Abstract
We report that the efficiency of reprogramming human somatic cells to induced pluripotent stem cells (hiPSCs) can be dramatically improved in a microfluidic environment. Microliter-volume confinement resulted in a 50-fold increase in efficiency over traditional reprogramming by delivery of synthetic mRNAs encoding transcription factors. In these small volumes, extracellular components of the TGF-β and other signaling pathways exhibited temporal regulation that appears critical to acquisition of pluripotency. The high quality and purity of the resulting hiPSCs (μ-hiPSCs) allowed direct differentiation into functional hepatocyte- and cardiomyocyte-like cells in the same platform without additional expansion.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Boyer, L.A. et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122, 947–956 (2005).
Vallier, L., Alexander, M. & Pedersen, R.A. Activin/Nodal and FGF pathways cooperate to maintain pluripotency of human embryonic stem cells. J. Cell Sci. 118, 4495–4509 (2005).
Vallier, L. et al. Activin/Nodal signaling maintains pluripotency by controlling Nanog expression. Development 136, 1339–1349 (2009).
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007).
Golipour, A. et al. A late transition in somatic cell reprogramming requires regulators distinct from the pluripotency network. Cell Stem Cell 11, 769–782 (2012).
Schlaeger, T.M. et al. A comparison of non-integrating reprogramming methods. Nat. Biotechnol. 33, 58–63 (2015).
Giobbe, G.G. et al. Functional differentiation of human pluripotent stem cells on a chip. Nat. Methods 12, 637–640 (2015).
Giulitti, S., Magrofuoco, E., Prevedello, L. & Elvassore, N. Optimal periodic perfusion strategy for robust long-term microfluidic cell culture. Lab Chip 13, 4430–4441 (2013).
Takahashi, K. et al. Induction of pluripotency in human somatic cells via a transient state resembling primitive streak-like mesendoderm. Nat. Commun. 5, 3678 (2014).
Luni, C., Michielin, F., Barzon, L., Calabrò, V. & Elvassore, N. Stochastic model-assisted development of efficient low-dose viral transduction in microfluidics. Biophys. J. 104, 934–942 (2013).
Warren, L. et al. Highly efficient reprogramming to pluripotency and directed differentiation of human cells with synthetic modified mRNA. Cell Stem Cell 7, 618–630 (2010).
Li, R. et al. A mesenchymal-to-epithelial transition initiates and is required for the nuclear reprogramming of mouse fibroblasts. Cell Stem Cell 7, 51–63 (2010).
Liu, X. et al. Sequential introduction of reprogramming factors reveals a time-sensitive requirement for individual factors and a sequential EMT–MET mechanism for optimal reprogramming. Nat. Cell Biol. 15, 829–838 (2013).
Yu, J. et al. Induced pluripotent stem cell lines derived from human somatic cells. Science 318, 1917–1920 (2007).
Zhou, T. et al. Generation of human induced pluripotent stem cells from urine samples. Nat. Protoc. 7, 2080–2089 (2012).
Xue, Y. et al. Generating a non-integrating human induced pluripotent stem cell bank from urine-derived cells. PLoS ONE 8, e70573 (2013).
Sharei, A. et al. A vector-free microfluidic platform for intracellular delivery. Proc. Natl. Acad. Sci. USA 110, 2082–2087 (2013).
Skelley, A.M., Kirak, O., Suh, H., Jaenisch, R. & Voldman, J. Microfluidic control of cell pairing and fusion. Nat. Methods 6, 147–152 (2009).
Anokye-Danso, F., Snitow, M. & Morrisey, E.E. How microRNAs facilitate reprogramming to pluripotency. J. Cell Sci. 125, 4179–4787 (2012).
Parchem, R.J. et al. Two miRNA clusters reveal alternative paths in late-stage reprogramming. Cell Stem Cell 14, 617–631 (2014).
Rais, Y. et al. Deterministic direct reprogramming of somatic cells to pluripotency. Nature 502, 65–70 (2013).
Gafni, O. et al. Derivation of novel human ground state naive pluripotent stem cells. Nature 504, 282–286 (2013).
Vidal, S.E., Amlani, B., Chen, T., Tsirigos, A. & Stadtfeld, M. Combinatorial modulation of signaling pathways reveals cell-type-specific requirements for highly efficient and synchronous iPSC reprogramming. Stem Cell Rep. 3, 574–584 (2014).
