Abstract
We describe ProteomeTools, a project building molecular and digital tools from the human proteome to facilitate biomedical research. Here we report the generation and multimodal liquid chromatography–tandem mass spectrometry analysis of >330,000 synthetic tryptic peptides representing essentially all canonical human gene products, and we exemplify the utility of these data in several applications. The resource (available at http://www.proteometools.org) will be extended to >1 million peptides, and all data will be shared with the community via ProteomicsDB and ProteomeXchange.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Zhang, Y., Fonslow, B.R., Shan, B., Baek, M.C. & Yates, J.R. III. Chem. Rev. 113, 2343–2394 (2013).
Ahrens, C.H., Brunner, E., Qeli, E., Basler, K. & Aebersold, R. Nat. Rev. Mol. Cell Biol. 11, 789–801 (2010).
Kusebauch, U. et al. Cell 166, 766–778 (2016).
Picotti, P. et al. Nature 494, 266–270 (2013).
Mallick, P. et al. Nat. Biotechnol. 25, 125–131 (2007).
Wilhelm, M. et al. Nature 509, 582–587 (2014).
Giansanti, P. et al. Cell Rep. 11, 1834–1843 (2015).
Marx, H. et al. Nat. Biotechnol. 31, 557–564 (2013).
Escher, C. et al. Proteomics 12, 1111–1121 (2012).
Cox, J. et al. J. Proteome Res. 10, 1794–1805 (2011).
Griss, J. et al. Nat. Methods 13, 651–656 (2016).
Savitski, M.M., Wilhelm, M., Hahne, H., Kuster, B. & Bantscheff, M. Mol. Cell. Proteomics 14, 2394–2404 (2015).
Gallien, S. et al. Mol. Cell. Proteomics 11, 1709–1723 (2012).
Lawrence, R.T., Searle, B.C., Llovet, A. & Villén, J. Nat. Methods 13, 431–434 (2016).
Guo, T. et al. Nat. Med. 21, 407–413 (2015).
Vizcaíno, J.A. et al. Nat. Biotechnol. 32, 223–226 (2014).
Bailey, D.J., McDevitt, M.T., Westphall, M.S., Pagliarini, D.J. & Coon, J.J. J. Proteome Res. 13, 2152–2161 (2014).
Kelchtermans, P. et al. Proteomics 14, 353–366 (2014).
Wang, J. et al. Nat. Methods 12, 1106–1108 (2015).
Krokhin, O.V. Anal. Chem. 78, 7785–7795 (2006).
Wenschuh, H. et al. Biopolymers 55, 188–206 (2000).
Rose, C.M. et al. J. Am. Soc. Mass Spectrom. 26, 1848–1857 (2015).
Shanmugam, A.K. & Nesvizhskii, A.I. J. Proteome Res. 14, 5169–5178 (2015).
Acknowledgements
The authors wish to thank numerous colleagues including S. Eluik, G. Tan, X. Sun, X. Liu (Thermo Fisher Scientific), A. Hubauer, J. Mergner, J. Zecha, P. Samaras and the entire Kuster team as well as H. Hahne (OmicScouts), R. Weise, D. Riehn, K. Schrödter, F. Schumacher, N. Kolls and R.A. Castro-Alvaro (JPT) for fruitful discussions and technical assistance. We thank D. Campbell and Z. Sun for their efforts and the entire Moritz group at ISB and the Aebersold group at ETH Zurich for their efforts in peptide selection and synthesis. This work was in part funded by the German Federal Ministry of Education and Research (BMBF; grant no. 031L0008A). A postdoctoral fellowship from the Alexander von Humboldt Foundation (to P.Y.) is also gratefully acknowledged. This work was also performed in part with federal funds from the American Recovery and Reinvestment Act (ARRA) funds through the NIH, from the National Human Genome Research Institute grant no. RC2HG005805 (to R.L.M.); the National Institute of General Medical Sciences grant nos. R01GM087221, S10RR027584 and 2P50 GM076547/Center for Systems Biology (to R.L.M.); the European Research Council grant nos. ERC-2008-AdG 233226 and ERC-2014-AdG 670821; and the Swiss National Science Foundation (grant no. 31003A-130530 to R.A.), and DAAD (fellowship to U.K.).
