Abstract
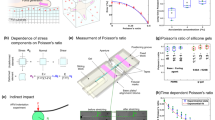
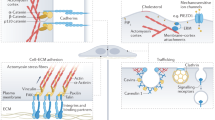
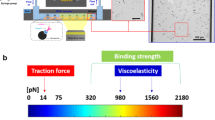
Forces generated by cells are critical regulators of cell adhesion, signaling, and function, and they are also essential drivers in the morphogenetic events of development. Over the past 20 years, several methods have been developed to measure these forces. However, despite recent substantial interest in understanding the contribution of these forces in biology, implementation and adoption of the developed methods by the broader biological community remain challenging because of the inherently multidisciplinary expertise required to conduct and interpret the measurements. In this review, we introduce the established methods and highlight the technical challenges associated with implementing each technique in a biological laboratory.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Bao, G. & Suresh, S. Cell and molecular mechanics of biological materials. Nat. Mater. 2, 715–725 (2003).
Bell, E., Ivarsson, B. & Merrill, C. Production of a tissue-like structure by contraction of collagen lattices by human fibroblasts of different proliferative potential in vitro. Proc. Natl. Acad. Sci. USA 76, 1274–1278 (1979).This study presents early observations of collagen hydrogel contraction as a measure of cell contractility.
Ehrlich, H.P. & Rajaratnam, J.B.M. Cell locomotion forces versus cell contraction forces for collagen lattice contraction: an in vitro model of wound contraction. Tissue Cell 22, 407–417 (1990).
Dallon, J.C. & Ehrlich, H.P. A review of fibroblast-populated collagen lattices. Wound Repair Regen. 16, 472–479 (2008).
Stopak, D. & Harris, A.K. Connective tissue morphogenesis by fibroblast traction. Dev. Biol. 90, 383–398 (1982).
Ngo, P., Ramalingam, P., Phillips, J.A. & Furuta, G.T. in Cell-Cell Interactions Vol. 341 (ed. Colgan, S.P.) 103–109 (Humana Press, 2006).
Smith, K.D., Wells, A. & Lauffenburger, D.A. Multiple signaling pathways mediate compaction of collagen matrices by EGF-stimulated fibroblasts. Exp. Cell Res. 312, 1970–1982 (2006).
Fernandez-Gonzalez, R. et al. Dynamics are regulated by tension in intercalating cells. Dev. Cell 17, 736–743 (2009).
Fernandez-Gonzalez, R. & Zallen, J.A. Wounded cells drive rapid epidermal repair in the early Drosophila embryo. Mol. Biol. Cell 24, 3227–3237 (2013).
Farhadifar, R., Röper, J.-C., Aigouy, B., Eaton, S. & Jülicher, F. The influence of cell mechanics, cell-cell interactions, and proliferation on epithelial packing. Curr. Biol. 17, 2095–2104 (2007).
Delvoye, P., Wiliquet, P., Leveque, J.-L., Nusgens, B.V. & Lapiere, C.M. Measurement of mechanical forces generated by skin fibroblasts embedded in a three-dimensional collagen gel. J. Invest. Dermatol. 97, 898–902 (1991).
Zimmermann, W.H. et al. Three-dimensional engineered heart tissue from neonatal rat cardiac myocytes. Biotechnol. Bioeng. 68, 106–114 (2000).This study combined the use of tissue contraction and cantilevers to assemble engineered tissues and allow force measurement.
Dennis, R.G. & Kosnik, P.E. Excitability and isometric contractile properties of mammalian skeletal muscle constructs engineered in vitro. In Vitro Cell. Dev. Biol. Anim. 36, 327–335 (2000).
Vandenburgh, H. et al. Automated drug screening with contractile muscle tissue engineered from dystrophic myoblasts. FASEB J. 23, 3325–3334 (2009).
Vandenburgh, H. et al. Drug-screening platform based on the contractility of tissue-engineered muscle. Muscle Nerve 37, 438–447 (2008).
Hansen, A. et al. Development of a drug screening platform based on engineered heart tissue. Circ. Res. 107, 35–44 (2010).
Zimmermann, W.-H. Tissue engineering of a differentiated cardiac muscle construct. Circ. Res. 90, 223–230 (2002).
Legant, W.R. et al. Microfabricated tissue gauges to measure and manipulate forces from 3D microtissues. Proc. Natl. Acad. Sci. USA 106, 10097–10102 (2009).
