Abstract
We describe epiGBS, a reduced representation bisulfite sequencing method for cost-effective exploration and comparative analysis of DNA methylation and genetic variation in hundreds of samples de novo. This method uses genotyping by sequencing of bisulfite-converted DNA followed by reliable de novo reference construction, mapping, variant calling, and distinction of single-nucleotide polymorphisms (SNPs) versus methylation variation (software is available at https://github.com/thomasvangurp/epiGBS). The output can be loaded directly into a genome browser for visualization and into RnBeads for analysis of differential methylation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
References
Suzuki, M.M. & Bird, A. Nat. Rev. Genet. 9, 465–476 (2008).
Beck, S. Nat. Biotechnol. 28, 1026–1028 (2010).
Gu, H. et al. Nat. Protoc. 6, 468–481 (2011).
Wang, J. et al. BMC Genomics 14, 11 (2013).
Boyle, P. et al. Genome Biol. 13, R92 (2012).
Elshire, R.J. et al. PLoS ONE 6, e19379 (2011).
Rigby, P.W.J., Dieckmann, M., Rhodes, C. & Berg, P. J. Mol. Biol. 113, 237–251 (1977).
Becker, C. et al. Nature 480, 245–249 (2011).
Pedersen, B.S., Eyring, K., De, S., Yang, I.V. & Schwartz, D.A. Preprint at http://arxiv.org/abs/1401.1129 (2014).
Garrison, E. & Marth, G. Preprint at http://arxiv.org/abs/1207.3907 (2012).
Liu, Y., Siegmund, K.D., Laird, P.W. & Berman, B.P. Genome Biol. 13, R61 (2012).
Assenov, Y. et al. Nat. Methods 11, 1138–1140 (2014).
Lister, R. et al. Cell 133, 523–536 (2008).
Cokus, S.J. et al. Nature 452, 215–219 (2008).
Harris, R.A. et al. Nat. Biotechnol. 28, 1097–1105 (2010).
Vandegehuchte, M.B. et al. Environ. Int. 35, 700–706 (2009).
Schrey, A.W. et al. Integr. Comp. Biol. 53, 340–350 (2013).
Vergeer, P., Wagemaker, N.C.A.M. & Ouborg, N.J. Biol. Lett. 8, 798–801 (2012).
Swillen, I., Vanoverbeke, J. & De Meester, L. Ecol. Evol. 5, 2712–2721 (2015).
Verhoeven, K.J.F., van Dijk, P.J. & Biere, A. Mol. Ecol. 19, 315–324 (2010).
Zhang, J., Kobert, K., Flouri, T. & Stamatakis, A. Bioinformatics 30, 614–620 (2014).
Li, H. et al. Bioinformatics 25, 2078–2079 (2009).
Edgar, R.C. Bioinformatics 26, 2460–2461 (2010).
Acknowledgements
We thank F. Johannes (Department of Plant Science, Technical University of Munich, Munich, Germany) and C. Becker (Max Planck Institute for Developmental Biology, Tübingen, Germany) for providing Arabidopsis samples, and we thank G. Maes, J. van Houdt and E. Schijlen for sequencing several epiGBS libraries as we developed the protocol. Daphnia magna samples were provided by L. De Meester (University of Leuven, Leuven, Belgium). Fallopia japonica and Fallopia × bohemica samples were provided by C. Richards (University of South Florida, Tampa, Florida, USA). Allium porrum and Daucus carota seeds were provided by Nunhems Netherlands BV–Bayer CropScience. Finally, we thank C. Bock, C. Richards, R. Elshire, W. van der Putten and A. Biere for critical comments and helpful suggestions on the manuscript. This study was funded by the Netherlands Organization for Scientific Research (NWO-ALW grants 864.10.008 and 820.01.025 to K.J.F.V.).