Bar-Nur, O. et al. Small molecules facilitate rapid and synchronous iPSC generation. Nat. Methods 11, 1170–1176 (2014).
Dimmeler, S., Ding, S., Rando, T.A. & Trounson, A. Translational strategies and challenges in regenerative medicine. Nat. Med. 20, 814–821 (2014).
Neuži, P., Giselbrecht, S., Länge, K., Huang, T.J. & Manz, A. Revisiting lab-on-a-chip technology for drug discovery. Nat. Rev. Drug Discov. 11, 620–632 (2012).
Luni, C., Serena, E. & Elvassore, N. Human-on-chip for therapy development and fundamental science. Curr. Opin. Biotechnol. 25, 45–50 (2014).
Zhou, T. et al. Generation of human induced pluripotent stem cells from urine samples. Nat. Protoc. 7, 2080–2089 (2012).
Unger, M.A., Chou, H.-P., Thorsen, T., Scherer, A. & Quake, S.R. Monolithic microfabricated valves and pumps by multilayer soft lithography. Science 288, 113–116 (2000).
Gómez-Sjöberg, R., Leyrat, A.A., Pirone, D.M., Chen, C.S. & Quake, S.R. Versatile, fully automated, microfluidic cell culture system. Anal. Chem. 79, 8557–8563 (2007).
Warren, L., Ni, Y., Wang, J. & Guo, X. Feeder-free derivation of human induced pluripotent stem cells with messenger RNA. Sci. Rep. 2, 657 (2012).
Singh, A. et al. Adhesion strength-based, label-free isolation of human pluripotent stem cells. Nat. Methods 10, 438–444 (2013).
Erban, R. & Chapman, S.J. Reactive boundary conditions for stochastic simulations of reaction–diffusion processes. Phys. Biol. 4, 16–28 (2007).
Moledina, F. et al. Predictive microfluidic control of regulatory ligand trajectories in individual pluripotent cells. Proc. Natl. Acad. Sci. USA 109, 3264–3269 (2012).
Kestin, J., Sokolov, M. & Wakeham, W.A. Viscosity of liquid water in the range −8 °C to 150 °C. J. Phys. Chem. Ref. Data 7, 941 (1978).
Russo, L., Berardi, V., Tardani, F., La Mesa, C. & Risuleo, G. Delivery of RNA and its intracellular translation into protein mediated by SDS-CTAB vesicles: potential use in nanobiotechnology. BioMed Res. Int. 2013, 734596 (2013).
Inui, M. et al. USP15 is a deubiquitylating enzyme for receptor-activated SMADs. Nat. Cell Biol. 13, 1368–1375 (2011).
Johnson, W.E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Acknowledgements
This research was supported by Progetti di Eccellenza Ca.Ri.Pa.Ro. and Progetto Strategico TRANSAC of University of Padova. E.S. was supported by Progetti Giovani Studiosi 2010 (DIRPRGR10) of University of Padova. We would like to thank M. Piccoli (Fondazione Istituto di Ricerca Pediatrica Città della Speranza) for teratoma assay, A. Zoso (University of Padova, Italy) for western blot optimization, D. Bellotti and J. Skripac for karyotype analyses at the Cytogenetic and Molecular Genetics Laboratory at the University of Brescia (Italy), M. Mora, S. Zanotti and Telethon Network of Genetic Biobanks for providing SkMF, and I. Ferrarotti for ATD samples (IRCCS, Pavia, Italy). Furthermore, we would like to thank S. Rüberg, S. Tomiuk and M. Knauel for microarray analysis, and S. Wild and M. Jurk, all at Miltenyi Biotec, for providing material and support for reprogramming technologies. We thank S. Piccolo and G. Martello (University of Padova, Italy) for critical revision of the manuscript.
Author information
Authors and Affiliations
Contributions
S.G., C.L. and N.E. designed and performed the experiments; C.L. performed computational analyses. S.G., E.S. and L.F. characterized hiPSCs. S.G. and F.M. developed microfluidic hPSC culture. G.G.G. designed hepatic differentiation. A.Z. and O.G. designed and realized the automated microfluidic platform. S.K. and A.B. contributed to microarray analysis and reprogramming design. C.L., S.G. and N.E. designed the study and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
C.L., S.G., and N.E. are coinventors on a patent application describing the reprogramming and differentiation processes in microfluidics, application number PD2013A000220. S.K. and A.B. are employees of Miltenyi Biotec.
