Abstract
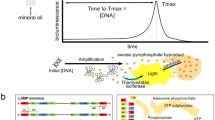
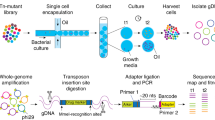
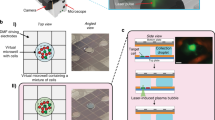
We have developed hydrogel-based virtual microfluidics as a simple and robust alternative to complex engineered microfluidic systems for the compartmentalization of nucleic acid amplification reactions. We applied in-gel digital multiple displacement amplification (dMDA) to purified DNA templates, cultured bacterial cells and human microbiome samples in the virtual microfluidics system, and demonstrated whole-genome sequencing of single-cell MDA products with excellent coverage uniformity and markedly reduced chimerism compared with products of liquid MDA reactions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Blainey, P.C. & Quake, S.R. Nat. Methods 11, 19–21 (2014).
Marshall, I.P.G., Blainey, P.C., Spormann, A.M. & Quake, S.R.A. Appl. Environ. Microbiol. 78, 8555–8563 (2012).
Wang, J., Fan, H.C., Behr, B. & Quake, S.R. Cell 150, 402–412 (2012).
Huggett, J.F., Cowen, S. & Foy, C.A. Clin. Chem. 61, 79–88 (2015).
Sykes, P.J. et al. Biotechniques 13, 444–449 (1992).
Vogelstein, B. & Kinzler, K.W. Proc. Natl. Acad. Sci. USA 96, 9236–9241 (1999).
Blainey, P.C. & Quake, S.R. Nucleic Acids Res. 39, e19 (2011).
Raghunathan, A. et al. Appl. Environ. Microbiol. 71, 3342–3347 (2005).
Zhang, K. et al. Nat. Biotechnol. 24, 680–686 (2006).
Fu, Y. et al. Proc. Natl. Acad. Sci. USA 112, 11923–11928 (2015).
Pamp, S.J., Harrington, E.D., Quake, S.R., Relman, D.A. & Blainey, P.C. Genome Res. 22, 1107–1119 (2012).
Dodsworth, J.A. et al. Nat. Comm. 4, 1854 (2013).
Hess, M. et al. Science 331, 463–467 (2011).
Love, K.R., Bagh, S., Choi, J. & Love, J.C. Trends Biotechnol. 31, 280–286 (2013).
Thorsen, T., Maerkl, S.J. & Quake, S.R. Science 298, 580–584 (2002).
Landry, Z.C., Giovanonni, S.J., Quake, S.R. & Blainey, P.C. Methods Enzymol. 531, 61–90 (2013).
de Bourcy, C.F.A. et al. PLoS ONE 9, e105585 (2014).
Marcy, Y. et al. PLoS Genet. 3, 1702–1708 (2007).
Thorsen, T., Roberts, R.W., Arnold, F.H. & Quake, S.R. Phys. Rev. Lett. 86, 4163–4166 (2001).
Mazutis, L. et al. Nat. Protoc. 8, 870–891 (2013).
Hindson, C.M. et al. Nat. Methods 10, 1003–1005 (2013).
Morinishi, L.S. & Blainey, P. J. Vis. Exp. 103, e52925 (2015).
Blainey, P.C. FEMS Microbiol. Rev. 37, 407–427 (2013).
Podar, M., Keller, M. & Hugenholtz, P. in Uncultivated Microorganisms, Vol. 10 (ed. Epstein, S.S.) 241–256 (Springer, Berlin Heidelberg, 2009).
Mitra, R.D. & Church, G.M. Nucleic Acids Res. 27, e34 (1999).
Allen, L.Z. et al. PLoS ONE 6, e17722 (2011).
Raeber, G.P., Lutolf, M.P. & Hubbell, J.A. Biophys. J. 89, 1374–1388 (2005).
Wu, Y., Joseph, S. & Aluru, N.R. J. Phys. Chem. B 113, 3512–3520 (2009).
Phelps, E.A. et al. Adv. Mater. 24, 64–70 (2011).
Dean, F.B. et al. Proc. Natl. Acad. Sci. USA 99, 5261–5266 (2002).
Marcy, Y. et al. Proc. Natl. Acad. Sci. USA 104, 11889–11894 (2007).
Woyke, T. et al. PLoS ONE 6, e26161 (2011).
Lasken, R.S. & Stockwell, T.B. BMC Biotechnol. 7, 19 (2007).
Segata, N. et al. Nat. Methods 9, 811–814 (2012).
Wu, M. & Scott, A.J. Bioinformatics 28, 1033–1034 (2012).