Author information
Authors and Affiliations
Contributions
T.P.v.G. conceived the method, performed lab work, developed software, analyzed data and wrote the paper. N.C.A.M.W. codeveloped the method, provided sample material, performed lab work and contributed to writing of the paper. B.W. codeveloped software and developed the RnBeads interface scripts. K.J.F.V. contributed to interpretation of the experiment and method and to writing of the paper. P.V. provided samples for the Scabiosa case study and contributed to interpretation of the experiment and method. J.N.J.O. contributed samples and to interpretation of the experiment and method.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Barcodes, barcoded adaptors and resulting expected nucleotide distributions and epiGBS clusters.
(a) The following forward and reverse inline barcodes were used to generate barcoded forward and reverse adapters (Supplementary Table 1b), (b) Barcoded adapter design is identical to the adapters used in Genotyping by Sequencing1 with the exception of the use of the B adapter barcode which is not present in GBS. Barcoded adapter sequences of both A and B adapters were generated using http://www.deenabio.com/services/gbs-adapters with 12 4-6 nucleotides barcodes for the A adapter and a subset of 8 4-nucleotide barcoded adapters were used for the B adapter. The sequence of Illumina PE-PCR primers 1 and 2 used to amplify the libraries are also listed. (c) Complimentary oligonucleotides with 5-methylcytosines instead of cytosines are annealed to form adapters. The four base 5’-3’ overhang complements the restriction site overhang generated by PstI but can be modified depending on the specific enzyme used. Adapter A is identical to the barcoded adapter used in GBS. 4-6 nucleotide barcodes are designed to maximize diversity over the first cycles of the reads. Adapter B is identical to the common adapter used in GBS, with the exception that a barcoded sequence is placed before the enzyme overhang. For Csp6I we used 5’-CCTA-3’ as forward and 5’-CTGG-3’ as reverse barcode. The overhang of both Csp6I adapters was modified with respect to PstI, instead of 5’-TGCA-3’ we designed adapters with 5’-AT-3’ complementary to the overhang generated by Csp6I digestion. (d) Given equal representation of all 12 forward barcodes the expected per cycle nucleotide composition of the forward read is depicted. Up to position 5 the composition is mostly unbiased, aiding the calculation of proper run-specific parameters during Illumina sequencing and thus preventing phasing and pre-phasing detection errors that can lead to low quality sequencing libraries. (e) Per species the total number of paired-end sequencing reads, individuals per species, percentage of merged reads, number of de novo discovered clusters, CG content, average size and number of clusters having gene hits (see online methods) is shown.
1Robert J Elshire et al., "A Robust, Simple Genotyping-by-Sequencing (GBS) Approach for High Diversity Species.," PLoS One 6, no. 5 (2011): e19379, doi:10.1371/journal.pone.0019379.
2Martin Kircher, Patricia Heyn, and Janet Kelso, "Addressing Challenges in the Production and Analysis of Illumina Sequencing Data.," BMC Genomics 12 (2011): 382, doi:10.1186/1471-2164-12-382.
Supplementary Figure 2 De novo reference creation.
De novo reference creation was performed at the species level. (1) paired-end reads are either merged using PEAR1 or concatenated (Supplementary Fig. 2), after which Watson and Crick reads are split. For concatenated pairs only the ~120 nucleotides corresponding to the read length minus the barcode on both ends can be recovered. (2) Methylation variation is removed in silico by C to T conversion in Watson and G to A conversion in Crick reads. (3) Dereplication merges identical reads. (4) For each remaining Watson and Crick read a binary AT only representation is created by simultaneous C to T and G to A conversion (binarization). (5) Identical binary representative sequences are paired, yielding AT-only pairs containing both Watson and Crick reads. (6) The original Watson and Crick reads are paired using the AT-only pairs as template. Based on the aligned Watson Crick read-pair the original non-converted sequence is reconstructed. A and T are identical in the alignment, Watson G / Crick A yields a G whereas Watson T / Crick C yields a C. (7) Finally, consensus sequences are clustered with 95% identity to yield reference sequences, allowing for insertions, substitutions and partial matches.
1 Jiajie Zhang et al., "PEAR: a Fast and Accurate Illumina Paired-End reAd mergeR.," Bioinformatics 30, no. 5 (March 1, 2014): 614–20, doi:10.1093/bioinformatics/btt593.