Integrated supplementary information
Supplementary Figure 1 Extrinsic Signaling during Reprogramming.
a, In human pluripotent stem cells, exogenously- (medium) and endogenously-promoted (cell-secreted) FGF and TGF-β pathways sustain the master genetic network of NANOG, POU5F1 and SOX2. b, The extra-cellular soluble components of the signaling pathways were identified at the intersection of genes related to the extra-cellular region and genes involved in signaling pathways. The temporal expression of these components during reprogramming was analyzed from data reported by Takahashi et al.18. Multiple of these genes that are progressively up- (red) or down-regulated (green) during reprogramming take part in signaling pathways implicated in pluripotency maintenance and self-renewal. c, In conventional well culture systems, endogenous factors get diluted in the medium bulk by diffusion; on the contrary, reducing medium volume, the concentration of endogenous factors is maintained high in close proximity to the cells.
Supplementary Figure 2 Extrinsic signaling pathway components show dynamic behavior during reprogramming.
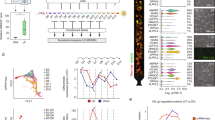
a, Hierarchical clustering based on microarray expression data (GSE50206) of genes belonging to the intersection of GO:Extracellular region with TGF-β, WNT, MAPK, HIF and AKT-PI3K pathways. Temporal evolution of gene expression during reprogramming shows convergence to pluripotent profiles. Color bar: relative median-centered gene expression presented as a green-black-red-colored heatmap (log2). b, K-means clustering of the same genes as in (a) into three main transcriptional patterns (invariant, down-regulation and up-regulation) during reprogramming (49 days). Cells at the end of reprogramming show transcriptional profiles consistent with stable hiPSC and hESC.
Supplementary Figure 3 Technological development of microfluidic cell culturing for reprogramming.
a, Schematic of the two classes of substrates tested: adsorbed extra-cellular matrix (ECM) only by physical interaction, and cross-linked ECM by covalent chemical bonding. b, Eight different surface treatments were tested within microfluidic channels: A: adsorbed 0.6% Gel on glass, B: adsorbed 0.1% Gel, C: glass silanization by APTES, D: methacrylated glass via TMSPMA, E: 2% GelMA, covalently bound to methacrylated glass, F: adsorbed Fn, G: 0.1% Gel, covalently bound to APTES treated-glass via glutaraldehyde, H: 0.1% GelMA, covalently bound to methacrylated glass. Error bars, mean ± s.d. (n 3). c, Representative images of BJ cultures on differently functionalized surfaces. Time points (day): A, D4; B, D9; C, D12; D and E, D14; F, D16; G and H, D24. Nomenclature A-H: same as in (b). Scale bars, 500 μm. d, Reprogramming microfluidic chip design with complete manual management is performed by pipetting solutions into a reservoir. By picking up exausted medium from the outlet, fresh medium introduced in culture channel. Each chip is built on common microscope glass slides and can be placed in a standard Petri dish with a phosphate buffer bath to ensure a sterile humidified envi- ronment. No other equipment is needed. e, Exploded and assembled representation of the microfluidic platform with an integrated automated medium distribution system; colors highlight culture channels and medium by-passes (pink; thick and thin, respectively), medium distribution network (black), and pneumatic control circuit (gray). Below, the section of a single channel is shown schematically to display automated operation mechanism (layers not on scale): the medium distribution system works by pneumatic control with pulsed periodic medium changes within the culture channel during the day.
Supplementary Figure 4 Efficiency of a single and multiple daily transfections of nGFP mmRNA on BJ cells.
a, Transfections were performed either for 4 h or overnight; fluorescence was detected 24 h after beginning of transfection, different transfection reagents were used: Stemfect (SF), StemMACS (SM), and RNAiMAX (RiM); cells transfected in microfluidics show higher efficiency in terms of percentage of nGFP+ cells and much lower mmRNA consumption; error bars, mean±s.d. (n=4). b-c, mmRNAs were delivered every 24 h for three times by SF or SM on BJ cells with a seeding density of 120 cells/mm2, either for 4 h or overnight. The two different mmRNA amounts indicated are expressed per 100 cells. Scale bar, 250 μm. After 72 h nGFP+ cells were nearly 90% using SF. Employing SM reagent, a smaller amount of transfected cells was visible, although markedly expressing nGFP. Error bars, mean ± s.d. (n=3).