Serrao, G.W. et al. Myocyte-depleted engineered cardiac tissues support therapeutic potential of mesenchymal stem cells. Tissue Eng. Part A 18, 1322–1333 (2012).
Boudou, T. et al. A microfabricated platform to measure and manipulate the mechanics of engineered cardiac microtissues. Tissue Eng. Part A 18, 910–919 (2012).
Sakar, M.S. et al. Formation and optogenetic control of engineered 3D skeletal muscle bioactuators. Lab Chip 12, 4976–4985 (2012).
Hinson, J.T. et al. Titin mutations in iPS cells define sarcomere insufficiency as a cause of dilated cardiomyopathy. Science 349, 982–986 (2015).
Whitesides, G.M., Ostuni, E., Takayama, S., Jiang, X. & Ingber, D.E. Soft lithography in biology and biochemistry. Annu. Rev. Biomed. Eng. 3, 335–373 (2001).
Bower, A.F. Applied Mechanics of Solids (CRC Press, 2009).
Harris, A.K., Wild, P. & Stopak, D. Silicone rubber substrata: a new wrinkle in the study of cell locomotion. Science 208, 177–179 (1980).A seminal demonstration of the use of synthetic materials to observe cellular forces.
Harris, A.K., Stopak, D. & Wild, P. Fibroblast traction as a mechanism for collagen morphogenesis. Nature 290, 249–251 (1981).
Lee, J. Traction forces generated by locomoting keratocytes. J. Cell Biol. 127, 1957–1964 (1994).
Dembo, M., Oliver, T., Ishihara, A. & Jacobson, K. Imaging the traction stresses exerted by locomoting cells with the elastic substratum method. Biophys. J. 70, 2008–2022 (1996).
Legant, W.R. et al. Multidimensional traction force microscopy reveals out-of-plane rotational moments about focal adhesions. Proc. Natl. Acad. Sci. USA 110, 881–886 (2013).
Hur, S.S., Zhao, Y., Li, Y.-S., Botvinick, E. & Chien, S. Live cells exert 3-dimensional traction forces on their substrata. Cell. Mol. Bioeng. 2, 425–436 (2009).
Maskarinec, S.A., Franck, C., Tirrell, D.A. & Ravichandran, G. Quantifying cellular traction forces in three dimensions. Proc. Natl. Acad. Sci. USA 106, 22108–22113 (2009).
Balaban, N.Q. et al. Force and focal adhesion assembly: a close relationship studied using elastic micropatterned substrates. Nat. Cell Biol. 3, 466–472 (2001).The authors implement a TFM method to investigate the relationship between cell traction forces and the molecular structure of cell adhesions.
Beningo, K.A., Dembo, M., Kaverina, I., Small, J.V. & Wang, Y.L. Nascent focal adhesions are responsible for the generation of strong propulsive forces in migrating fibroblasts. J. Cell Biol. 153, 881–888 (2001).
Dembo, M. & Wang, Y.-L. Stresses at the cell-to-substrate interface during locomotion of fibroblasts. Biophys. J. 76, 2307–2316 (1999).An important study that used TFM to characterize the forces generated by migrating fibroblasts.
Jannat, R.A., Dembo, M. & Hammer, D.A. Traction forces of neutrophils migrating on compliant substrates. Biophys. J. 101, 575–584 (2011).
Plotnikov, S.V., Pasapera, A.M., Sabass, B. & Waterman, C.M. Force fluctuations within focal adhesions mediate ECM-rigidity sensing to guide directed cell migration. Cell 151, 1513–1527 (2012).The authors use high-resolution TFM to characterize the distribution of forces within single focal adhesions.
Engler, A.J., Sen, S., Sweeney, H.L. & Discher, D.E. Matrix elasticity directs stem cell lineage specification. Cell 126, 677–689 (2006).
Kraning-Rush, C.M., Califano, J.P. & Reinhart-King, C.A. Cellular traction stresses increase with increasing metastatic potential. PLoS ONE 7, e32572 (2012).
Oliver, T., Jacobson, K. & Dembo, M. Design and use of substrata to measure traction forces exerted by cultured cells. Methods Enzymol. 298, 497–521 (1998).
Beningo, K.A. & Wang, Y.-L. Flexible substrata for the detection of cellular traction forces. Trends Cell Biol. 12, 79–84 (2002).
