Abstract
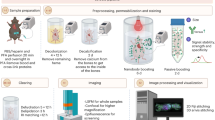
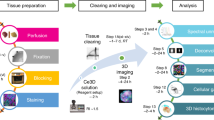
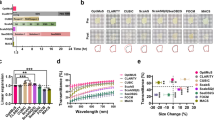
Recent tissue-clearing approaches have become important alternatives to standard histology approaches. However, light scattering in thick tissues and the size restrictions on samples that can be imaged with standard light-sheet microscopy pose limitations for analyzing large samples such as an entire rodent body. We developed 'ultimate DISCO' (uDISCO) clearing to overcome these limitations in volumetric imaging. uDISCO preserves fluorescent proteins over months and renders intact organs and rodent bodies transparent while reducing their size up to 65%. We used uDISCO to image neuronal connections and vasculature from head to toe over 7 cm and to perform unbiased screening of transplanted stem cells within the entire body of adult mice. uDISCO is compatible with diverse labeling methods and archival human tissue, and it can readily be used in various biomedical applications to study organization of large organ systems throughout entire organisms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Yang, B. et al. Single-cell phenotyping within transparent intact tissue through whole-body clearing. Cell 158, 945–958 (2014).
Hama, H. et al. ScaleS: an optical clearing palette for biological imaging. Nat. Neurosci. 18, 1518–1529 (2015).
Chung, K. et al. Structural and molecular interrogation of intact biological systems. Nature 497, 332–337 (2013).
Tainaka, K. et al. Whole-body imaging with single-cell resolution by tissue decolorization. Cell 159, 911–924 (2014).
Ertürk, A. et al. Three-dimensional imaging of the unsectioned adult spinal cord to assess axon regeneration and glial responses after injury. Nat. Med. 18, 166–171 (2011).
Ertürk, A. et al. Three-dimensional imaging of solvent-cleared organs using 3DISCO. Nat. Protoc. 7, 1983–1995 (2012).
Kuwajima, T. et al. ClearT: a detergent- and solvent-free clearing method for neuronal and non-neuronal tissue. Development 140, 1364–1368 (2013).
Susaki, E.A. et al. Whole-brain imaging with single-cell resolution using chemical cocktails and computational analysis. Cell 157, 726–739 (2014).
Ke, M.T., Fujimoto, S. & Imai, T. SeeDB: a simple and morphology-preserving optical clearing agent for neuronal circuit reconstruction. Nat. Neurosci. 16, 1154–1161 (2013).
Susaki, E.A. et al. Advanced CUBIC protocols for whole-brain and whole-body clearing and imaging. Nat. Protoc. 10, 1709–1727 (2015).
Richardson, D.S. & Lichtman, J.W. Clarifying tissue clearing. Cell 162, 246–257 (2015).
Tuchin, V.V. Tissue optics and photonics: light-tissue interaction. Journal of Biomedical Photonics & Engineering 1, 98–134 (2015).
Lichtman, J.W. & Conchello, J.A. Fluorescence microscopy. Nat. Methods 2, 910–919 (2005).
Dodt, H.U. et al. Ultramicroscopy: three-dimensional visualization of neuronal networks in the whole mouse brain. Nat. Methods 4, 331–336 (2007).
Susaki, E.A. & Ueda, H.R. Whole-body and whole-organ clearing and imaging techniques with single-cell resolution: toward organism-level systems biology in mammals. Cell Chem. Biol. 23, 137–157 (2016).
Liu, Z. et al. Immune homeostasis enforced by co-localized effector and regulatory T cells. Nature 528, 225–230 (2015).
Espinosa-Medina, I. et al. Neurodevelopment. Parasympathetic ganglia derive from Schwann cell precursors. Science 345, 87–90 (2014).
Oshimori, N., Oristian, D. & Fuchs, E. TGF-β promotes heterogeneity and drug resistance in squamous cell carcinoma. Cell 160, 963–976 (2015).