Supplementary Figure 5 Reprogramming factors delivery efficiency.
a, Schematic representation of the physical processes accounted for in the stochastic mathematic model: cells are represented as discrete areas on the culture surface, and RNA-containing vesicles move in the medium above by Brownian motion within a lattice. Pjump, probability of a movement between neighboring positions on the lattice or towards cell surface; Pads, probability of vesicle adsorption into a cell. Dimensions are not on scale. b, Experimental (large dots) and mathematical model-derived (small dots) transfection efficiency for different incubation periods (RiM). Error bars, mean ± s.d. (n>4). c, In silico simulation of mmRNA content distribution in the cell population, as a function of mmRNA amount and transfection duration. d, Protein expression in early reprogramming phase. BJ cells were daily transfected with 5.5 pg/100 cells of 7-mmRNA mix, including nGFP mmRNA, during reprogramming. Images were taken 24 h and 48 h after the first transfection. Scale bar, 100 μm. e, Quantification of nuclei fluorescence intensity determined by analysis of images obtained as in (d). To account for substrate differences (polystyrene vs. glass), fluorescence was normalized by background intensity. Cells transfected in microfluidics (μF) exhibit a significantly higher fluorescence intensity of nGFP compared to those in well. Error bars, mean ± s.d. (n>600 cells). f, Cell lysates from the same samples, both wells and single microfluidic channels, were analyzed by Western blot for POUF5F1 (OCT4) and KLF4. Cells transfected in microfluidics express a higher protein level also of these reprogramming factors. g, Fibroblast morphological mesenchymal-epithelial transition and transfection progression in microfluidics during reprogramming verified by nGFP expression. Scale bars, 500 and 150 μm (macro) refer to each column of figures.
Supplementary Figure 6 Extraction methods of patient-specific freshly generated hiPSC from microfluidics.
Manual picking: a single colony (arrow) was mechanically dissected and picked with a glass tool after coring the PDMS surface with a biopsy punch (both the hole and the core flipped upside-down are shown). Colony fragments were seeded on MEF in a well and conventionally expanded. Shear stress-based extraction: hiPSC (left, arrows) were detached and collected at the outlet of the microfluidic channel by applying DPBS perfusion at high rate. Gathered cells were then seeded on MEF. Opening of the microfluidic channel and subsequent manual picking of single colonies results in a straightforward method to maintain hiPSC clonality upon isolation and cell line generation. While shear stress can dissociate each original hiPSC colony in multiple smaller clusters, seeding into a well in sparse condition permits to randomly choose and isolate hiPSC arbitrary clones few days later. Alternatively, it is possible to perfuse colonies twice for 5 min each with 0.5 mM ethylenediaminetetraacetic acid, a standard solution for expansion of pluripotent stem cells in feeder-free conditions; Loosen hiPSC are then detached and collected perfusing with hiPSC medium. Scale bars, 750 μm (channels), 250 μm (on MEF).
Supplementary Figure 7 In situ and off-chip characterization of hiPSC derived from BJ and HFF fibroblasts.
a, Immunofluorescence of early reprogramming phase (day 6) in a microfluidics. E-CAD antibody was used to detect BJ cells undergoing mesenchymal-epithelial transition. After 5 daily transfections most of the cells express epithelial cadherin. Scale bars, 250 μm. b-e, Characterization of BJ-derived hiPSC extracted from microfluidic channels. b, pluripotency markers by immunofluorescence and alkaline phosphatase expression. Scale bars, 100 μm. c, pluripotency genes expression by RT-PCR. d, upon differentiation stimuli, hiPSCs are able to generate embryoid bodies randomly expressing markers of three germ layers. e, normal karyotype. f, Characterization of HFF-derived hiPSC. In situ immunofluorescence demonstrates obtained colonies are NANOG+, OCT-4+ and SSEA-4+; analyses were performed 3 days after the last reprogramming transfection. Karyotype analysis does not show abnormalities in HFF-derived hiPSC after 12 passages (e, μF-hiPSC #357). Scale bars, 200 μm. g, Upon subcutaneous injection in Rag2-/-γc-/- mice, hiPSC derived in microfluidics underwent spontaneous differentiation generating teratomas with structures typical of the three germ layers.
