Abstract
Many protein interactions are mediated by small linear motifs interacting specifically with defined families of globular domains. Quantifying the specificity of a motif requires measuring and comparing its binding affinities to all its putative target domains. To this end, we developed the high-throughput holdup assay, a chromatographic approach that can measure up to 1,000 domain-motif equilibrium binding affinities per day. After benchmarking the approach on 210 PDZ-peptide pairs with known affinities, we determined the affinities of two viral PDZ-binding motifs derived from human papillomavirus E6 oncoproteins for 209 PDZ domains covering 79% of the human 'PDZome'. We obtained sharply sequence-dependent binding profiles that quantitatively describe the PDZome recognition specificity of each motif. This approach, applicable to many categories of domain-ligand interactions, has wide potential for quantifying the specificities of interactomes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Nash, P.D. Why modules matter. FEBS Lett. 586, 2572–2574 (2012).
Diella, F. et al. Understanding eukaryotic linear motifs and their role in cell signaling and regulation. Front. Biosci. 13, 6580–6603 (2008).
Lievens, S., Eyckerman, S., Lemmens, I. & Tavernier, J. Large-scale protein interactome mapping: strategies and opportunities. Expert Rev. Proteomics 7, 679–690 (2010).
Tonikian, R. et al. Bayesian modeling of the yeast SH3 domain interactome predicts spatiotemporal dynamics of endocytosis proteins. PLoS Biol. 7, e1000218 (2009).
Jadwin, J.A., Ogiue-Ikeda, M. & Machida, K. The application of modular protein domains in proteomics. FEBS Lett. 586, 2586–2596 (2012).
Doyle, D.A. et al. Crystal structures of a complexed and peptide-free membrane protein-binding domain: molecular basis of peptide recognition by PDZ. Cell 85, 1067–1076 (1996).
Luck, K., Charbonnier, S. & Travé, G. The emerging contribution of sequence context to the specificity of protein interactions mediated by PDZ domains. FEBS Lett. 586, 2648–2661 (2012).
Nourry, C., Grant, S.G. & Borg, J.P. PDZ domain proteins: plug and play! Sci. STKE 2003, RE7 (2003).
Iden, S. & Collard, J.G. Crosstalk between small GTPases and polarity proteins in cell polarization. Nat. Rev. Mol. Cell Biol. 9, 846–859 (2008).
Javier, R.T. & Rice, A.P. Emerging theme: cellular PDZ proteins as common targets of pathogenic viruses. J. Virol. 85, 11544–11556 (2011).
Charbonnier, S., Zanier, K., Masson, M. & Travé, G. Capturing protein-protein complexes at equilibrium: the holdup comparative chromatographic retention assay. Protein Expr. Purif. 50, 89–101 (2006).
Luck, K. et al. Putting into practice domain-linear motif interaction predictions for exploration of protein networks. PLoS ONE 6, e25376 (2011).
Stiffler, M.A. et al. PDZ domain binding selectivity is optimized across the mouse proteome. Science 317, 364–369 (2007).
Kranjec, C. & Banks, L. A systematic analysis of human papillomavirus (HPV) E6 PDZ substrates identifies MAGI-1 as a major target of HPV type 16 (HPV-16) and HPV-18 whose loss accompanies disruption of tight junctions. J. Virol. 85, 1757–1764 (2011).
Zanier, K. et al. Solution structure analysis of the HPV16 E6 oncoprotein reveals a self-association mechanism required for E6-mediated degradation of p53. Structure 20, 604–617 (2012).
Atchley, W.R., Zhao, J., Fernandes, A.D. & Drüke, T. Solving the protein sequence metric problem. Proc. Natl. Acad. Sci. USA 102, 6395–6400 (2005).
Belotti, E. et al. The human PDZome: a gateway to PDZ mediated functions. Mol. Cell. Proteomics 12, 2587–2603 (2013).
Cassonnet, P. et al. Benchmarking a luciferase complementation assay for detecting protein complexes. Nat. Methods 8, 990–992 (2011).
Beroukhim, R. et al. The landscape of somatic copy-number alteration across human cancers. Nature 463, 899–905 (2010).