Author information
Authors and Affiliations
Contributions
R.A., H.W., T.M., A.H., U.R. and B.K. conceived the study. D.P.Z., M. Wilhelm, K.S., J.Z., T.K., B.D., K.K., U.K., R.L.M. and B.K. designed experiments. D.P.Z., M. Wilhelm, K.S., J.Z., T.K., B.D., D.J.B., P.Y., K.K., E.W.D. and T.S. performed the experiments and analyzed data. M. Wilhelm, S.G., H.-C.E., M. Weininger, J.S., T.S. and S.A. extended the web resource. D.P.Z., M. Wilhelm and B.K. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
M. Wilhelm and B.K. are founders and shareholders of OmicScouts, which operates in the field of proteomics. They have no operational role in the company. R.A. is a cofounder and shareholder of Biognosys, which operates in the field of proteomics. He has no operational involvement in the company. K.S., J.Z., T.K., H.W. and U.R. are employees of JPT. B.D., D.J.B, T.M. and A.H. are employees of Thermo Fisher Scientific. S.G., H.-C.E., J.S. and S.A. are employees of SAP SE. Retention time peptides will be commercialized by JPT.
Integrated supplementary information
Supplementary Figure 1 Schematic representation of the peptide pool design process for the SRMAtlas peptide set
Starting with peptides in individual cavities in 96 well plates, peptides were manually pooled to form a mixture of 95 peptides (a plate pool). To create measurement pools of ~1,000 peptides, either 10 (for tryptic peptides) or 14 (non-tryptic peptides) plate pools were combined. To avoid bias in any peptide pool towards a particular MW (molecular weight) or HI (hydrophobicity index), a pooling scheme was computed to best mimic the overall MW and HI distribution of the entire set. Starting with a particular pool (top left panel; black line, here plate pool 1), all the remaining plate pools were tested in silico to generate a combined mixture, where MW or HI would best resemble the overall set (blue line). After determining the best next plate pool to use (here plate pool 313), the resulting mixture was tested again (middle panel) and the process was repeated until the desired number of plate pools was reached (using an iterative greedy approach). The resulting MW and HI distribution (black line) in comparison to that of the total set (blue line) is shown in the bottom right panel. In the example shown, a near perfect overlay of HI and a good approximation of MW distributions was achieved.
Supplementary Figure 2 Determination of retention times and retention time indices
(a) Retention time stability of 71 selected retention time standard peptides (66 RT peptides + 5 QC peptides) across ~1,200 LC-MS/MS runs (identification in n runs and median indicated). The median standard deviation of observed retention time differences for individual peptides was 48.7 seconds without (middle panel) and 12.4 seconds with retention time adjustment (lower panel) using RT indices calculated based on the peptides ISLGEHEGGGK and YDTAIDFGLFK. Data were filtered for Andromeda scores of >100. (b) The scatter plot of two retention time indices calculated based on different reference peptides shows that a conversion between them is possible without losing accuracy (R2 = 0.9996). Retention time index 1 (iRT1) is calculated based on the early eluting peptide ISLGEHEGGGK and late eluting peptide YDTAIDFGLFK (same as in (a); indicated by purple dashed lines). Retention time index 2 (iRT2) is calculated based on SYASDFGSSAK and GFVIDDGLITK (red dashed line). Each dot represents one of the 71 selected peptides identified in one of the ~1200 LC-MS/MS runs. (c) Similar to (b), here a third retention time index (iRT3) was calculated based on GSGGFTEFDLK and LTDELLSEYYK (orange dashed line) which span only a narrow part of the gradient. The linear fit shows that retention time indices can still be converted with very high accuracy (R2 = 0.9977) indicating that any high confident identifications (not necessarily peptides used for retention time calculation) can be used for retention time index calculation and thus conversion.
