Abstract
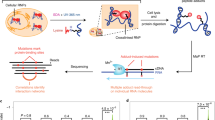
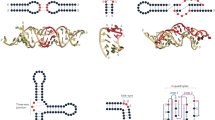
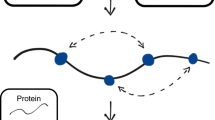
RNA regulates many biological processes; however, identifying functional RNA sequences and structures is complex and time-consuming. We introduce a method, mutational interference mapping experiment (MIME), to identify, at single-nucleotide resolution, the primary sequence and secondary structures of an RNA molecule that are crucial for its function. MIME is based on random mutagenesis of the RNA target followed by functional selection and next-generation sequencing. Our analytical approach allows the recovery of quantitative binding parameters and permits the identification of base-pairing partners directly from the sequencing data. We used this method to map the binding site of the human immunodeficiency virus-1 (HIV-1) Pr55Gag protein on the viral genomic RNA in vitro, and showed that, by analyzing permitted base-pairing patterns, we could model RNA structure motifs that are crucial for protein binding.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Wan, Y., Kertesz, M., Spitale, R.C., Segal, E. & Chang, H.Y. Understanding the transcriptome through RNA structure. Nat. Rev. Genet. 12, 641–655 (2011).
Tome, J.M. et al. Comprehensive analysis of RNA-protein interactions by high-throughput sequencing–RNA affinity profiling. Nat. Methods 11, 683–688 (2014).
Lambert, N. et al. RNA Bind-n-Seq: quantitative assessment of the sequence and structural binding specificity of RNA-binding proteins. Mol. Cell 54, 887–900 (2014).
Buenrostro, J.D. et al. Quantitative analysis of RNA-protein interactions on a massively parallel array reveals biophysical and evolutionary landscapes. Nat. Biotechnol. 32, 562–568 (2014).
Fowler, D.M. et al. High-resolution mapping of protein sequence–function relationships. Nat. Methods 7, 741–746 (2010).
Pitt, J.N. & Ferré-D'Amaré, A.R. Rapid construction of empirical RNA fitness landscapes. Science 330, 376–379 (2010).
Zykovich, A., Korf, I. & Segal, D.J. Bind-n-Seq: high-throughput analysis of in vitro protein-DNA interactions using massively parallel sequencing. Nucleic Acids Res. 37, e151 (2009).
Guenther, U.-P. et al. Hidden specificity in an apparently nonspecific RNA-binding protein. Nature 502, 385–388 (2013).
Auyeung, V.C., Ulitsky, I., McGeary, S.E. & Bartel, D.P. Beyond secondary structure: primary-sequence determinants license pri-miRNA hairpins for processing. Cell 152, 844–858 (2013).
D'Souza, V. & Summers, M.F. How retroviruses select their genomes. Nat. Rev. Microbiol. 3, 643–655 (2005).
Didierlaurent, L. et al. Role of HIV-1 RNA and protein determinants for the selective packaging of spliced and unspliced viral RNA and host U6 and 7SL RNA in virus particles. Nucleic Acids Res. 39, 8915–8927 (2011).
McBride, M.S., Schwartz, M.D. & Panganiban, A.T. Efficient encapsidation of human immunodeficiency virus type 1 vectors and further characterization of cis elements required for encapsidation. J. Virol. 71, 4544–4554 (1997).
Clever, J.L., Miranda, D., Parslow, T.G., Miranda, D. Jr. & Parslow, T.G. RNA structure and packaging signals in the 5′ leader region of the human immunodeficiency virus type 1 genome. J. Virol. 76, 12381–12387 (2002).
Wilkinson, K.A. et al. High-throughput SHAPE analysis reveals structures in HIV-1 genomic RNA strongly conserved across distinct biological states. PLoS Biol. 6:e96 (2008).
Watts, J.M. et al. Architecture and secondary structure of an entire HIV-1 RNA genome. Nature 460, 711–716 (2009).
