Abstract
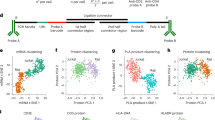
To enable the detection of expression signatures specific to individual cells, we developed PLAYR (proximity ligation assay for RNA), a method for highly multiplexed transcript quantification by flow and mass cytometry that is compatible with standard antibody staining. When used with mass cytometry, PLAYR allowed for the simultaneous quantification of more than 40 different mRNAs and proteins. In primary cells, we quantified multiple transcripts, with the identity and functional state of each analyzed cell defined on the basis of the expression of a separate set of transcripts or proteins. By expanding high-throughput deep phenotyping of cells beyond protein epitopes to include RNA expression, PLAYR opens a new avenue for the characterization of cellular metabolism.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Jaitin, D.A. et al. Massively parallel single-cell RNA-seq for marker-free decomposition of tissues into cell types. Science 343, 776–779 (2014).
Hashimshony, T., Wagner, F., Sher, N. & Yanai, I. CEL-Seq: single-cell RNA-Seq by multiplexed linear amplification. Cell Rep. 2, 666–673 (2012).
Islam, S. et al. Highly multiplexed and strand-specific single-cell RNA 5′ end sequencing. Nat. Protoc. 7, 813–828 (2012).
Islam, S. et al. Quantitative single-cell RNA-seq with unique molecular identifiers. Nat. Methods 11, 163–166 (2014).
Ramsköld, D. et al. Full-length mRNA-seq from single-cell levels of RNA and individual circulating tumor cells. Nat. Biotechnol. 30, 777–782 (2012).
Sasagawa, Y. et al. Quartz-Seq: a highly reproducible and sensitive single-cell RNA sequencing method, reveals non-genetic gene-expression heterogeneity. Genome Biol. 14, R31 (2013).
Shalek, A.K. et al. Single-cell transcriptomics reveals bimodality in expression and splicing in immune cells. Nature 498, 236–240 (2013).
Wu, A.R. et al. Quantitative assessment of single-cell RNA-sequencing methods. Nat. Methods 11, 41–46 (2014).
Deng, Q., Ramsköld, D., Reinius, B. & Sandberg, R. Single-cell RNA-seq reveals dynamic, random monoallelic gene expression in mammalian cells. Science 343, 193–196 (2014).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Tang, F., Lao, K. & Surani, M.A. Development and applications of single-cell transcriptome analysis. Nat. Methods 8, S6–S11 (2011).
Fan, H.C., Fu, G.K. & Fodor, S.P.A. Expression profiling. Combinatorial labeling of single cells for gene expression cytometry. Science 347, 1258367 (2015).
Dvinge, H. et al. Sample processing obscures cancer-specific alterations in leukemic transcriptomes. Proc. Natl. Acad. Sci. USA 111, 16802–16807 (2014).
Grün, D. & van Oudenaarden, A. Design and analysis of single-cell sequencing experiments. Cell 163, 799–810 (2015).
Bauman, J.G., Bayer, J.A. & van Dekken, H. Fluorescent in-situ hybridization to detect cellular RNA by flow cytometry and confocal microscopy. J. Microsc. 157, 73–81 (1990).
Patterson, B.K. et al. Detection of HIV-1 DNA and messenger RNA in individual cells by PCR-driven in situ hybridization and flow cytometry. Science 260, 976–979 (1993).
Belloc, F. & Durrieu, F. Detection of mRNA species by flow cytometry. Methods Cell Biol. 42, 59–69 (1994).
Borzì, R.M. et al. A fluorescent in situ hybridization method in flow cytometry to detect HIV-1 specific RNA. J. Immunol. Methods 193, 167–176 (1996).
Lalli, E., Gibellini, D., Santi, S. & Facchini, A. In situ hybridization in suspension and flow cytometry as a tool for the study of gene expression. Anal. Biochem. 207, 298–303 (1992).
Just, T., Burgwald, H. & Broe, M.K. Flow cytometric detection of EBV (EBER snRNA) using peptide nucleic acid probes. J. Virol. Methods 73, 163–174 (1998).
Larsson, C., Grundberg, I., Söderberg, O. & Nilsson, M. In situ detection and genotyping of individual mRNA molecules. Nat. Methods 7, 395–397 (2010).