Lafkas, D. et al. Therapeutic antibodies reveal Notch control of transdifferentiation in the adult lung. Nature 528, 127–131 (2015).
Renier, N. et al. iDISCO: a simple, rapid method to immunolabel large tissue samples for volume imaging. Cell 159, 896–910 (2014).
Belle, M. et al. A simple method for 3D analysis of immunolabeled axonal tracts in a transparent nervous system. Cell Rep. 9, 1191–1201 (2014).
Renier, N. et al. Mapping of brain activity by automated volume analysis of immediate early genes. Cell 165, 1789–1802 (2016).
Feng, G. et al. Imaging neuronal subsets in transgenic mice expressing multiple spectral variants of GFP. Neuron 28, 41–51 (2000).
Schwarz, M.K. et al. Fluorescent-protein stabilization and high-resolution imaging of cleared, intact mouse brains. PLoS One 10, e0124650 (2015).
Treweek, J.B. et al. Whole-body tissue stabilization and selective extractions via tissue-hydrogel hybrids for high-resolution intact circuit mapping and phenotyping. Nat. Protoc. 10, 1860–1896 (2015).
Ascenzi, A. & Fabry, C. Technique for dissection and measurement of refractive index of osteones. J. Biophys. Biochem. Cytol. 6, 139–142 (1959).
Genina, E.A., Bashkatov, A.N. & Tuchin, V.V. Optical clearing of cranial bone. Adv. Opt. Technol. 2008, 1–8 (2008).
De Miguel, M.P. et al. Immunosuppressive properties of mesenchymal stem cells: advances and applications. Curr. Mol. Med. 12, 574–591 (2012).
D'souza, N. et al. Mesenchymal stem/stromal cells as a delivery platform in cell and gene therapies. BMC Med. 13, 186 (2015).
Guenoun, J. et al. In vivo quantitative assessment of cell viability of gadolinium or iron-labeled cells using MRI and bioluminescence imaging. Contrast Media Mol. Imaging 8, 165–174 (2013).
Leibacher, J. & Henschler, R. Biodistribution, migration and homing of systemically applied mesenchymal stem/stromal cells. Stem Cell Res. Ther. 7, 7 (2016).
Nemeth, K., Mayer, B., Sworder, B.J., Kuznetsov, S.A. & Mezey, E. A practical guide to culturing mouse and human bone marrow stromal cells. Curr. Protoc. Immunol. 102, Unit 22F.12 (2013).
Rosen, A.B. et al. Finding fluorescent needles in the cardiac haystack: tracking human mesenchymal stem cells labeled with quantum dots for quantitative in vivo three-dimensional fluorescence analysis. Stem Cells 25, 2128–2138 (2007).
Goldmacher, G.V. et al. Tracking transplanted bone marrow stem cells and their effects in the rat MCAO stroke model. PLoS One 8, e60049 (2013).
Detante, O. et al. Intravenous administration of 99mTc-HMPAO-labeled human mesenchymal stem cells after stroke: in vivo imaging and biodistribution. Cell Transplant. 18, 1369–1379 (2009).
Chen, F., Tillberg, P.W. & Boyden, E.S. Optical imaging. Expansion microscopy. Science 347, 543–548 (2015).
Chozinski, T.J. et al. Expansion microscopy with conventional antibodies and fluorescent proteins. Nat. Methods 13, 485–488 (2016).
Chen, F. et al. Nanoscale imaging of RNA with expansion microscopy. Nat. Methods 13, 679–684 (2016).
Treweek, J.B. & Gradinaru, V. Extracting structural and functional features of widely distributed biological circuits with single cell resolution via tissue clearing and delivery vectors. Curr. Opin. Biotechnol. 40, 193–207 (2016).
Chen, B.C. et al. Lattice light-sheet microscopy: imaging molecules to embryos at high spatiotemporal resolution. Science 346, 1257998 (2014).
Bareyre, F.M. et al. The injured spinal cord spontaneously forms a new intraspinal circuit in adult rats. Nat. Neurosci. 7, 269–277 (2004).
