviews Genetics (2024)
-
NAP-seq reveals multiple classes of structured noncoding RNAs with regulatory functions
Nature Communications (2024)