Supplementary Figure 8 Feeder-dependent reprogramming in microfluidics.
a, Highly efficient reprogramming within the microfluidic setup. A binary image of NANOG+ (green) and TRA-1-60+ (red) colonies, detected by fluorescence microscopy 48 h after the last reprogramming mmRNA transfection, was overlapped with the bright-field image of the corresponding 10 microfluidic channels. The same channels of Fig. 3b are shown. b, Image of 10 channels within a microfluidic chip at day 18 of reprogramming with feeders. Primary derived hiPSC are visible as white translucent spots inside each channel. Scale bar, 3 mm. A magnification of channel 4 shows 4 colonies. Channel width is 1.5 mm. c, Reprogramming efficiency as a function of mmRNA dosage, maintained constant during the process. 1× refers to the optimal transfection dosages in well and microfluidic systems, 24 and 4 pg/100 cells, respectively.
Supplementary Figure 9 Feeder-free reprogramming of BJ cells performed with SF and SM.
BJs were seeded at 10 cell/mm2 and transfected for 18 days (SF) or 11 days (SM). a, Time course of nGFP expression during the first 6 days of reprogramming. Scale bar, 250 μm. b, TRA-1-60+ colonies are schematically marked inside each channel at their actual position. Compared to SF, feeder-free reprogramming with SM generated a significantly higher number of colonies with a more homogeneous distribution along each microfluidic channel.
Supplementary Figure 10 Feeder-free reprogramming of BJ cells with optimized conditions.
a, Nested ANOVA analysis considering all reprogrammed samples (biological and technical independent replicates) reprogrammed in optimized feeder-free conditions with incremental mmRNA dosage during the first days. Variability between different cell batches or microfluidic chips is not statistically significant and results in similar p-values. b, Reprogramming in feeder-free condition in microfluidics generates a considerably higher amount of hiPSC colonies per unit area compared to a standard well plate, both when performed at optimal initial cell density (5 cell/mm2), and with a 10-fold higher initial cell density (50 cell/mm2). In the latter case, daily mmRNA amount has been doubled during the first week to accommodate exponential growth of a higher amount of cells. Despite this, the amount of hiPSC colonies generated per unit area is drastically reduced. Error bars, mean ± s.d. n=6. c, Morphological cell changes during reprogramming progression performed in microfluidics as described in (b). Conventional seeding density in microfluidic optimized reprogramming is 4.88 ± 1.60 cell/mm2, n=32, consistent with the expected value of 5 cell/mm2.
Supplementary Figure 11 hiPSC derived from renal epithelial (RE) cells isolated from urine.
a, RE cells at day 13 of the reprogramming protocol, after daily mmRNA transfections, within a microfluidic channel (phase contrast and nGFP fluorescence). b, Extracted hiPSC at passage 1 (p1) directly cultured in feeder-free conditions and expanded at passage 3 (p3). c, Immunofluorescence characterization of RE-derived hiPSC confirms the expression of endogenous NANOG, OCT-4 (POU5F1), SSEA-4, and TRA-1-60. Scale bars 200 μm, except (b, left) 25 μm.
Supplementary Figure 12 Comparison of hiPSC colony expression profiles.
Microarray datasets from hiPSC colonies obtained in this work either in microfluidics or in well, are compared to datasets in GEO and Synapse databases (GSE50206, GSE42445, syn1447097, syn1449098). Principal component analysis shows mmRNA colonies obtained in this work (p0, p3) have expression profiles similar to pluripotent stem cell samples from literature. Moreover, differences among hiPSC colonies obtained in this work in well plates or in microfluidics (μF) are within the biological variability of pluripotent stem cells obtained by other groups.
Supplementary Figure 13 Downscale of mmRNA transfection in systems characterized by different medium height.
mmRNA concentration used in well is maintained constant and mmRNA quantity rescale accordingly to medium height. The 200-μm-tall microfluidic channel represent the standard used in this paper and promotes high concentration of endogenous factors. By decreasing the medium height, no substantial reprogramming efficiency reduction is observed, while transfection (day 8) efficiency decreases with the reduction of mmRNA quantity introduced in the system. Reprogramming performed in 1,000-μm-tall channels with the same mmRNA quantity (0.5 ng mm-2) used in 200 μm height microfluidic optimised condition (Fig. 3a; efficiency of 121 ± 38) has a ~100-fold reduction in the reprogramming efficiency index.