Supplementary Figure 3 Creation of merged and concatenated reads.
(a) A typical epiGBS library shows a peak on a Bioanalyzer image corresponding to fragments of ~300 basepairs (bp), although larger fragments are also obtained. A 300 bp fragment corresponds to an epiGBS fragment (insert) of 163 bp, as adapters are not sequenced. (b) Barcoded adapters A and B can be ligated on both ends of each fragment, so for any given genomic locus both forward and reverse oriented inserts exist. Fragments derive from either Watson or Crick strand. (c, d) Overlapping paired-end reads are merged whereas non overlapping paired-end reads are concatenated to facilitate de novo reference reconstruction. Concatenation is performed by adding 10 N’s between the forward and reverse read. Concatenated reads can be used to determine the sequence on both ends of inserts larger than ~1,9x the read length.
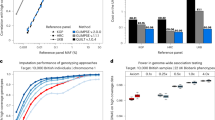
Supplementary Figure 4 Intergenerational methylation rate comparisons for different sequence contexts in epiGBS and WGBS.
Methylation rates are compared for CG, CHG and CHH context between generation 31 and 32 for WGBS vs WGBS data and WGBS vs epiGBS data. Only positions with 10 or more informative reads in both datasets were taken into account. (a) In the Becker et al1 WGBS data for line 49 we selected the subset of loci covered by PstI. Methylation rates between cytosines in generation 31 and 32 plants are well correlated. CG and CHG contexts have Pearson R2 values of respectively 0.94 and 0.92 whereas rates in CHH context show a lower R2 of 0.80. Heatmap legends on the right-hand side of all figures in panel a-c indicate the number of data points per 0.25*0.25 block in each scatter plot. (b) Methylation rates are compared in all sequence contexts between PstI-epiGBS data for line 29 generation 32 with the Becker et al WGBS data for generation 31. Compared to the WGBS vs WGBS correlations shown in panel a, the CG methylation correlates are equally good, whereas CHG and CHH correlations are slightly lower, with R2 values of 0.74 and 0.59 respectively. The methylation sensitivity of PstI in CHG context could be causing this lower correlation, as CHG methylation in Arabidopsis is spatially auto-correlated 2. Given partial methylation of CHG sites non-digestion of methylated PstI-sites could give rise to underestimating true methylation rates of cytosines inside these fragments as internal CHG and CHH methylation rates could be spatially correlated to methylation rates of the two CHG positions present in the PstI recognition site (‘5- C_TGCA^G-3’). (c) To assess whether the methylation sensitivity of PstI contributed to the lower methylation estimates and lower R2 for CHG and CHH context we have generated one additional epiGBS library with the methylation insensitive enzyme Csp6I for Arabidopsis line 29. Given the same coverage threshold (more than 10 informative reads) as used for the WGBS-WGBS and WGBS-PstI-epiGBS comparison shown in panel (a) and (b) we have obtained fewer positions with lower median coverage. This in turn leads to R2 values for CG, CHG and CHH context of 0.89, 0.76 and 0.61. (d) To test whether the lower coverage of the Csp6I epiGBS library contributed to the lower R2 values obtained compared to the WGBS data we calculated R2 values between WGBS generation 31 and subsets of the generation 32 Csp6I epiGBS data with increasingly stringent minimum read coverage criteria. R2 values increase asymptotically to the same levels as obtained for the WGBS vs WGBS comparisons made in panel a. Thus, given sufficient coverage, epiGBS estimates of methylation ratios with methylation insensitive enzymes are as accurate as WGBS based estimates. (e) Median coverage for positions in CHG context for WGBS data in line 49 gen 31 and for epiGBS data in line 29 gen 32 for PstI and Csp6I. Median coverage for other contexts was comparable to that of CHG. Median coverage was 151 in the epiGBS PstI library, whereas median coverage was 17 for the Csp6I data. (f) For positions having unequal inter- generational methylation rates we divided the positions in two groups per sequence context, one having a higher methylation estimate in epiGBS compared to WGBS whereas the other group shows the opposite pattern. For PstI, the group with positions showing a lower epiGBS rate is overrepresented for the CHG and CHH context. For Csp6I the groups are more or less of equal size, suggesting that the methylation sensitivity of PstI contributed to the overall lower methylation estimates obtained for CHG and CHH context.