Weibrecht, I. et al. In situ detection of individual mRNA molecules and protein complexes or post-translational modifications using padlock probes combined with the in situ proximity ligation assay. Nat. Protoc. 8, 355–372 (2013).
Player, A.N., Shen, L.P., Kenny, D., Antao, V.P. & Kolberg, J.A. Single-copy gene detection using branched DNA (bDNA) in situ hybridization. J. Histochem. Cytochem. 49, 603–612 (2001).
Porichis, F. et al. High-throughput detection of miRNAs and gene-specific mRNA at the single-cell level by flow cytometry. Nat. Commun. 5, 5641 (2014).
Bendall, S.C. et al. Single-cell mass cytometry of differential immune and drug responses across a human hematopoietic continuum. Science 332, 687–696 (2011).
Lin, Y., Sohn, C.H., Dalal, C.K., Cai, L. & Elowitz, M.B. Combinatorial gene regulation by modulation of relative pulse timing. Nature 527, 54–58 (2015).
Chubb, J.R., Trcek, T., Shenoy, S.M. & Singer, R.H. Transcriptional pulsing of a developmental gene. Curr. Biol. 16, 1018–1025 (2006).
Cai, L., Friedman, N. & Xie, X.S. Stochastic protein expression in individual cells at the single molecule level. Nature 440, 358–362 (2006).
Fredriksson, S. et al. Protein detection using proximity-dependent DNA ligation assays. Nat. Biotechnol. 20, 473–477 (2002).
Söderberg, O. et al. Direct observation of individual endogenous protein complexes in situ by proximity ligation. Nat. Methods 3, 995–1000 (2006).
Lizardi, P.M. et al. Mutation detection and single-molecule counting using isothermal rolling-circle amplification. Nat. Genet. 19, 225–232 (1998).
Leuchowius, K.-J. et al. Parallel visualization of multiple protein complexes in individual cells in tumor tissue. Mol. Cell. Proteomics 12, 1563–1571 (2013).
Untergasser, A. et al. Primer3—new capabilities and interfaces. Nucleic Acids Res. 40, e115 (2012).
Jurka, J. et al. Repbase Update, a database of eukaryotic repetitive elements. Cytogenet. Genome Res. 110, 462–467 (2005).
O'Gorman, W.E. et al. Single-cell systems-level analysis of human Toll-like receptor activation defines a chemokine signature in patients with systemic lupus erythematosus. J. Allergy Clin. Immunol. 136, 1326–1336 (2015).
Newell, E.W., Sigal, N., Bendall, S.C., Nolan, G.P. & Davis, M.M. Cytometry by time-of-flight shows combinatorial cytokine expression and virus-specific cell niches within a continuum of CD8+ T cell phenotypes. Immunity 36, 142–152 (2012).
Amir, el-A.D. et al. viSNE enables visualization of high dimensional single-cell data and reveals phenotypic heterogeneity of leukemia. Nat. Biotechnol. 31, 545–552 (2013).
DeForge, L.E. & Remick, D.G. Kinetics of TNF, IL-6, and IL-8 gene expression in LPS-stimulated human whole blood. Biochem. Biophys. Res. Commun. 174, 18–24 (1991).
Agarwal, S., Piesco, N.P., Johns, L.P. & Riccelli, A.E. Differential expression of IL-1 beta, TNF-alpha, IL-6, and IL-8 in human monocytes in response to lipopolysaccharides from different microbes. J. Dent. Res. 74, 1057–1065 (1995).
Irish, J.M. et al. Single cell profiling of potentiated phospho-protein networks in cancer cells. Cell 118, 217–228 (2004).
Gaudillière, B. et al. Clinical recovery from surgery correlates with single-cell immune signatures. Sci. Transl. Med. 6, 255ra131 (2014).
Jaye, D.L., Bray, R.A., Gebel, H.M., Harris, W.A.C. & Waller, E.K. Translational applications of flow cytometry in clinical practice. J. Immunol. 188, 4715–4719 (2012).
Kaleem, Z. et al. Flow cytometric analysis of acute leukemias. Diagnostic utility and critical analysis of data. Arch. Pathol. Lab. Med. 127, 42–48 (2003).
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096–1098 (2013).
Schulte, R. et al. Index sorting resolves heterogeneous murine hematopoietic stem cell populations. Exp. Hematol. 43, 803–811 (2015).