Zaric, J. et al. Identification of MAGI1 as a tumor-suppressor protein induced by cyclooxygenase-2 inhibitors in colorectal cancer cells. Oncogene 31, 48–59 (2012).
Lee, S.J. et al. MAGI-3 competes with NHERF-2 to negatively regulate LPA2 receptor signaling in colon cancer cells. Gastroenterology 140, 924–934 (2011).
Valiente, M. et al. Binding of PTEN to specific PDZ domains contributes to PTEN protein stability and phosphorylation by microtubule-associated serine/threonine kinases. J. Biol. Chem. 280, 28936–28943 (2005).
Kotelevets, L. et al. Implication of the MAGI-1b/PTEN signalosome in stabilization of adherens junctions and suppression of invasiveness. FASEB J. 19, 115–117 (2005).
Ellenbroek, S.I., Iden, S. & Collard, J.G. Cell polarity proteins and cancer. Semin. Cancer Biol. 22, 208–215 (2012).
Roberts, S., Delury, C. & Marsh, E. The PDZ protein discs-large (DLG): the 'Jekyll and Hyde' of the epithelial polarity proteins. FEBS J. 279, 3549–3558 (2012).
Ong, D.C. et al. LARG at chromosome 11q23 has functional characteristics of a tumor suppressor in human breast and colorectal cancer. Oncogene 28, 4189–4200 (2009).
Thomas, M. & Banks, L. PDZRN3/LNX3 is a novel target of human papillomavirus type 16 (HPV-16) and HPV-18 E6. J. Virol. 89, 1439–1444 (2015).
Hu, H. et al. A map of WW domain family interactions. Proteomics 4, 643–655 (2004).
Landgraf, C. et al. Protein interaction networks by proteome peptide scanning. PLoS Biol. 2, E14 (2004).
Kaushansky, A. et al. Quantifying protein-protein interactions in high throughput using protein domain microarrays. Nat. Protoc. 5, 773–790 (2010).
Babault, N. et al. Peptides targeting the PDZ domain of PTPN4 are efficient inducers of glioblastoma cell death. Structure 19, 1518–1524 (2011).
Perkins, J.R., Diboun, I., Dessailly, B.H., Lees, J.G. & Orengo, C. Transient protein-protein interactions: structural, functional, and network properties. Structure 18, 1233–1243 (2010).
Gerber, D., Maerkl, S.J. & Quake, S.R. An in vitro microfluidic approach to generating protein-interaction networks. Nat. Methods 6, 71–74 (2009).
Hause, R.J. Jr. et al. Comprehensive binary interaction mapping of SH2 domains via fluorescence polarization reveals novel functional diversification of ErbB receptors. PLoS ONE 7, e44471 (2012).
Velázquez Campoy, A. & Freire, E. ITC in the post-genomic era...? Priceless. Biophys. Chem. 115, 115–124 (2005).
Rich, R.L. & Myszka, D.G. Advances in surface plasmon resonance biosensor analysis. Curr. Opin. Biotechnol. 11, 54–61 (2000).
Weiser, A.A. et al. SPOT synthesis: reliability of array-based measurement of peptide binding affinity. Anal. Biochem. 342, 300–311 (2005).
Tropea, J.E., Cherry, S., Nallamsetty, S., Bignon, C. & Waugh, D.S. A generic method for the production of recombinant proteins in Escherichia coli using a dual hexahistidine-maltose-binding protein affinity tag. Methods Mol. Biol. 363, 1–19 (2007).
Nominé, Y. et al. A strategy for optimizing the monodispersity of fusion proteins: application to purification of recombinant HPV E6 oncoprotein. Protein Eng. 14, 297–305 (2001).
Vincentelli, R. et al. High-throughput protein expression screening and purification in Escherichia coli. Methods 55, 65–72 (2011).
Saez, N.J. & Vincentelli, R. High-throughput expression screening and purification of recombinant proteins in E. coli. Methods Mol. Biol. 1091, 33–53 (2014).
Fournane, S. et al. Surface plasmon resonance analysis of the binding of high-risk mucosal HPV E6 oncoproteins to the PDZ1 domain of the tight junction protein MAGI-1. J. Mol. Recognit. 24, 511–523 (2011).