Supplementary Figure 14 Characterization of cells undergoing reprogramming in microfluidics.
a, Three conditions were tested: non-treated control (-), inhibition of TGF-β1 signaling (SB, 2 μM) and exogenous TGF- b1. Images were taken at day 7 after the first transfection. Scale bar, 50 mm. Quantification of the percentage of E-CAD+ cells is shown below. Expression of the epithelial marker E-CAD is high in each conditions, although TGF- b1-treated cells do not give rise to any hiPSC colony (Figure 4h). b, Reprogramming at day 12. While exogenous TGF-β1 induces fibroblasts elongation counteracting mesenchymal to epithelial transition, treatment with SB (either 2 or 10 μM) promotes the definition of an epithelial-like cell layer without substantial cell aggregation and rise of small hiPSC clusters (-).
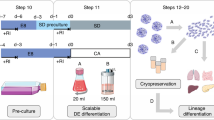
Supplementary Figure 15 Optimization of reprogramming and straightforward differentiation.
a, Comparison of requirements for reprogramming in microfluidics and well (left) and downscaling strategy of the conventional process to obtain functional cell types from hiPSC in single-seeding procedure (right). b, Freshly-derived hiPSC were differentiated for 5 days in EB medium allowing the derivation of different cell phenotypes derived from the same colony. RT-PCR analysis (right) reveals the presence of markers of the three germ-layers. c, Freshly-derived hiPSC were also differentiated towards a functional cardiac phenotype for 14 days. At the end of the differentiation protocol, extensive areas within the culture channel show tissue rearrangement (left). Scale bar, 750 μm. RT-PCR (right), from single-channel total RNA extraction, evidenced the presence of advanced cardiac development and functional maturation. RT, reverse transcriptase; c+, samples used as a positive control.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15 and Supplementary Tables 1 and 2 (PDF 2497 kb)
Supplementary Table 3
List of single-experiment conditions and reprogramming efficiency results. List of characterized clones and performed analyses. (XLSX 55 kb)
Supplementary Data 1
Bioinformatics analysis of microarray expression data (GSE50206). List of genes belonging to the intersection of GO: Extracellular region with TGF-beta, WNT, MAPK, HIF and AKT-PI3K pathways along human fibroblast reprogramming. (XLSX 45 kb)
Supplementary Data 2
Bioinformatics analysis of microarray data (GSE59534). Expression profiles of differentially expressed genes between the four conditions: p0 μF, p0 well, p3 μF, and p3 well. (XLSX 31476 kb)
Supplementary Data 3
Bioinformatics analysis of microarray data (GSE59534). Functional annotation analysis of differentially expressed genes between the four conditions: p0 μF, p0 well, p3 μF, and p3 well. (XLSX 31292 kb)
Supplementary Software
MATLAB code for the stochastic mathematical model of mmRNA transfections. LabView code for controlling the automated microfluidic platform. (ZIP 10971 kb)
Automated reprogramming chip
Validation of the automated microfluidic platform with integrated medium distribution system by food dyes. (MP4 872 kb)
Cells reprogrammed in microfluidics
Morphological appearance of freshly obtained hiPSC colonies within 10 microfluidic channels at day 18 from beginning of reprogramming. (MP4 7675 kb)
Reprogramming progression in microfluidics
Time lapse of a feeder-free reprogramming process of human fibroblasts in microfluidics. Frames were captured every 12 h for 17 days. Day 0 represents the day of the first mmRNA transfection. At the end of the process, cells were fixed and stained for NANOG immunofluorescence detection. (MP4 8761 kb)
Reprogramming progression in microfluidics
Enlargement of a selected area from Supplementary Video 4. (MP4 10831 kb)
Differentiation of freshly-derived hiPSC
Contracting culture of cardiomyocytes. Freshly obtained microfluidic hiPSC colonies were directly differentiated in well towards the cardiac lineage. (MP4 965 kb)
Rights and permissions
About this article
Cite this article
Luni, C., Giulitti, S., Serena, E. et al. High-efficiency cellular reprogramming with microfluidics. Nat Methods 13, 446–452 (2016). https://doi.org/10.1038/nmeth.3832
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3832
This article is cited by
-
Cellular population dynamics shape the route to human pluripotency
Nature Communications (2023)
-
Synchronization between peripheral circadian clock and feeding-fasting cycles in microfluidic device sustains oscillatory pattern of transcriptome
Nature Communications (2021)
-
Highly parallelized human embryonic stem cell differentiation to cardiac mesoderm in nanoliter chambers on a microfluidic chip
Biomedical Microdevices (2021)
-
Microfluidic reprogramming to pluripotency of human somatic cells
Nature Protocols (2019)
-
In vitro metabolic zonation through oxygen gradient on a chip
Scientific Reports (2019)