Abbink, T.E.M., Ooms, M., Haasnoot, P.C.J. & Berkhout, B. The HIV-1 leader RNA conformational switch regulates RNA dimerization but does not regulate mRNA translation. Biochemistry 44, 9058–9066 (2005).
Siegfried, N.A., Busan, S., Rice, G.M., Nelson, J.A.E. & Weeks, K.M. RNA motif discovery by SHAPE and mutational profiling (SHAPE-MaP). Nat. Methods 11, 959–965 (2014).
Paillart, J.C. et al. First snapshots of the HIV-1 RNA structure in infected cells and in virions. J. Biol. Chem. 279, 48397–48403 (2004).
Huthoff, H. & Berkhout, B. Two alternating structures of the HIV-1 leader RNA. RNA 7, 143–157 (2001).
Stephenson, J.D. et al. Three-dimensional RNA structure of the major HIV-1 packaging signal region. Structure 21, 951–962 (2013).
Lu, K. et al. NMR detection of structures in the HIV-1 5′-leader RNA that regulate genome packaging. Science 334, 242–245 (2011).
Damgaard, C.K., Dyhr-Mikkelsen, H. & Kjems, J. Mapping the RNA-binding sites for human immunodeficiency virus type-1 gag and NC proteins within the complete HIV-1 and -2 untranslated leader regions. Nucleic Acids Res. 26, 3667–3676 (1998).
Skripkin, E., Paillart, J.C., Marquet, R., Ehresmann, B. & Ehresmann, C. Identification of the primary site of the human immunodeficiency virus type 1 RNA dimerization in vitro. Proc. Natl. Acad. Sci. USA 91, 4945–4949 (1994).
Paillart, J.C., Skripkin, E., Ehresmann, B., Ehresmann, C. & Marquet, R. A loop-loop 'kissing' complex is the essential part of the dimer linkage of genomic HIV-1 RNA. Proc. Natl. Acad. Sci. USA 93, 5572–5577 (1996).
Paillart, J.-C., Shehu-Xhilaga, M., Marquet, R. & Mak, J. Dimerization of retroviral RNA genomes: an inseparable pair. Nat. Rev. Microbiol. 2, 461–472 (2004).
Nikolaitchik, O., Rhodes, T.D., Ott, D. & Hu, W.S. Effects of mutations in the human immunodeficiency virus type 1 gag gene on RNA packaging and recombination. J. Virol. 80, 4691–4697 (2006).
Heng, X. et al. Identification of a minimal region of the HIV-1 5′ leader required for RNA dimerization, NC binding and packaging. J. Mol. Biol. 417, 224–239 (2012).
Sakuragi, J., Iwamoto, A. & Shioda, T. Dissociation of genome dimerization from packaging functions and virion maturation of human immunodeficiency virus type 1. J. Virol. 76, 959–967 (2002).
Sakuragi, J.-I., Sakuragi, S. & Shioda, T. Minimal region sufficient for genome dimerization in the human immunodeficiency virus type 1 virion and its potential roles in the early stages of viral replication. J. Virol. 81, 7985–7992 (2007).
Lever, A., Gottlinger, H., Haseltine, W. & Sodroski, J. Identification of a sequence required for efficient packaging of human immunodeficiency virus type 1 RNA into virions. J. Virol. 63, 4085–4087 (1989).
Clavel, F. & Orenstein, J.M. A mutant of human immunodeficiency virus with reduced RNA packaging and abnormal particle morphology. J. Virol. 64, 5230–5234 (1990).
Abd El-Wahab, E.W. et al. Specific recognition of the HIV-1 genomic RNA by the gag precursor. Nat. Commun. 5:4304 (2014).
Kutluay, S.B. et al. Global changes in the RNA-binding specificity of HIV-1 gag regulate virion genesis. Cell 159, 1096–1109 (2014).
Houzet, L. et al. HIV controls the selective packaging of genomic, spliced viral and cellular RNAs into virions through different mechanisms. Nucleic Acids Res. 35, 2695–2704 (2007).