Charbonnier, S. et al. The structural and dynamic response of MAGI-1 PDZ1 with noncanonical domain boundaries to the binding of human papillomavirus E6. J. Mol. Biol. 406, 745–763 (2011).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
Acknowledgements
This work was supported by institutional support from CNRS, Université de Strasbourg, Infrastructures Biologie Santé et Agronomie (Ibisa) and grants from the Association pour la Recherche contre le Cancer (ARC) (no. 3171), Agence Nationale de la Recherche (ANR-MIME-2007 EPI-HPV-3D), US National Institutes of Health (NIH grant R01CA134737), Ligue contre le Cancer and Alsace contre le Cancer. K.L. was supported by grants of Région Alsace and ARC. J. Poirson was supported by a grant of the Ligue Régionale contre le Cancer. J.-P.B. is supported in part by grants from La Ligue Contre le Cancer, Institut paoli-Calmettes, Cancéropôle PACA and SIRIC (INCa-DGOS-Inserm 6038). J.-P.B. is supported as a scholar of Institut Universitaire de France. We thank the CCSB (Harvard University) for kindly providing clones of the human ORFeome, and T. Rolland, M.A. Delsuc, D. Altschuh, B. Kieffer, A. Dejaegere and all members of the “Oncoproteins” team for helpful discussions and advice.
Author information
Authors and Affiliations
Contributions
G.T. conceived the project and raised funding for it. G.T., S.C. and R.V. supervised the project. R.V. and S.C. developed the robotized holdup assay. K.L. designed and performed all computational data treatment and bioinformatics analysis. R.V., M.B., J.A., F.I., J.T. and K.R. performed high-throughput protein cloning and expression, and high-throughput holdup assays. S.C., M.-L.S. and K.L. curated the data. J. Poirson, A.F., P.C., Y.J. and M.M. performed the GPCA experiments. J. Polanowska, J.R. and J.-P.B. provided the 'A' PDZome resource. Y.N. provided expertise in data treatment. G.T., K.L., S.C. and N.W. interpreted the data. K.L., S.C., J. Poirson and G.T. prepared the figures. K.L., S.C., R.V., J. Poirson and G.T. wrote the Online Methods. K.L., G.T. and J. Poirson prepared the supplementary information. G.T. wrote the main text.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Comparison of holdup assays performed with either purified PDZ domains or bacterial overexpression extracts.
Correlation plot of binding intensities measured for a set of 5 His-MBP-PDZ domain constructs against 42 biotinylated peptides. Binding intensities (BI) of holdup experiments performed with IMAC purified His-MBP-PDZ domains or bacterial overexpression extracts are plotted on the x-axis and y-axis, respectively. The plot shows that the correlation for high-affinity binders (BIs between 0.8 and 1.0) are very good and with decreasing BI the divergence from the ideal diagonal stays in the error range of +/- 0.2 units. A general tendency towards obtaining slightly higher BIs when performing holdup experiments with bacterial overexpression extracts can be observed. Both observations of the good correlation between purified and unpurified His-MBP-PDZ constructs and the apparent higher BIs for unpurified His-MBP- PDZ constructs suggest that for well-produced and well folded His-MBP-PDZ domains, no purification is required for performing high-throughput holdup assays. This increases both speed and accuracy of the measurement.
Supplementary Figure 2 Virtual gel representation of capillary electrophoresis measurements of the overexpressed PDZome.
(a) Electropherograms (represented as virtual gels) of overexpression lysates of the 223 PDZ domains produced in 24 deep-well plate cultures and used for the E6-PDZome holdup screen in the present publication. The data shown correspond to the biotin resin filtrates used as negative controls in the screen. (b) Lysates of PDZ domains that were also overexpressed in 24-well cultures but did not meet all requirements (sequence, data quality, and reproducibility) for being retained in the final interaction data. (c) PDZ domains that required overexpression in 50 ml cultures in flasks and subsequent purification for being detected. (d) List of PDZ domains that could not be produced even after changing the construct and/or the expression conditions.