Wahl, A.S. et al. Neuronal repair. Asynchronous therapy restores motor control by rewiring of the rat corticospinal tract after stroke. Science 344, 1250–1255 (2014).
Reinhardt, R.L., Khoruts, A., Merica, R., Zell, T. & Jenkins, M.K. Visualizing the generation of memory CD4 T cells in the whole body. Nature 410, 101–105 (2001).
Pan, C., Cai, R., Quacquarelli, F.P., Ghasemi, A. & Ertürk, A. Whole organ and organism tissue clearing by uDISCO. Protocol Exchange http://dx.doi.org/10.1038/protex.2016.055 (2016).
Kilkenny, C., Browne, W.J., Cuthill, I.C., Emerson, M. & Altman, D.G. Improving bioscience research reporting: the ARRIVE guidelines for reporting animal research. PLoS Biol. 8, e1000412 (2010).
Acar, M. et al. Deep imaging of bone marrow shows non-dividing stem cells are mainly perisinusoidal. Nature 526, 126–130 (2015).
Ertürk, A., Lafkas, D. & Chalouni, C. Imaging cleared intact biological systems at a cellular level by 3DISCO. J. Vis. Exp. 89, e51382 (2014).
Zacharaki, D. et al. Characterization of in vitro expanded bone marrow-derived mesenchymal stem cells isolated from experimental autoimmune encephalomyelitis mice. J. Mol. Neurosci. 51, 282–297 (2013).
Preibisch, S., Saalfeld, S. & Tomancak, P. Globally optimal stitching of tiled 3D microscopic image acquisitions. Bioinformatics 25, 1463–1465 (2009).
Pietzsch, T., Preibisch, S., Tomancák, P. & Saalfeld, S. ImgLib2–generic image processing in Java. Bioinformatics 28, 3009–3011 (2012).
Jahr, W., Schmid, B., Schmied, C., Fahrbach, F.O. & Huisken, J. Hyperspectral light sheet microscopy. Nat. Commun. 6, 7990 (2015).
Gonzalez, R. Digital Image Processing 3rd edn. (India, Pearson Prentice Hall, 2006).
Arganda-Carreras, I. et al. Consistent and elastic registration of histological sections using vector-spline regularization. in Computer Vision Approaches to Medical Image Analysis (eds. Beichel, R.R. & Sonka, M.) 85–95 (Springer, 2006).
Acknowledgements
This work was supported by the Vascular Dementia Research Foundation, Synergy Excellence Cluster Munich (SyNergy), ERA-Net Neuron (01EW1501A to A.E. and N.P.), and the European Union's Horizon 2020 research and innovation programme (grant agreement no. 666881, SVDs@target, M.D.). A.L. and N.P. were supported by a Marie Curie Intra European Fellowship grant (FP7-PEOPLE-2013-IEF, project no. 625970). We thank M. Hübener and F. Voss (Max Planck Institute of Neurobiology, Munich) for providing mice; D. Trauner and O. Thorn-Seshold for helpful discussions; A. Weingart for illustrations; and C. Hojer, S. Tappan and T. Misgeld for critical reading of the manuscript. C.P. and R.C. are members of the Graduate School of Systemic Neurosciences (GSN), Ludwig Maximilian University of Munich. Human tissues were provided by the brain bank of the Institute of Anatomy, University of Leipzig.