McKinstry, W.J. et al. Expression and purification of soluble recombinant full-length HIV-1 Pr55Gag protein in Escherichia coli. Protein Expr. Purif. 100, 10–18 (2014).
Acknowledgements
This work was supported by grants from Sidaction and the French Agence Nationale de Recherches sur le SIDA et les Hépatites Virales (ANRS) to R.M. R.P.S. was supported by fellowships from the ANRS and from the Initiative d'excellence (IDEX), Par-delà les frontières, l'Université de Strasbourg. J.M. was funded by grants from the US National Institutes of Health (NIH), the Australian National Health and Medical Research Council (NHMRC) and the Australian Research Council (ARC). M.v.K. acknowledges funding from the German Ministry for Education & Science (BMBF) through grant number 031A307 and from the Einstein Center for Mathematics Berlin (ECMath) through project CH5. We thank A. Tomasini and P. Romby (Institut de Biologie Moléculaire et Cellulaire du Centre National de la Recherche Scientifique, Université de Strasbourg) for the gift of the MS2 phage coat protein.
Author information
Authors and Affiliations
Contributions
R.P.S. and R.M. designed the study. R.P.S. generated the libraries for sequencing and performed the MIME experiments. R.P.S., L.D., F.J. and M.v.K. developed bioinformatic tools. M.v.K. developed binding models and statistical tools. M.H. and J.M. expressed and purified the Pr55Gag protein. S.B. characterized the Pr55Gag protein. G.H. and L.W. performed DNA sequencing. R.P.S., J.-C.P., M.v.K. and R.M. analyzed the data. R.P.S., M.v.K. and R.M. wrote the paper with contributions from the other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Raw mutation rate of DNA libraries
Two independent mutated libraries were generated by PCR mutagenesis. Sequencing of these mutant libraries showed that we had introduced a mutation rate (μ) of approximately 0.006 mutations per nucleotide, respectively. Coefficient of variation was 37% in both libraries. Wild-type DNA was sequenced to measure errors introduced during library preparation and sequencing. Wild-type DNA showed a mutation rate of 0.0013 mutations per nucleotide with a coefficient of variation of 117%.
Supplementary Figure 2 Comparison of relative Kd values obtained from experimental replicates
Left: Significant median Kdm(max)/Kdw values computed from technical replicate 1 (experiment 1) compared to replicate/experiment 2, showing the Pearson correlation between the quantitative Kd estimates and the corresponding P-value for assessing non-zero correlation. ‘Median’ denotes the median of the re-sampling distribution obtained after computational analysis (see eq. (6) in the Online Methods. Significance is determined using eqs. (7)-(8) in the Online Methods after multiple test correction. The vertical- and horizontal dashed lines separate Kdm(max)/Kdw estimates that are significantly larger- or smaller than 1 in the respective experiments and thus allow a qualitative assessment of the respective relative Kd estimates between experimental replicates. For example, in 1% of all depicted relative Kd estimates, replicate 2 yielded Kdm(max)/Kdw > 1 vs. Kdm(max)/Kdw < 1 in experimental replicate 1. The diagonal dashed line indicates the line of unity. Right: Cross-tabulation of the estimated number of positions significantly altering (increasing/decreasing) Pr55Gag binding. Congruent estimates, in terms of labeling the position as significantly vs. not significantly altering Pr55Gag binding, are found on the diagonal. Incongruent estimates are found on the anti-diagonal.