Supplementary Figure 3 LC-MS Data acquisition scheme and qualitative peptide identification analysis
(a) Data acquisition scheme used for all peptide pools. After an initial 1h survey run using HCD fragmentation with Orbitrap readout and CID fragmentation with ion trap readout, an inclusion list was generated. The three subsequent LC-MS runs from every pool utilized the inclusion list to target fully synthesized peptides by the indicated fragmentation techniques and collision energies. (b) Violin plots of the average Andromeda score (top panel), average number of matched fragment ions (middle panel) and the average intensity that could be explained by Andromeda in the tandem MS spectra (lower panel). Only identifications with an Andromeda score >100 were considered here. The numbers on top of each violin indicates the number of peptides.
Supplementary Figure 4 Peptide identifications (score >100) across different acquisition methods
(a) Venn diagram for the comparison of identifications with an Andromeda Score >100 across five major fragmentation types. The upper bar chart displays the number of peptide identifications for each fragmentation type, the lower bar chart shows peptide identifications only if the corresponding method reached a score of at least 90% of the highest score observed for that peptide (indicating the number of peptides for which the respective fragmentation technique gave the best identification result). We note that even though the various ETD versions were less successful than CID or HCD, there are still thousands of peptides for which ETD is the best fragmentation technique. (d) Venn diagram for the comparison of identifications using different HCD collision energies with an Andromeda Score >100. The upper bar chart displays peptide identifications for every collision energy, the lower bar chart shows peptide identifications only if the corresponding collision energy experiment reached a score of at least 90% of the highest score observed for that peptide.
Supplementary Figure 5 Andromeda score distributions for different peptide precursor charge states and the 11 tandem MS methods used in this study
Violin plots of the maximum Andromeda score distributions for peptides over the respective charge states. The number of peptide sequences is indicated above every violin, the median score is indicated as a white circle inside the violin. As expected, ETD based fragmentation techniques gives good results for peptides with higher charge states. Interestingly, resonance and beam type CID still yield higher absolute peptide identifications at any charge state but ETD often generates higher identification scores for peptides of higher charge.
Supplementary Figure 6 HCD fragmentation spectra of YYLIQLLEDDAQR with Orbitrap readout at different collision energies.
Fragmentation spectra of the peptide YYLIQLLEDDAQR for all six HCD methods used in this study (normalized collision energies of NCE 20, 23, 25, 28, 30, 35 respectively). All annotated spectra are the best identification from Andromeda (i.e. highest score) for the respective fragmentation mode.
Supplementary Figure 7 Fragmentation spectra of YYLIQLLEDDAQR using resonance type CID, HCD and versions of ETD
Fragmentation spectra of the peptide YYLIQLLEDDAQR for ETD, EThcD and ETciD (all Orbitrap readout) as well as HCD with 28 NCE with ion trap readout and CID with 35 NCE and ion trap readout. All annotated spectra are the best identification from Andromeda (i.e. highest score) for the respective fragmentation mode.
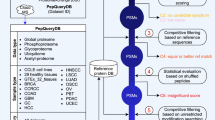
Supplementary Figure 8 Logistics of data handling and effect of different database search strategies
(a) Schematic representation of the data handling pipeline governed by the internal pipeline/database used for the ProteomeTools project. After pool design and peptide synthesis, an initial survey acquisition run followed by an automatic MaxQuant search was used to identify the desired full length peptides. The results were imported into the internal database which then automatically prepared the acquisition methods for the HCD, IT and ETD acquisition runs (see Supplementary Information for details). These subsequent acquisitions were again automatically searched and imported into the database for quality control and data organization. (b) Comparison of database searches for peptide identification. Upper panel: Analysis of 20 pools from the “proteotypic” set in separate searches or searched together (combined). It is evident that shorter peptide identifications are lost when combining peptide pools for database searching. Lower panel: Analysis of 96 pools from the “proteotypic” set, searched either with tryptic or unspecific digestion of the database. It is evident that searching without tryptic specificity results in lower peptide identifications. We note that both these are issues of current database search algorithms that need addressing.