Khetan, S. et al. Degradation-mediated cellular traction directs stem cell fate in covalently crosslinked three-dimensional hydrogels. Nat. Mater. 12, 458–465 (2013).
Kim, I.L., Khetan, S., Baker, B.M., Chen, C.S. & Burdick, J.A. Fibrous hyaluronic acid hydrogels that direct MSC chondrogenesis through mechanical and adhesive cues. Biomaterials 34, 5571–5580 (2013).
Hall, M.S. et al. Toward single cell traction microscopy within 3D collagen matrices. Exp. Cell Res. 319, 2396–2408 (2013).
Style, R.W. et al. Traction force microscopy in physics and biology. Soft Matter 10, 4047 (2014).
Wang, J.H.-C. & Lin, J.-S. Cell traction force and measurement methods. Biomech. Model. Mechanobiol. 6, 361–371 (2007).
Tseng, Q. et al. Spatial organization of the extracellular matrix regulates cell-cell junction positioning. Proc. Natl. Acad. Sci. USA 109, 1506–1511 (2012).
Schwarz, U.S. et al. Calculation of forces at focal adhesions from elastic substrate data: the effect of localized force and the need for regularization. Biophys. J. 83, 1380–1394 (2002).
Hur, S.S. et al. Roles of cell confluency and fluid shear in 3-dimensional intracellular forces in endothelial cells. Proc. Natl. Acad. Sci. USA 109, 11110–11115 (2012).
Sabass, B., Gardel, M.L., Waterman, C.M. & Schwarz, U.S. High resolution traction force microscopy based on experimental and computational advances. Biophys. J. 94, 207–220 (2008).
Franck, C., Hong, S., Maskarinec, S.A., Tirrell, D.A. & Ravichandran, G. Three-dimensional full-field measurements of large deformations in soft materials using confocal microscopy and digital volume correlation. Exp. Mech. 47, 427–438 (2007).
Franck, C., Maskarinec, S.A., Tirrell, D.A. & Ravichandran, G. Three-dimensional traction force microscopy: a new tool for quantifying cell-matrix interactions. PLoS ONE 6, e17833 (2011).
del Álamo, J.C. et al. Three-dimensional quantification of cellular traction forces and mechanosensing of thin substrata by Fourier traction force microscopy. PLoS ONE 8, e69850 (2013).
Baker, B.M. & Chen, C.S. Deconstructing the third dimension—how 3D culture microenvironments alter cellular cues. J. Cell Sci. 125, 3015–3024 (2012).
Koch, T.M., Münster, S., Bonakdar, N., Butler, J. & Fabry, B. III Traction forces in cancer cell invasion. PLoS ONE 7, e33476 (2012).
Bloom, R.J., George, J.P., Celedon, A., Sun, S.X. & Wirtz, D. Mapping local matrix remodeling induced by a migrating tumor cell using three-dimensional multiple-particle tracking. Biophys. J. 95, 4077–4088 (2008).
Steinwachs, J. et al. Three-dimensional force microscopy of cells in biopolymer networks. Nat. Methods 13, 171–176 (2016).
Legant, W.R. et al. Measurement of mechanical tractions exerted by cells in three-dimensional matrices. Nat. Methods 7, 969–971 (2010).This study characterized the traction stresses generated by cells embedded in 3D PEG hydrogels.
Galbraith, C.G. & Sheetz, M.P. A micromachined device provides a new bend on fibroblast traction forces. Proc. Natl. Acad. Sci. USA 94, 9114–9118 (1997).An example of a packaged microfabricated platform designed to measure cellular forces in real time.
Rajagopalan, J., Tofangchi, A. & Saif, M.T.A. Linear high-resolution BioMEMS force sensors with large measurement range. J. Microelectromech. Syst. 19, 1380–1389 (2010).
Rajagopalan, J. & Saif, M.T.A. MEMS sensors and microsystems for cell mechanobiology. J. Micromech. Microeng. 21, 54002 (2011).
Park, J. et al. Real-time measurement of the contractile forces of self-organized cardiomyocytes on hybrid biopolymer microcantilevers. Anal. Chem. 77, 6571–6580 (2005).
Feinberg, A.W. et al. Muscular thin films for building actuators and powering devices. Science 317, 1366–1370 (2007).