1. Claude Becker et al., "Spontaneous Epigenetic Variation in the Arabidopsis Thaliana Methylome.," Nature 480, no. 7376 (December 8, 2011): 245–49, doi:10.1038/nature10555.
2. Shawn J Cokus et al., "Shotgun Bisulphite Sequencing of the Arabidopsis Genome Reveals DNA Methylation Patterning.," Nature 452, no. 7184 (March 13, 2008): 215–19, doi:10.1038/nature06745.
Supplementary Figure 5 Venn diagrams of DMPs detected in pairwise between-line comparisons of PstI-epiGBS versus WGBS data and of WGBS versus WGBS data.
Venn diagrams of DMPs detected in pairwise between-line comparisons (a) Pair-wise comparisons show the overlap and differences between DMPs detected between four genotypes using both WGBS generation 31 and epiGBS generation 32 data. Per diagram, only positions having sufficient informative read coverage in both WGBS and epiGBS data were taken into account. (b) Pair-wise comparisons show the overlap and differences between DMPs detected between the Arabidopsis line 39 and 49 using both WGBS generation 31 and WGBS generation 32 data. Per diagram, only positions having sufficient informative read coverage in both WGBS generation 31 and WGBS generation 32 data were taken into account.
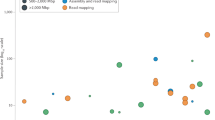
Supplementary Figure 6 Average coverage and methylation rate of the Arabidopsis thaliana genome as measured with PstI-epiGBS data.
Per cytosine, average coverage and methylation rate were calculated per 1MB windows for all four Arabidopsis lines sequenced in PstI epiGBS for the Arabidopsis reference genome TAIR10 on chromosome 1-5. Only positions covered by at least 10 reads were taken into account. Average methylation is higher in centromeric regions but per cytosine coverage is lower in these regions, reflecting sensitivity of PstI to 5mC methylation.
Supplementary Figure 7 Visualization of DNA methylation differences in the IGV Genome Browser.
For both Watson and Crick strands the coverage and polymorphic sites are shown for all samples combined. Every line containing white and red blocks represents an individual. In this example, Group “G” has a lower average methylation level for this cluster compared to group “S”. Per cytosine the methylation level is calculated and visualized. Dark red blocks indicate full methylation whereas white blocks indicate no methylation is present.
Supplementary Figure 8 Percentage of positions identified as DMPs in CHG and CG contexts.
The upper panel shows the percentage of cytosines in CHG context detected as DMP, whereas the lower panel shows the same for cytosines in CG context. Results are split for gene related and non-gene related clusters. Data for all positions per species is added as an excel file (Supplementary data 2). DMP detection was carried out as described in online methods section “DMP detection in all species”. For Arabidopsis we found that 11% of cytosines in CG context were differentially methylated in genes and 8% in non-genes. These values are lower than the 17.8% of cytosines in CG context reported as DMPs in Becker et al 2011, We suspect that our values are lower due to the stringent selection criteria requiring methylation to be present in both forward and reverse reads on both Watson and Crick strand. This might lead to under-estimation of actual DMP levels at lower sequencing coverage. Indeed, by selecting a higher minimum read coverage threshold of 50 forward and 50 reverse informative reads the percentage of CG dinucleotides found in genes was higher with 14.5 % and in non- genes 11.3 %. For most species, CG-DMPs are more prevalent in genes, with the exception of Scabiosa columbaria and Daucus carota, for which more DMPs were found in clusters not having significant blast hits to genes. CHG DMPs are absent in Daphnia magna and are most prevalent in Taraxacum officinale, consistent with the observations made for the absence of CHG methylation in Daphnia magna and the higher presence of CHG methylation in Taraxacum officinale (see Fig. 2b).