Author information
Authors and Affiliations
Contributions
A.E. designed and led all aspects of the project. C.P., R.C., and F.P.Q. performed most of the experiments. A.G. performed the image rendering and developed algorithms for data analysis. C.P., R.C., F.P.Q., and A.G. analyzed the data. A.L. interpreted data and performed the BMSC cultures, characterization, and transplantations; F.H. performed virus tracing; P.M. assisted first-clearing experiments; N.P. and M.D. supervised A.L. and F.H., respectively. A.E., C.P., R.C., F.P.Q., and A.G. wrote the paper. All authors edited the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–30, Supplementary Tables 1 and 2, and Supplementary Protocol (PDF 15451 kb)
3D visualization of microglia in CX3CR1-EGFP mouse brain
A ~3 months old CX3CR1-EGFP mouse brain was cleared with uDISCO and imaged with the light-sheet microscopy using the 4x corrected objective. The individual microglia throughout the entire brain are evident. (MOV 21448 kb)
uDISCO clearing of limbs and bones
Surface reconstruction and 3D maximum intensity projection of a paw with forearm from 4 months old C57BL/6N mouse labeled with Texas Red Dextran after whole-body uDISCO clearing. uDISCO renders the bones fully transparent, allowing the visualization of the vascular details in the entire limbs. (MOV 12455 kb)
Details of vasculature in the forelimb
3D visualization and orthoslice of the forearm with vasculature labeling in Supplementary Video 2. Fine details of the vasculature are clear throughout the entire scan. (MOV 3988 kb)
3D rendering of vasculature in the entire rat CNS
The vasculature of a 4-weeks old rat was labeled with Texas Red Dextran. We imaged the entire CNS (the brain plus spinal cord) after whole-body clearing of the rat. Both the large and small vessels are clearly visible throughout the entire scan of ≥13 cm CNS rat tissue. (MOV 15658 kb)
Head-to-limb imaging of the nervous system
3D visualization of neuronal connections throughout the intact mouse CNS in Figure 4 (GFP-M mouse, 4 months old). The fine details of the neuronal connections are evident from head to the nerves invading the hind limbs. (MOV 29200 kb)
Tracing of the CNS axons over several centimeters
A single traced axons (red) in the spinal cord of the sample from Figure 4 is shown. The trajectories of individual axons in the entire CNS of the adult mice can be determined. Note that occasional curling of long spinal cord axons does not interfere with the tracing of their entire trajectories. The pseudo-colored traced axon is shown thicker than its actual size for easier visualization. (MOV 10308 kb)
3D visualization of neuronal connections in the intact brain in Figure 4
A GFP-M mouse brain (4 months old) was cleared with uDISCO and imaged with the light-sheet microscopy. The whole-brain images were obtained using the 4x corrected objective, and high-magnification images in the second part of the video were obtained using the 20x corrected objective. (MOV 29413 kb)
Light-sheet microscopy images of the dendritic spines after uDISCO clearing
A light-sheet microscopy stack acquired on uDISCO cleared GFP-M mouse brain at 0.5 – 1 mm depth using Zeiss CLARITY objective (25x, NA 1.0), which was directly immersed into BABB-D. The individual dendritic spines are evident throughout the scan. (MOV 1672 kb)
Visualization of Qdot-positive BMSCs IV-injected lung
The video shows the 3D reconstruction of the densely distributed BMSCs throughout the entire lung. uDISCO rendered the intact lungs fully transparent, allowing to assess the distribution of individual BMSCs. (MOV 23884 kb)
Rights and permissions
About this article
Cite this article
Pan, C., Cai, R., Quacquarelli, F. et al. Shrinkage-mediated imaging of entire organs and organisms using uDISCO. Nat Methods 13, 859–867 (2016). https://doi.org/10.1038/nmeth.3964
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3964
This article is cited by
-
Whole-body cellular mapping in mouse using standard IgG antibodies
Nature Biotechnology (2024)
-
Current and future applications of light-sheet imaging for identifying molecular and developmental processes in autism spectrum disorders
Molecular Psychiatry (2024)
-
Reflective multi-immersion microscope objectives inspired by the Schmidt telescope
Nature Biotechnology (2024)
-
Pathological hemodynamic changes and leukocyte transmigration disrupt the blood–spinal cord barrier after spinal cord injury
Journal of Neuroinflammation (2023)
-
Three-dimensional visualization of neural networks inside bone by Osteo-DISCO protocol and alteration of bone remodeling by surgical nerve ablation
Scientific Reports (2023)