Supplementary Figure 9 Success of full length peptide identifications in the three peptides sets generated in this study
We measured the success of each synthesis by determining the fraction of peptides in a pool that could be identified by LC-MS/MS (the different fragmentation modes are indicated in each plot, all HCD collision energies were combined). Apart from a 1% peptide FDR, no additional score cutoff was applied here. For the ‘proteotypic’ set (top panel), recoveries are generally very high (average ~95 %) and only decrease for very long peptides (high pool numbers) presumably because it becomes increasingly difficult to obtain a full length peptide. For the ‘missing gene’ set (middle panel), recoveries were lower (average ~80 %) likely because of lower success in the LC-MS/MS analysis (e. g. solubility, ionization efficiency, fragmentation efficiency). We note that this was expected given the fact that these peptides were predicted from the protein sequences regardless of any prior observation from biological sources. The recovery of the ‘SRMAtlas’ set (bottom panel) was also lower (average ~65 %) possibly (among other potential factors) because these peptides had been synthesized ~6 years prior to our analysis and because this set contains peptides representing N-linked glycosylation sites after PNGase F digestion which we did not account for in the database search.
Supplementary Figure 10 Assessment of peptide purity and side product profile
(a) Using an unspecific MaxQuant search with the “depended peptides” option enabled, a synthesis tree view of the peptide ESQLKDLEAENRR was constructed that displays the estimated relative yield of the desired full length peptide product (85%) as well as other side products in the synthesis. (b) Same as panel (a) but for the peptide LVFVDAVAFLTGK what displays an estimated relative yield of the desired full length peptide product of 52%. The tree lists all identified truncation and by-products and their relative contribution to the entire signal intensity attributable to these molecular species. By-products with less than 1% estimated yield are omitted from the visualization fro clarity. Annotation from bottom to top: Peptide sequence identified, potential modification, mass error to annotated modification in ppm, delta mass compared to the full length peptide (e. g. mass of missing amino acid or additional protection group) and percentage of the total intensity of the identified synthesis products. The correct full length product is marked in green.
Supplementary Figure 11 Comparison of Orbitrap Lumos beam-type CID spectra with QTOF beam-type CID spectra
(a) Bar chart showing the total (red) number of Lumos spectra matched against the QTOF (5600 TripleTOF) spectrum library (Guo et al.) and the number of best matching spectra (blue) for a particular normalized collision energy (NCE). Data were filtered for an Andromeda score of > 100. (b) Histograms of Pearson spectrum correlations between QTOF spectra and the corresponding Orbitrap Fusion Lumos spectra acquired at different NCEs. The highest median correlation is observed at 28 and 30 NCE. (c) Boxplot of Pearson spectrum correlation coefficients between spectra acquired on a 5600 TripleTOF mass spectrometer and the best matching spectra acquired on an Orbitrap Fusion Lumos at different Andromeda scores. While the analysis in (a) and throughout the manuscript used a conservative score cutoff of 100, the distribution of correlation coefficients here suggests that spectra with an Andromeda score between 60-100 are also suitable as reference spectra. (d) Example for low correlating spectra: experimental beam-type CID QTOF mass spectrum of the peptide ILIEDSDQNLK/2+ (top) compared to the corresponding beam-type CID spectrum at 28 NCE of the synthesized reference peptide standard acquired on an Orbitrap Fusion Lumos (Andromeda score 171). Both, the low signal-to-noise and near zero Pearson correlation suggest that the upper spectrum is a false positive in the QTOF library. (e) Example for high correlating spectra: experimental beam-type CID QTOF mass spectrum of the peptide EVGTPHGIILDSVDAAFICPGSSR/3+ (top) compared to the corresponding beam-type CID spectrum at 35 NCE of the synthesized reference peptide standard (bottom) acquired on an Orbitrap Fusion Lumos (Andromeda score 169) showing very good overall agreement.