Supplementary Figure 3 Detailed PDZome binding profiles of two independent negative controls.
For studying the internal variability of the holdup assay and for defining a threshold for significant experimental results we performed holdup experiments using beads either coated with a biotinylated peptide called “none” (Biotin-GSNSGNGNS), which is designed not to bind to any PDZ domain, or simply coated with biotin, the usual negative control for a standard holdup experiment. Upper panel: NONE peptide-PDZ domain binding intensities (BI) ranked by decreasing values using biotin as a negative control reference. Lower panel: Biotin-PDZ domain BIs ranked by decreasing values using biotinylated none peptide as a negative control reference. The error bars correspond to 2-3 repetitions for the A and B PDZ constructs and up to 9 repetitions for the standard C constructs (see Online Methods for a detailed description of the A, B and C His-MBP-PDZ constructs). Note that in both panels 100% of the BIs are below a value of 0.2 and 98% below 0.1.
Supplementary Figure 4 Detailed PDZome binding profiles of HPV16 E6 and HPV18 E6 PBMs.
Upper panel: PDZome interaction profile of HPV16 E6 PBM (Biotin-SSRTRRETQL). Binding affinities (BI) are ranked from highest to lowest. Middle panel: PDZome interaction profile of HPV18 E6 PBM (Biotin-RLQRRRETQV). PDZ domains are listed in the same order as for the HPV16 E6 PBM ranking in the upper panel. Lower panel: PDZome interaction profile against HPV18 E6 PBM. BIs are ranked from highest to lowest. The detection threshold of 0.2 BI units is drawn as a blue line. Dark violet bars indicate PDZ domains that were previously published to interact with the motif and light violet bars indicate the strongest binding PDZ domain from a previously published interacting PDZ domain containing protein. All BI values for these plots are listed in Supplemental Table 3.
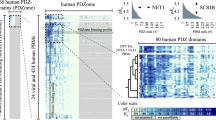
Supplementary Figure 5 Heat-map representation of the HPV16 and HPV18 E6 PBM – PDZome holdup screen.
PDZ domains are listed in alphabetical order. The left column shows experimental BIs for HPV16 E6 PBM and the right column for HPV18 E6 PBM. BIs ranging from 0 – 1 are color coded according to the color scale depicted on the lower right corner of the figure. Bold black with a green background labels PDZ domains published to bind HPV16 or HPV18. Bold black labels the PDZ domains that bound best to HPV16 or 18 E6 PBM from full-length PDZ domain containing proteins that had been published to bind to HPV16 or HPV18 E6. Black labels other PDZ domains from PDZ domain containing proteins that are known to bind to HPV16 or HPV18 E6. Grey color labels PDZ domains where no published information is available. This table shows that all the published interactions figure among the higher affinity binders and that many low affinity interactions were only detected using the holdup assay.
Supplementary Figure 6 Multiple sequence alignment of the 209 PDZ domains measured by holdup, ranked according to their strongest binding intensity to either HPV16 or HPV18 E6.
The alignment is presented in four panels in the following order: upper left, upper right, lower left, lower right. The source Fasta file for this alignment is provided in Supplementary Data 3.
Supplementary Figure 7 Extracting sequence-structure-activity correlations from holdup data: identifying PDZ domain positions that determine binding to E6.
Using a statistical approach (see Online Methods) we exploited the quantitative E6-binding data obtained on 209 PDZ domains to identify PDZ domain positions, where the occurrence of particular residues correlates with E6 binding (see Supplemental Table 5). The positions where the best correlations were found are represented in Fig. 6a on the structure of MAGI1-2/6. His119 and Val123 (numberings refer to PDZ alignment used for this analysis, see Supplemental Fig. 6) are among the residues that display highest correlation with E6 binding. In the 3D structure these two positions are facing residue Thr-2 of the peptide. Accordingly, in PDZ-binding motifs, the nature of the residue at position -2 is a well-known determining factor for PDZ selectivity.