Wilson, N.K. et al. Combined single-cell functional and gene expression analysis resolves heterogeneity within stem cell populations. Cell Stem Cell 16, 712–724 (2015).
Angelo, M. et al. Multiplexed ion beam imaging of human breast tumors. Nat. Med. 20, 436–442 (2014).
Wernig, M. et al. A drug-inducible transgenic system for direct reprogramming of multiple somatic cell types. Nat. Biotechnol. 26, 916–924 (2008).
Krutzik, P.O. & Nolan, G.P. Intracellular phospho-protein staining techniques for flow cytometry: monitoring single cell signaling events. Cytometry A 55, 61–70 (2003).
Zunder, E.R. et al. Palladium-based mass tag cell barcoding with a doublet-filtering scheme and single-cell deconvolution algorithm. Nat. Protoc. 10, 316–333 (2015).
Acknowledgements
We thank L. Lanier (UCSF, San Francisco, California, USA) for providing the NKL cell line, and A. Trejo and A. Jager for technical assistance. A.P.F. is supported by a Fellowship for Prospective Researchers from the Swiss National Science Foundation, an EMBO Long-Term Fellowship and a Marie Curie International Outgoing Fellowship. P.F.G. is a Howard Hughes Medical Institute Fellow of the Life Sciences Research Foundation. F.-A.B. is supported by a Human Frontier Science Program Long-Term Fellowship. This work was supported by the US National Institutes of Health (grants U19 AI057229, 1U19AI100627, R01CA184968, 1R33CA183654-01, R33CA183692, 1R01GM10983601, 1R21CA183660, 1R01NS08953301, OPP1113682, 5UH2AR067676, 1R01CA19665701 and R01HL120724 to G.P.N.), US Department of Defense Congressionally Directed Medical Research Programs (grants OC110674 and 11491122 to G.P.N.), the Northrop-Grumman Corporation (G.P.N.) and the Rachford & Carlotta A. Harris Endowed Chair (G.P.N.).
Author information
Authors and Affiliations
Contributions
A.P.F., F.-A.B. and P.F.G. conceived the work, performed experiments, analyzed data and wrote the manuscript. E.R.Z. provided help with mouse embryonic stem cell experiments. E.W.Y.H. and S.-Y.C. provided help with cytokine induction experiments. G.P.N. supervised the work and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
G.P.N. has a personal financial interest in the company Fluidigm, the manufacturer of the mass cytometer used in this study.
Integrated supplementary information
Supplementary Figure 1 Validation of different PLAYR insert systems used for multiplexed detection of transcripts.
The three transcripts ACTB, GAPDH, and PPIA were detected using 2 probe pairs per gene on 7 different insert systems used for multiplexed detection of transcripts. Rolling Circle Amplification was carried out for 30min, 1h, 2h, 4h or over night (16h) and samples were analyzed by mass cytometry.
Supplementary Figure 2 Graphical display of the PLAYRDesign software tool for user-friendly design of PLAYR probe pairs.
Each potential probe is represented by a red rectangle. The Primer3 score of each probe is represented by a color gradient from light pink to red, where red probes have higher scores and are preferred over light red probes. The position of probes along the transcript is represented together with sequence features that can guide probe selection. Different graphs represent: maximum sequence identity of BLAST matches to other transcripts (blue top), maximum sequence identity of BLAST matches to a database of repetitive sequences (red); predicted melting temperature in a window of 20 residues (green); number of ESTs that skip an exon but include the exons flanking it (blue bottom). The actual melting temperature of probes is independently calculated by Primer3, while the purpose of the green graph is to give an indication on whether certain regions of the transcript have a melting temperature that is too low or too high to be amenable for probe design. Blue and red graphs represent sequence features that are not considered in the scoring of Primer3 probes. Secondary RNA structure is not considered for probe design and it is always recommended to simultaneously use multiple probe pairs that target different regions of a transcript to increase the sensitivity of the assay and to minimize experimental artifacts due to transcript accessibility. The software does not explicitly deal with alternative splicing. If one is interested in targeting a specific splice variant the best approach is to use the sequence of the isoform as input to the program. The aim of displaying exons that are skipped in publicly available EST data is to give the user the possibility to primarily target constitutive exons. Repetitive sequences should always be avoided during probe design since probe pairs can otherwise lead to unspecific signals.