Supplementary Figure 12 Properties of amino acids and fragmentation efficiency of amino acid pairs across different HCD collision energies
(a-f) Heatmap of median y-ion fragment intensity ranks at 20 (a), 23 (b), 25 (c), 28 (d), 30 (e) and 35 (f) normalized collision energy (NCE) of all possible amino acids combinations N-terminal and C-terminal of the fragmentation position. Note that rank 1 (dark blue) corresponds to the highest fragment ion intensity in a spectrum. The top three rows indicate charge, polarity and class of the amino acid. In case the fragmentation occurs N-terminal of P (row), high intense fragments are generated. In contrast, if the fragmentation occurs C-terminal of G (column), generally low intense fragments are generated.
Supplementary Figure 13 Generation of collision energy-specific fragmentation prediction models based on multiple spectra of peptides
(a-d) Plots showing the relative intensity of y-type fragment ions occurring between particular amino acid pairs (here Q-R in (a), Y-L in (b), D-D in (c) and D-P in (d)) as a function of the relative position of the y-ion within the peptide sequence (0 = C-terminus; 1 = N-terminus). For each amino acid pair N- and C-terminal of the fragmentation position, a normalized collision energy-dependent LOESS regression (red line) was used to model the relative fragment length (y-ion divided by total peptide length) and relative fragment intensity (normalized to base peak intensity of the MS2 spectrum) for later prediction. The number of observations of each fragment ion are shown in blue (the darker, the more observations). The shape of the LOESS fits varies greatly between different normalized collision energies, relative positions and amino acids pairs indicating vastly different fragmentation behaviors. For example, the pair Q-R (fragmentation C-terminal of Q, but N-terminal of R) shows only low intensity and low mass y-ions at low collision energies but increasing to almost 20% relative intensity at higher collision energies. In fully cleaved tryptic peptides, Q-R occurs very rarely, hence there are only few occurrences in the plot. Other amino acid combinations are much more frequent and, therefore lead to much more data in each plot. This information was used to train models predicting the fragment ion intensity of peptides given their amino acid sequence.
Supplementary Figure 14 Prediction of fragmentation spectra based on y-ion collision energy-specific fragmentation models
Panels (a) and (b) show examples for the prediction of fragmentation patterns for two different peptides at different collision energies. The upper two panels in each plot show the observed median relative fragment ion intensities of y-fragments across six different collision energies. The box plots in the panel below display the reproducibility and number of observations (number on top) of the relative intensity of the different y-ions across all acquired tandem mass spectra for this peptide and a given normalized collision energy (here 20, 30 and 35). The panels below show the predicted y-ion fragment spectra for each normalized collision energy including the Pearson correlation between the predicted and observed spectrum. (c) Histogram of Pearson correlations between predicted and observed spectrum at (left to right) 20, 23, 25, 28, 30 and 35 normalized collision energy (NCE). It is evident, that our classifier can correctly predict the intensity of fragment ions within a tandem mass spectrum in most cases.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14 and Supplementary Notes 1–4 (PDF 3148 kb)
Supplementary Table 1
Peptide pool mapping (XLSX 18030 kb)
Supplementary Table 2
Peptide identifications (XLSX 62969 kb)
Supplementary Table 3
Overlap with mouse (XLSX 4467 kb)
Rights and permissions
About this article
Cite this article
Zolg, D., Wilhelm, M., Schnatbaum, K. et al. Building ProteomeTools based on a complete synthetic human proteome. Nat Methods 14, 259–262 (2017). https://doi.org/10.1038/nmeth.4153
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4153
This article is cited by
-
Accurate de novo peptide sequencing using fully convolutional neural networks
Nature Communications (2023)
-
Prediction of peptide mass spectral libraries with machine learning
Nature Biotechnology (2023)
-
Peptide collision cross sections of 22 post-translational modifications
Analytical and Bioanalytical Chemistry (2023)
-
False discovery rate estimation using candidate peptides for each spectrum
BMC Bioinformatics (2022)
-
A learned embedding for efficient joint analysis of millions of mass spectra
Nature Methods (2022)