Supplementary Figure 3 Relative Kd values for each Pr55Gag concentration respectively vs. values obtained for the pooled (all concentrations) dataset
Left panels: Significant median log2(Kdm(max)/Kdw) QUOTE values of pooled data compared to (a) low Pr55Gag:RNA ratio (20nM Pr55Gag). (b) Equimolar Pr55Gag:RNA ratio (200nM Pr55Gag). (c) High Pr55Gag:RNA ratio (2000nM Pr55Gag), showing the Pearson correlation between the quantitative Kd estimates and the corresponding P-value for assessing non-zero correlation. ‘Median’ denotes the median of the re-sampling distribution obtained after computational analysis (see eq.(6) in the Online Methods). Significance is determined using eqs.(7)-(8) in the Online Methods after multiple test correction. Filled circles indicate Kdm(max)/Kdw estimates that are significantly smaller- or greater than 1 using either dataset, whereas red unfilled circles indicate Kdm(max)/Kdw estimates that are significantly smaller/greater 1 in the pooled dataset, but not in the dataset using the individual Pr55Gag concentration. Blue unfilled circles indicate Kdm(max)/Kdw estimates that are not significantly smaller/greater 1 in the pooled dataset, but which are estimated to be significantly altered when using the dataset with the individual Pr55Gag concentration. The vertical- and horizontal dashed lines separate Kdm(max)/Kdw estimates that are significantly larger- or smaller than 1 using the respective datasets and thus allow a qualitative assessment of the respective relative Kd estimates between datasets. The indicated percentages are computed on the bases of all Kdm(max)/Kdw estimates that are significantly smaller- or greater than 1 using either dataset (filled circles). Right panels: Cross-tabulation of the estimated number of positions significantly altering (increasing/decreasing) Pr55Gag binding. Congruent estimates, in terms of labeling the position as significantly vs. not significantly altering Pr55Gag binding, are found on the diagonal. Incongruent estimates are found on the anti-diagonal.
Supplementary Figure 4 Binding of Pr55Gag to the core-binding domain by filter binding assay
(a) Schematic of RNA (b) Binding of RNA (NL 1-532) and RNA corresponding to Pr55Gag core binding domain (NL 227-377) analyzed by filter binding assay.
Supplementary Figure 5 Interaction between MS2 coat protein and MS2 stem loop in non-cognate RNA
(a) Mapping of the effect of mutations on relative binding affinity, depicted as Kdm(max)/Kdw QUOTE , to a structural representation of the HIV-1 genome including the MS2 stem loop, which was inserted between the TAR and polyA hairpins. (b) Median effect of mutations on relative binding affinity, Kdm(max)/Kdw, in the HIV-1 genome containing the MS2 stem loop (red) and the HIV-1 genome without the MS2 stem loop (blue; negative control). Weak unspecific binding of the MS2 coat protein to the polyA and SL1 hairpins was detected irrespective of the presence of MS2 stem loop. (c) Effect of specific mutations on log2(Kdm/Kdw), for positions 52-92. MS2 stem loop spans positions 61-79. Box and whisker plots show effect of each class of mutation on relative binding affinity expressed as log2(Kdm/Kdw) where black and white circle shows median, box shows quartiles (25% and 75%) and whiskers show extremes (excluding outliers). Mutation classes are colour coded: red mutated to A; green mutated to C; blue mutated to G; yellow mutated to U. (d) Zoom on MS2 stem loop structure showing mutation with maximum effect on relative Kd, Kdm(max)/Kdw. Colour scale shows red with decreased binding affinity, blue increased binding affinity.
Supplementary Figure 6 Single variation analysis of Pr55Gag core binding domain
Positions where certain classes of mutations have statistically different effects on Pr55Gag binding. Green circle: structure-affecting mutations significantly impair binding and structure-preserving mutations impair binding significantly less than other possibilities. Yellow circles: structure-affecting mutations significantly impair binding and structure-preserving mutations impairs binding significantly less than one of the other possible mutations. Blue circles: structure-modulating mutations significantly impair Pr55Gag binding and where structure-preserving mutations improve Pr55Gag binding. Grey Circles: other sites of interest. Box and whisker plots show effect of each class of mutation on relative binding affinity expressed as QUOTE where black and white circle shows median, box shows quartiles (25% and 75%) and whiskers show extremes (excluding outliers). Mutation classes are colour coded: red mutated to A; green mutated to C; blue mutated to G; yellow mutated to U. Statistical tests are listed in Online Methods , P-values are listed in Supplementary Data 3.
