Abstract
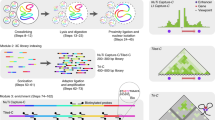
Methods for analyzing chromosome conformation in mammalian cells are either low resolution or low throughput and are technically challenging. In next-generation (NG) Capture-C, we have redesigned the Capture-C method to achieve unprecedented levels of sensitivity and reproducibility. NG Capture-C can be used to analyze many genetic loci and samples simultaneously. High-resolution data can be produced with as few as 100,000 cells, and single-nucleotide polymorphisms can be used to generate allele-specific tracks. The method is straightforward to perform and should greatly facilitate the investigation of many questions related to gene regulation as well as the functional dissection of traits examined in genome-wide association studies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Wang, Z., Gerstein, M. & Snyder, M. RNA-seq: a revolutionary tool for transcriptomics. Nat. Rev. Genet. 10, 57–63 (2009).
Mikkelsen, T.S. et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448, 553–560 (2007).
Robertson, G. et al. Genome-wide profiles of STAT1 DNA association using chromatin immunoprecipitation and massively parallel sequencing. Nat. Methods 4, 651–657 (2007).
Hesselberth, J.R. et al. Global mapping of protein-DNA interactions in vivo by digital genomic footprinting. Nat. Methods 6, 283–289 (2009).
Buenrostro, J.D., Giresi, P.G., Zaba, L.C., Chang, H.Y. & Greenleaf, W.J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
Tolhuis, B., Palstra, R.J., Splinter, E., Grosveld, F. & de Laat, W. Looping and interaction between hypersensitive sites in the active beta-globin locus. Mol. Cell 10, 1453–1465 (2002).
Noordermeer, D. et al. The dynamic architecture of Hox gene clusters. Science 334, 222–225 (2011).
Sanyal, A., Lajoie, B.R., Jain, G. & Dekker, J. The long-range interaction landscape of gene promoters. Nature 489, 109–113 (2012).
van de Werken, H.J. et al. Robust 4C-seq data analysis to screen for regulatory DNA interactions. Nat. Methods 9, 969–972 (2012).
de Laat, W. & Duboule, D. Topology of mammalian developmental enhancers and their regulatory landscapes. Nature 502, 499–506 (2013).
Hughes, J.R. et al. Analysis of hundreds of cis-regulatory landscapes at high resolution in a single, high-throughput experiment. Nat. Genet. 46, 205–212 (2014).
Pasquali, L. et al. Pancreatic islet enhancer clusters enriched in type 2 diabetes risk-associated variants. Nat. Genet. 46, 136–143 (2014).
Maurano, M.T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
Parker, S.C. et al. Chromatin stretch enhancer states drive cell-specific gene regulation and harbor human disease risk variants. Proc. Natl. Acad. Sci. USA 110, 17921–17926 (2013).
Rao, S.S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Jäger, R. et al. Capture Hi-C identifies the chromatin interactome of colorectal cancer risk loci. Nat. Commun. 6, 6178 (2015).
Schoenfelder, S. et al. The pluripotent regulatory circuitry connecting promoters to their long-range interacting elements. Genome Res. 25, 582–597 (2015).
Vernimmen, D., De Gobbi, M., Sloane-Stanley, J.A., Wood, W.G. & Higgs, D.R. Long-range chromosomal interactions regulate the timing of the transition between poised and active gene expression. EMBO J. 26, 2041–2051 (2007).
Hughes, J.R. et al. High-resolution analysis of cis-acting regulatory networks at the alpha-globin locus. Phil. Trans. R. Soc. Lond. B 368, 20120361 (2013).
Baù, D. et al. The three-dimensional folding of the α-globin gene domain reveals formation of chromatin globules. Nat. Struct. Mol. Biol. 18, 107–114 (2011).
Simonis, M. et al. Nuclear organization of active and inactive chromatin domains uncovered by chromosome conformation capture-on-chip (4C). Nat. Genet. 38, 1348–1354 (2006).