Supplementary Figure 3 Specificity control experiments for PLAYR.
Detection of the ACTB transcript in Jurkat cells. No signal is detected when PLAYR is performed in absence of probes (NO PROBES), in absence of insert (NO INSERT), in absence of backbone (NO BACKBONE), in absence of ligase (NO LIGATION), in absence of detection oligo (NO DETECTION OLIGO), in presence of probes directed against the anti-sense ACTB transcript (SENSE PROBES), in presence of probes with the same half of the insert-complementary sequence (ORIENTATION CONTROL), or in presence of non-cognate probe pairs targeting different transcripts (ACTB and GAPDH, GENE-SPECIFICITY CONTROL). Signals were detected by flow cytometry. 4 probe pairs were used per gene.
Supplementary Figure 4 Detection of specific transcripts in single cells by flow cytometry using multiple probe pairs.
a) Detection of CD10 and CD3E by PLAYR in three replicates. Jurkat and NALM-6 cells were incubated with the indicated number of probe pairs and analyzed by flow cytometry. The expression of CD3E and CD10 is specific to Jurkat and NALM-6 cells respectively. b) The intensity of PLAYR signals depends on the distance between PLAYR probe binding sites on a target transcript. Multiple adjacent probe pairs spanning a transcript were designed and tested in all possible pairwise combinations. The x-axis represents the distance between each pair of probes, and the y-axis represents the ratio between the signal obtained with a given combination and the signal obtained with the corresponding adjacent cognate probe (i.e., the one that was originally designed to be used in the pair).
Supplementary Figure 5 Protocol optimization for RNA detection with simultaneous antibody staining.
Several surface antigens are disrupted by methanol permeabilization, which requires antibody staining to be performed before this step in the PLAYR protocol. In addition, we have found that crosslinking with the BS3 crosslinker is necessary to preserve antibody staining during the PLAYR protocol, as determined in multiple experiments with different antibodies (data not shown). However, RNA is progressively degraded during the incubations that are necessary to perform the steps mentioned above. This can be avoided by permeabilization of cells in the presence of RNAse inhibitors that inhibit endogenous and exogenous RNAses. Shown are signal intensities for the ACTB transcript in monocytes as measured by PLAYR when antibody staining is performed in different buffers containing RNAsin Ribonuclease Inhibitor (RNAsin) only or in combination with ribonucleoside vanadyl complex (RVC) and polyvinylsulfonic acid (PVS). BD Perm/Wash buffer is a commercially available saponin-based buffer and was replaced with PBS + 2% saponin that performed comparably. Signals obtained when cells are permeabilized directly without antibody staining are shown as a control. We decided to omit RVC in this step from the final protocol because the reagent interferes with the staining of some antibodies. Experiments were performed on a fluorescence flow cytometer with healthy PBMCs and monocytes were gated based on Forward and Side Scatter.
Supplementary Figure 6 Reproducibility of PLAYR measurements across technical replicates.
The plot displays the quantification of 22 different transcripts including negative controls across 16 technical replicates in mouse embryonic stem cells as measured by mass cytometry. Points represent mean values of replicates and bars indicate the full range of values for each transcript.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 (PDF 917 kb)
Supplementary Table 1
PLAYR probes and backbone-insert systems (XLSX 90 kb)
Supplementary Software
PLAYR Design software for probe design (ZIP 72 kb)
Rights and permissions
About this article
Cite this article
Frei, A., Bava, FA., Zunder, E. et al. Highly multiplexed simultaneous detection of RNAs and proteins in single cells. Nat Methods 13, 269–275 (2016). https://doi.org/10.1038/nmeth.3742
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3742
This article is cited by
-
The CRISPR-Cas13a Gemini System for noncontiguous target RNA activation
Nature Communications (2024)
-
Extending the dynamic range of biomarker quantification through molecular equalization
Nature Communications (2023)
-
CODEX multiplexed tissue imaging with DNA-conjugated antibodies
Nature Protocols (2021)
-
Functional patient-derived cellular models for neuropsychiatric drug discovery
Translational Psychiatry (2021)
-
Extended live-cell barcoding approach for multiplexed mass cytometry
Scientific Reports (2021)