Supplementary Figure 7 Binding of Pr55Gag to mutant RNA
HIV-1 genomic RNA containing point mutations was tested by filter binding experiments. These mutations were selected to be representative of the MIME data, including positions predicted to be single stranded, double stranded, positions showing strong evidence of RNA structure, and several positions where MIME indicates that co-variation maintains Pr55Gag binding.
Supplementary Figure 8 Comparison of relative dissociation constants obtained from filter binding experiments vs. MIME
Comparison of relative dissociation constants (expressed as log2(Kdm/Kdw)) for binding of Pr55Gag to HIV-1 genomic RNA, containing single point mutations, obtained from the filter binding experiments displayed in Supplementary Figure 7 vs. MIME. The left panel (a) compares the relative dissociation constants quantitatively. The black circles have the median log2(Kdm/Kdw) from the filter binding experiments vs. MIME as x- and y coordinates. The grey vertical bars show the quartiles of the MIME prediction, whereas the horizontal grey bars show the quartiles of the log2(Kdm/Kdw) estimate obtained from the filter binding experiment. The line of unity is indicated by a diagonal red dashed line and the vertical- and horizontal dashed black lined separate filter binding and MIME estimates that increase or decrease binding respectively. The right panel (b) shows a cross-tabulation comparing the qualitative outcome of the two assays. E.g. the upper left entry shows the number of Kdm/Kdw values that were estimated to be significantly different from 1 by MIME, but not by the filter binding experiment, whereas the upper right corner shows the number of Kdm/Kdw values that were estimated to be significantly different from 1 by both assays.
Supplementary Figure 9 Mapping of effects of mutations on binding affinity to the structure of the HIV genomic RNA
Effect of mutations mapped to the structure of the HIV-1 genome proposed by Siegfried et al. Mutation with maximum effect on Kd expressed as log2(Kdm(max)/Kdw). Postitions in red significantly impair Pr55Gag binding when mutated. Positions in blue significantly improve Pr55Gag binding when mutated. Positions with no significant change are shown in grey.
Siegfried, N. A., Busan, S., Rice, G. M., Nelson, J. A. E. & Weeks, K. M. RNA motif discovery by SHAPE and mutational profiling (SHAPE-MaP). Nat. Methods (2014). doi:10.1038/nmeth.3029
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9, Supplementary Tables 1 and 2 and Supplementary Notes 1–4 (PDF 4821 kb)
Supplementary Data 1
Table containing complete dataset of relative Kd values for each genome position. (XLSX 39 kb)
Supplementary Data 2
Table containing complete dataset of relative Kd values for each class of mutation and each genome position. (XLSX 94 kb)
Supplementary Data 3
Table containing results of statistical test to assess whether different classes of mutation have comparable effects on Pr55Gag binding affinity. (XLSX 20 kb)
Supplementary Data 4
Table containing stems predicted to be important for Pr55Gag binding. (XLSX 10 kb)
Rights and permissions
About this article
Cite this article
Smyth, R., Despons, L., Huili, G. et al. Mutational interference mapping experiment (MIME) for studying RNA structure and function. Nat Methods 12, 866–872 (2015). https://doi.org/10.1038/nmeth.3490
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3490
This article is cited by
-
Nano-DMS-MaP allows isoform-specific RNA structure determination
Nature Methods (2023)
-
Short- and long-range interactions in the HIV-1 5′ UTR regulate genome dimerization and packaging
Nature Structural & Molecular Biology (2022)
-
HIV-1 integrase binding to genomic RNA 5′-UTR induces local structural changes in vitro and in virio
Retrovirology (2021)
-
RNA structure-wide discovery of functional interactions with multiplexed RNA motif library
Nature Communications (2020)
-
New windows into retroviral RNA structures
Retrovirology (2018)