Grosberg, A., Alford, P.W., McCain, M.L. & Parker, K.K. Ensembles of engineered cardiac tissues for physiological and pharmacological study: heart on a chip. Lab Chip 11, 4165 (2011).
Bhatia, S.N. & Ingber, D.E. Microfluidic organs-on-chips. Nat. Biotechnol. 32, 760–772 (2014).
Huh, D., Hamilton, G.A. & Ingber, D.E. From 3D cell culture to organs-on-chips. Trends Cell Biol. 21, 745–754 (2011).
Tan, J.L. et al. Cells lying on a bed of microneedles: an approach to isolate mechanical force. Proc. Natl. Acad. Sci. USA 100, 1484–1489 (2003).An early report of micropillar arrays used to measure the forces generated by cells as a function of substrate stiffness and cell shape.
Fu, J. et al. Mechanical regulation of cell function with geometrically modulated elastomeric substrates. Nat. Methods 7, 733–736 (2010).
Trichet, L. et al. Evidence of a large-scale mechanosensing mechanism for cellular adaptation to substrate stiffness. Proc. Natl. Acad. Sci. USA 109, 6933–6938 (2012).
du Roure, O. et al. Force mapping in epithelial cell migration. Proc. Natl. Acad. Sci. USA 102, 2390–2395 (2005).
Ganz, A. et al. Traction forces exerted through N-cadherin contacts. Biol. Cell 98, 721–730 (2006).
Liu, Z. et al. Mechanical tugging force regulates the size of cell-cell junctions. Proc. Natl. Acad. Sci. USA 107, 9944–9949 (2010).
Maruthamuthu, V., Sabass, B., Schwarz, U.S. & Gardel, M.L. Cell-ECM traction force modulates endogenous tension at cell-cell contacts. Proc. Natl. Acad. Sci. USA 108, 4708–4713 (2011).
Saez, A., Ghibaudo, M., Buguin, A., Silberzan, P. & Ladoux, B. Rigidity-driven growth and migration of epithelial cells on microstructured anisotropic substrates. Proc. Natl. Acad. Sci. USA 104, 8281–8286 (2007).
Ghassemi, S. et al. Cells test substrate rigidity by local contractions on submicrometer pillars. Proc. Natl. Acad. Sci. USA 109, 5328–5333 (2012).
Yang, M.T., Fu, J., Wang, Y.-K., Desai, R.A. & Chen, C.S. Assaying stem cell mechanobiology on microfabricated elastomeric substrates with geometrically modulated rigidity. Nat. Protoc. 6, 187–213 (2011).
Stabley, D.R., Jurchenko, C., Marshall, S.S. & Salaita, K.S. Visualizing mechanical tension across membrane receptors with a fluorescent sensor. Nat. Methods 9, 64–67 (2011).
Liu, Y., Yehl, K., Narui, Y. & Salaita, K. Tension sensing nanoparticles for mechano-imaging at the living/nonliving interface. J. Am. Chem. Soc. 135, 5320–5323 (2013).
Morimatsu, M., Mekhdjian, A.H., Adhikari, A.S. & Dunn, A.R. Molecular tension sensors report forces generated by single integrin molecules in living cells. Nano Lett. 13, 3985–3989 (2013).
Liu, Y. et al. Nanoparticle tension probes patterned at the nanoscale: impact of integrin clustering on force transmission. Nano Lett. 14, 5539–5546 (2014).
Wang, X. & Ha, T. Defining single molecular forces required to activate integrin and Notch signaling. Science 340, 991–994 (2013).The authors implemented DNA-based sensors to measure the forces applied to single integrin molecules during early adhesion.
Blakely, B.L. et al. A DNA-based molecular probe for optically reporting cellular traction forces. Nat. Methods 11, 1229–1232 (2014).
Zhang, Y., Ge, C., Zhu, C. & Salaita, K. DNA-based digital tension probes reveal integrin forces during early cell adhesion. Nat. Commun. 5, 5167 (2014).
Grashoff, C. et al. Measuring mechanical tension across vinculin reveals regulation of focal adhesion dynamics. Nature 466, 263–266 (2010).A seminal study in which an engineered cell-adhesion protein is used to measure the forces in focal adhesions.
Hoffman, B.D., Grashoff, C. & Schwartz, M.A. Dynamic molecular processes mediate cellular mechanotransduction. Nature 475, 316–323 (2011).