Supplementary Figure 9 Nick translation.
(a) Due to the use of non-phosphorylated adapters, epiGBS libraries contain nicks between the 3’ fragment overhang and the 5’ non-phosphorylated adapter nucleotide (on the bottom strand in this Figure). To prevent the partial loss of adapter sequence during bisulfite treatment the nick was repaired by DNA polymerase I. DNA polymerase I is able to repair single stranded nicks in dsDNA in two steps. (b) first it removes the nucleotide (in this case C) at the 5’ side of the nick by 5’-3’ exonuclease activity after which (c) the 3’ hydroxyl terminus is elongated with a single dNTP (in this case 5mC) using the nucleotide from the opposite strand (here G) as template. DNA polymerase I moves one nucleotide downstream towards the 3’ end of the bottom adapter strand. Steps (b) and (c) are repeated until the 3’ end of the adapter is reached, leaving no nicks in the adapter sequence.
Supplementary Figure 10 Bioinformatics protocol.
A) Mapping
(1) Because part of the sequenced fragment originates from the methylated adapter sequence, this part needs to be excluded from the analysis. Therefore, the first 4 bases of merged and forward Watson reads as well as the last 4 of merged and first four of reverse Crick reads are removed.
(2) Reads are mapped against either de novo obtained or existing reference using BWA-METH1 as it is more sensitive / accurate compared to similar bisulfite sequence-aligners and allows for easy transfer or read group tags from the sequence name to the bam output file1.
B) Variant calling
(3) Variant calling is done using Freebayes2 separately for both Watson and Crick reads for all samples simultaneously. The settings force every position to be called for all samples. Freebayes is used because it allows for indel realignment and sensitive variant calling.
(4) By simultaneously iterating over both Watson and Crick variant call files (VCF) SNPs and methylation polymorphisms can be distinguished. C/T polymorphisms in Watson combined with C on the Crick strand indicate a methylation polymorphism on the Watson strand whereas a G/A polymorphism on Crick combined with a G on the Watson strand indicates a methylation polymorphism on the Crick strand. Where combined SNP and methylation lead to a C/T or G/A polymorphism on both Watson and Crick strand only the SNP is called as the methylation ratio cannot be determined.
C) Visualization
(5) SNPs and methylation polymorphisms are exported in VCF format. Furthermore, methylation ratios are calculated and exported in a tab separated IGV specific format (.IGV) including the context (CG, CHG or CHH). Datasets for all species studied are available on genomespace (see https://gsui.genomespace.org/jsui/gsui.html?pathOrUrl=/Home/thomasvangurp/epiGBS%20Nature%20Methods/).
D) Annotation
(6) Usearch blastx3 is performed against reference protein sequences related to the species sequenced.
(7) Resulting blastx reads are imported into blast2go for mapping to gene ontology terms and enzyme codes. A list of all annotated genes is exported. Optionally, a list of contigs mapping to specific genes or with specific GO terms can be exported allowing for a focused analysis in RnBeads4.
E) Analysis
(8) A pipeline for processing the sample specific methylation bed files and experimental details like treatment and/or sample groups allows for comparing differential methylation between combinations of treatments, including all possible 2-way interactions between treatment groups using RnBeads4 or other methylation analysis tools.
1 Brent S Pedersen et al., "Fast and Accurate Alignment of Long Bisulfite-Seq Reads," arXiv:1401.1129v2, January 6, 2014.
2 Erik Garrison and Gabor Marth, "Haplotype-Based Variant Detection From Short-Read Sequencing," arXiv:1207.3907v2, July 17, 2012.
3 Robert C Edgar, "Search and Clustering Orders of Magnitude Faster Than BLAST.," Bioinformatics 26, no. 19 (October 1, 2010): 2460–61, doi:10.1093/bioinformatics/btq461.
4 Yassen Assenov et al., "Comprehensive Analysis of DNA Methylation Data with RnBeads.," Nature Methods 11, no. 11 (November 2014): 1138–40, doi:10.1038/nmeth.3115.
Supplementary Figure 11 De novo clustering algorithm.