Acknowledgements
The authors thank S. Kim, D. Feldman and A. Kulesa for advice on bioinformatics, microscopy and image analysis; N. Ranu and the Hung lab (Broad Institute) for bacterial samples; L. Griffith, G. Lagoudas, J. Borrajo and L. Morinishi for helpful discussions; and members of the Griffith lab (MIT), especially C. Chopko, J. Valdez and H. Lee, for hydrogel expertise. This work was supported in part by a Lawrence Summers Fellowship from the Broad Institute (L.X.), a Career Award at the Scientific Interface from the Burroughs Welcome Fund (P.C.B.) and grants from the Center for Microbiome Informatics and Therapeutics at MIT; a National Human Genome Research Institute, grant number U54HG003067, to the Broad Institute; the Center for Environmental Health Sciences at MIT and the Fijian Ministry of Health. The Broad Institute and MIT may seek to commercialize aspects of this work, and related applications for intellectual property have been filed.
Author information
Authors and Affiliations
Contributions
P.C.B. and L.X. conceived the concept for this study. L.X. designed and implemented experiments and conducted data analysis. I.L.B. and E.J.A. provided Fiji microbiome samples. I.L.B. conducted analysis of gut microbe data. L.X., P.C.B. and I.L.B. wrote and all authors approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The Broad Institute and MIT may seek to commercialize aspects of this work and related applications for intellectual property have been filed.
Integrated supplementary information
Supplementary Figure 1 Real time dMDA for quantification of cluster growth and count with time
The cluster radius curve shows mean radius ± SEM. The zero time point cluster count and radius data points reflect the properties of fluorescent contaminants.
Supplementary Figure 2 E. coli MDA cluster quantification
E. coli MDA cluster’s DNA content was calibrated against (unamplified) hydrogel-embedded mammalian cells (HEK 293) by comparing SYTOX Orange fluorescence intensities. DNA content in individual product clusters and mammalian cells was approximated by integrating pixel intensities under the same hydrogel and staining conditions. The observed integrated fluorescence was comparable across MDA product clusters and unamplified mammalian cells, leading us to conclude that the average DNA content per MDA product cluster was approximately equal to the average DNA content of an unsynchronized mammalian cell population, or on the order of 10 pg. The centerline represents the mean value.
Supplementary Figure 3 Digital single-molecule PCR in hydrogel (Lambda phage DNA)
Measured cluster number per field of view versus calculated cluster number based on template concentration. Reaction conditions are described in methods (PCR). The dotted line indicates the ideal situation when measured cluster number equals to the theoretical cluster number.
Supplementary Figure 4 MDA cluster size decreases with increasing DNA template concentration with the same gel condition
Data is shown as 5 % - 95 % box plot with outliers scattered and the centerline as the median. (n = 2 fields of view at each concentration, number of clusters for each field of view is: n = 43, 62, 89, 88, 191, 167, 321, 305, 478, 542, 711, 833).
Supplementary Figure 5 Cluster size and location correlation analysis
Five reference clusters with the largest radii and five reference clusters with the smallest radii were chosen among clusters in one field of view (100+ clusters) of one MDA hydrogel sample. a) By zooming out from selected clusters, the number of other clusters encountered at successively greater distances was plotted. b) The total volume of other clusters encountered was plotted. c) and d) zoom-in views of a) and b) at 0 to 32 µm from reference clusters. We hypothesized that large clusters consume local resources and thus, would reduce the number and/or size of surrounding clusters. Although slight enhancements in the number and size of clusters surrounding the set of small reference clusters versus the set of large reference clusters exist, the effect is very small. Error bars are SEM.
Supplementary Figure 6 NGS data analysis schematic for E. coli and S. aureus
The analysis workflow is shown with a combination of bioinformatic tools, python scripts and MATLAB scripts.
Supplementary Figure 7 Mapping single-cell genomes to references
Single-cell samples were mapped to E. coli and S. aureus reference sequences with mean percent pair aligned and standard deviation shown. (n = 5 for E. coli punches, n = 7 for S. aureus punches, n = 7 for negative punches).
Supplementary Figure 8 MDA chimera frequency with different insert sizes, E. coli and S. aureus data
a) 1 – 3 kb. b) 3 kb – 10 kb. c) Larger than 10 kb. Centerlines represent the mean value.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Tables 1–8 and Supplementary Notes 1–5 (PDF 3624 kb)
Supplementary Table 9
Features of 117 FijiCOMP single-cell assemblies. (XLSX 69 kb)
Supplementary Software
Supplementary software for data analysis (ZIP 28 kb)
Rights and permissions
About this article
Cite this article
Xu, L., Brito, I., Alm, E. et al. Virtual microfluidics for digital quantification and single-cell sequencing. Nat Methods 13, 759–762 (2016). https://doi.org/10.1038/nmeth.3955
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3955
This article is cited by
-
Agarose gel microcapsules enable easy-to-prepare, picolitre-scale, single-cell genomics, yielding high-coverage genome sequences
Scientific Reports (2022)
-
Validation of the application of gel beads-based single-cell genome sequencing platform to soil and seawater
ISME Communications (2022)
-
Examining horizontal gene transfer in microbial communities
Nature Reviews Microbiology (2021)
-
Single-cell genomics of uncultured bacteria reveals dietary fiber responders in the mouse gut microbiota
Microbiome (2020)
-
Microbial single-cell omics: the crux of the matter
Applied Microbiology and Biotechnology (2020)