Conway, D.E. et al. Fluid shear stress on endothelial cells modulates mechanical tension across VE-cadherin and PECAM-1. Curr. Biol. 23, 1024–1030 (2013).
Borghi, N. et al. E-cadherin is under constitutive actomyosin-generated tension that is increased at cell-cell contacts upon externally applied stretch. Proc. Natl. Acad. Sci. USA 109, 12568–12573 (2012).
Meng, F. & Sachs, F. Visualizing dynamic cytoplasmic forces with a compliance-matched FRET sensor. J. Cell Sci. 124, 261–269 (2011).
Smith, M.L. et al. Force-induced unfolding of fibronectin in the extracellular matrix of living cells. PLoS Biol. 5, e268 (2007).
Cost, A.-L., Ringer, P., Chrostek-Grashoff, A. & Grashoff, C. How to measure molecular forces in cells: a guide to evaluating genetically-encoded FRET-based tension sensors. Cell. Mol. Bioeng. 8, 96–105 (2015).
Mammoto, T. & Ingber, D.E. Mechanical control of tissue and organ development. Development 137, 1407–1420 (2010).
Campàs, O. et al. Quantifying cell-generated mechanical forces within living embryonic tissues. Nat. Methods 11, 183–189 (2014).
Paszek, M.J. et al. Tensional homeostasis and the malignant phenotype. Cancer Cell 8, 241–254 (2005).
McBeath, R., Pirone, D.M., Nelson, C.M., Bhadriraju, K. & Chen, C.S. Cell shape, cytoskeletal tension, and RhoA regulate stem cell lineage commitment. Dev. Cell 6, 483–495 (2004).
Rape, A.D., Guo, W.H. & Wang, Y.L. The regulation of traction force in relation to cell shape and focal adhesions. Biomaterials 32, 2043–2051 (2011).
Thavandiran, N. et al. Design and formulation of functional pluripotent stem cell-derived cardiac microtissues. Proc. Natl. Acad. Sci. USA 110, E4698–E4707 (2013).
Yang, M.T., Reich, D.H. & Chen, C.S. Measurement and analysis of traction force dynamics in response to vasoactive agonists. Integr. Biol. (Camb.) 3, 663 (2011).
Butler, J.P., Tolic-Norrelykke, I.M., Fabry, B. & Fredberg, J.J. Traction fields, moments, and strain energy that cells exert on their surroundings. Am. J. Physiol. Cell Physiol. 282, C595–C605 (2002).
Wang, N. et al. Cell prestress. I. Stiffness and prestress are closely associated in adherent contractile cells. Am. J. Physiol. Cell Physiol. 282, C606–C616 (2002).
Pelham, R.J. & Wang, Y.L. Cell locomotion and focal adhesions are regulated by substrate flexibility. Proc. Natl. Acad. Sci. USA 94, 13661–13665 (1997).
Meyers, M.A., Chen, P.-Y., Lin, A.Y.-M. & Seki, Y. Biological materials: structure and mechanical properties. Prog. Mater. Sci. 53, 1–206 (2008).
Acknowledgements
We thank A. Chopra and M. Kutys for helpful discussions. This work was supported in part by the US National Institutes of Health (NIH) (grants EB00262 and GM74048 to C.S.C.), the National Science Foundation (grant CMMI-1462710 to C.S.C.), and the RESBIO Technology Resource for Polymeric Biomaterials (grant P41-EB001046 to C.S.C.). W.J.P. acknowledges financial support from the NIH through the Organ Design and Engineering Training program (T32 EB16652).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Polacheck, W., Chen, C. Measuring cell-generated forces: a guide to the available tools. Nat Methods 13, 415–423 (2016). https://doi.org/10.1038/nmeth.3834
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3834
This article is cited by
-
Hydrogel-based molecular tension fluorescence microscopy for investigating receptor-mediated rigidity sensing
Nature Methods (2023)
-
Organization, dynamics and mechanoregulation of integrin-mediated cell–ECM adhesions
Nature Reviews Molecular Cell Biology (2023)
-
The extracellular matrix mechanics in the vasculature
Nature Cardiovascular Research (2023)
-
Tension-tuned receptors for synthetic mechanotransduction and intercellular force detection
Nature Biotechnology (2023)
-
Contact Guidance Drives Upward Cellular Migration at the Mesoscopic Scale
Cellular and Molecular Bioengineering (2023)