Kang, J.H. et al. Genomic organization, tissue distribution and deletion mutation of human pyridoxine 5′-phosphate oxidase. Eur. J. Biochem. 271, 2452–2461 (2004).
Love, M.I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Klein, F.A. et al. FourCSeq: analysis of 4C sequencing data. Bioinformatics 31, 3085–3091 (2015).
Thongjuea, S., Stadhouders, R., Grosveld, F.G., Soler, E. & Lenhard, B. r3Cseq: an R/Bioconductor package for the discovery of long-range genomic interactions from chromosome conformation capture and next-generation sequencing data. Nucleic Acids Res. 41, e132 (2013).
Osborne, C.S. et al. Active genes dynamically colocalize to shared sites of ongoing transcription. Nat. Genet. 36, 1065–1071 (2004).
Noordermeer, D. et al. Variegated gene expression caused by cell-specific long-range DNA interactions. Nat. Cell Biol. 13, 944–951 (2011).
Bernet, A. et al. Targeted inactivation of the major positive regulatory element (HS-40) of the human alpha-globin gene locus. Blood 86, 1202–1211 (1995).
Anguita, E. et al. Deletion of the mouse alpha-globin regulatory element (HS-26) has an unexpectedly mild phenotype. Blood 100, 3450–3456 (2002).
de Wit, E. & de Laat, W. A decade of 3C technologies: insights into nuclear organization. Genes Dev. 26, 11–24 (2012).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Magoč, T. & Salzberg, S.L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Kent, W.J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Raney, B.J. et al. Track data hubs enable visualization of user-defined genome-wide annotations on the UCSC Genome Browser. Bioinformatics 30, 1003–1005 (2014).
Acknowledgements
J.O.J.D. thanks the Wellcome Trust for funding his work (Wellcome Trust Clinical Research Training Fellowship reference 098931/Z/12/Z). The work was also supported by a Wellcome Trust Strategic Award (reference 106130/Z/14/Z) and the Medical Research Council (MRC Core Funding and Centenary Award reference 4050189188). We thank E. Repapi for statistical advice and L. Hanssen, M. Oudelaar, D. Jeziorska, B. Graham, M. Kassouf, M. Suciu, H. Long, S. Pasricha, V. Buckle, T. Milne, T. Fulga, T. Sauka-Spengler, D. Downes and A. Drakesmith for their critique of the manuscript. We thank S. Thongjuea for discussions on analysis.
Author information
Authors and Affiliations
Contributions
J.O.J.D. performed the experiments, analyzed the data and wrote the manuscript. J.R.H. designed the experiments, assisted with the bioinformatic analysis and wrote the manuscript. N.A.R. assisted with the experiments. J.M.T. analyzed the data. S.J.M. and S.T. assisted with the bioinformatics analysis and prepared the software for public release. D.R.H. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–22 and Supplementary Note (PDF 20583 kb)
Supplementary Data
NG Capture-C supplementary data file (XLSX 55 kb)
Rights and permissions
About this article
Cite this article
Davies, J., Telenius, J., McGowan, S. et al. Multiplexed analysis of chromosome conformation at vastly improved sensitivity. Nat Methods 13, 74–80 (2016). https://doi.org/10.1038/nmeth.3664
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3664
This article is cited by
-
Bivalent activity of super-enhancer RNA LINC02454 controls 3D chromatin structure and regulates glioma sensitivity to temozolomide
Cell Death & Disease (2024)
-
Multiple Fra-1-bound enhancers showing different molecular and functional features can cooperate to repress gene transcription
Cell & Bioscience (2023)
-
Transcription modulates chromatin dynamics and locus configuration sampling
Nature Structural & Molecular Biology (2023)
-
Determining chromatin architecture with Micro Capture-C
Nature Protocols (2023)
-
MLL-AF4 cooperates with PAF1 and FACT to drive high-density enhancer interactions in leukemia
Nature Communications (2023)