1)
a) Paired-end reads contain in-line barcodes in both forward and reverse reads. Barcodes are removed while sample specific tags are added to the read name. The original strand, Watson or Crick, is determined based on the (non)conversion of the enzyme recognition site at both reads (Fig. 1b).
b) Overlapping paired-end reads are merged using Pear1 to (i) avoid adapter remnants for reads derived from fragment with an insert size shorter than the number of read cycles as they can lead to false polymorphisms or mapping errors, (ii) avoid double calling of bases from the same molecule in the overlapping region of the pair, and (iii) lowering the error rate by selecting the highest quality base from the two reads in the overlapping region. Non-overlapping reads are concatenated to allow for matching Watson and Crick pairs (Supplementary Fig. 3).
2) Methylation polymorphisms are removed by converting all C’s in Watson to T and all G’s in Crick to A to allow for clustering of same-origin reads with methylation polymorphisms.
3) Identical Watson and Crick reads are dereplicated using usearch2. Only Watson and Crick reads that occur twice or more are retained to avoid sequencing errors.
4) Binarization: To aid pairing of same-origin Watson and Crick reads all C’s are converted to T’s in Watson reads while G’s are converted to A’s in Crick reads rendering a binary AT only sequence output, that is identical for same origin Watson and Crick reads (see Fig. 1b).
5) Using usearch dereplication binary AT only watson and crick reads are paired.
6) Based on the alignment and orientation of the binary Watson and Crick reads the original unconverted Watson and Crick reads are aligned in a bam file. The reference sequence of each pair is called using combined bash piped queries using the following components of samtools3: mpileup and bcftools for variant calling followed by vcfutils.pl vcf2fq for creating the Watson and Crick consensus sequence. These consensus sequences are used to recreate the original sequence. In case of Watson:T / Crick:C a C is added, whereas in case of Watson:G / Crick:A a G is added. In all other cases where nucleotides do not match an ambiguous nucleotide (N) is added.
7) Consensus sequences with 95 % ore higher identity are clustered, leading to the final reference sequence.
1 Jiajie Zhang et al., "PEAR: a Fast and Accurate Illumina Paired-End reAd mergeR.," Bioinformatics 30, no. 5 (March 1, 2014): 614–20, doi:10.1093/bioinformatics/btt593.
2 Robert C Edgar, "Search and Clustering Orders of Magnitude Faster Than BLAST.," Bioinformatics 26, no. 19 (October 1, 2010): 2460–61, doi:10.1093/bioinformatics/btq461.
3 H. Li et al., "The Sequence Alignment/Map Format and SAMtools," Bioinformatics 25, no. 16 (August 7, 2009): 2078–79, doi:10.1093/bioinformatics/btp352.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11 (PDF 1960 kb)
Supplementary Data 1
Detailed sample descriptions (XLSX 21 kb)
Supplementary Data 2
DMPs of all species (XLSX 2844 kb)
Supplementary Data 3
Data for Supplementary Figures (XLSX 9378 kb)
Source data
Rights and permissions
About this article
Cite this article
van Gurp, T., Wagemaker, N., Wouters, B. et al. epiGBS: reference-free reduced representation bisulfite sequencing. Nat Methods 13, 322–324 (2016). https://doi.org/10.1038/nmeth.3763
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3763
This article is cited by
-
Exploitation of epigenetic variation of crop wild relatives for crop improvement and agrobiodiversity preservation
Theoretical and Applied Genetics (2022)
-
Epigenetics of wheat–rust interaction: an update
Planta (2022)
-
The methylome of Biomphalaria glabrata and other mollusks: enduring modification of epigenetic landscape and phenotypic traits by a new DNA methylation inhibitor
Epigenetics & Chromatin (2021)
-
Family-effects in the epigenomic response of red blood cells to a challenge test in the European sea bass (Dicentrarchus labrax, L.)
BMC Genomics (2021)
-
AlphaBeta: computational inference of epimutation rates and spectra from high-throughput DNA methylation data in plants
Genome Biology (2020)