(a) Plot showing the cumulative count of PDZ domains having a His at position 119. 107 out of 209 PDZs in our study have His119. The PDZ domains are ranked based on their highest holdup binding intensity obtained for 18E6 or 16E6, from the weakest (left) to the strongest binders (right). The experimental cumulative count curve is displayed as a fat line, colored in heat map mode according to the E6-binding intensities of the PDZ domains. For comparison, 100 randomly generated cumulative counts are shown, where the occurrence of His at position 119 has been randomly shuffled over the 209 PDZs. While all the random curves roughly approximate a straight line, the experimental curve displays a concave shape. This illustrates the bias for the preferential occurrence of His 119 in E6-binding PDZ domains.
(b) The concave shape of the experimental His 119 cumulative count curve can be numerically distinguished from the straight shape of the randomized curves by its lower AUC (Area Under Curve). The stronger the requirement of His119 for E6 binding, the more concave the curve, the lower the AUC. The diagram shows the distribution of 10,000 areas under the curve (AUC) obtained from randomized cumulative counts of 107 His residues over 209 PDZs. The AUC obtained for the cumulative count of His119 was 83. This AUC value is very unlikely to be found by chance, as illustrated by its statistical deviation from the AUC values of the randomized curves (p-value < 1.0E-4; z-score = -5.4).
Supplementary Figure 8 The 60 best E6-binding PDZ domains, ranked by ΔBI between HPV18 and HPV16 E6.
Positions whose physico-chemical properties correlate with preferences for HPV18 vs HPV16 (or vice versa) are indicated. The source Fasta file for this alignment is provided in Supplementary Data 3.
Supplementary Figure 9 Extracting sequence-structure-activity correlations from holdup data: identifying PDZ domain positions that determine preferential binding to either HPV16 or HPV18 E6.
Using a statistical approach (see Online Methods) we identified for the 60 E6-binding PDZ domains 13 domain positions whose physico-chemical properties significantly correlated with differences (ΔBI) in their holdup binding intensities for HPV18 E6 vs HPV16 E6 (see Supplemental Tables 6 and 7). To analyze the significance of these 13 correlated positions, we generated 100,000 random data sets in which the experimental ΔBI values were randomly re-attributed to the 60 PDZ domains. In each dataset, we counted the numbers of domain positions whose physico-chemical properties correlated with ΔBI values. The plot shows the distribution of numbers of correlated domain positions found in the 100,000 randomized datasets. At best, we observed 9 correlated positions, and this was only observed twice. By contrast, we identified 13 correlated positions in the real data. This number obtains a high z-score value (13.5) relative to the randomized data distribution. Therefore, these 13 correlated positions are very unlikely to be found by chance.
These positions showing the highest binding preference to HPV18 vs HPV16 E6 have been plotted on the 3D structure of PDZ DLG2-2/3 in Fig. 6b. These positions are found to cluster around the binding pocket that accommodates peptide position p0. Accordingly, p0 displays the most important sequence difference between the two E6 C-termini with respect to PDZ binding (L for HPV16; V for HPV18).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 (PDF 2231 kb)
Supplementary Table 1
Benchmarking holdup (HU) data vs. Biacore data previously obtained on 210 PDZ-peptide interactions (Luck et al., PlosOne 2011) (XLSX 57 kb)
Supplementary Table 2
Cloning results and sequences of the human PDZ domains used in this study (XLSX 102 kb)
Supplementary Table 3
Data of the E6-PDZome holdup screen in 384-well format (XLSX 49 kb)
Supplementary Table 4
Ranking of human PDZ domains according to E6-binding intensities (XLS 54 kb)
Supplementary Table 5
PDZ residue-position pairs preferred or disliked for E6 binding (XLSX 51 kb)
Supplementary Table 6
The 60 PDZ domains that bound at least one of the two E6 peptides with a BI >= 0.1 in the holdup assay (XLS 41 kb)
Supplementary Table 7
Search for correlations of ΔBI values with physico-chemical property indices for the 60 E6-binding PDZ domains revealed by holdup assay (XLS 69 kb)
Supplementary Table 8
Curated HPV E6 - PDZ interactions from the literature (XLSX 54 kb)
Supplementary Table 9
Comparison of holdup vs yeast two-hybrid results on PDZ-peptide interactions obtained with a 10 or 15-residue long peptide representing the HPV16 E6 C-terminus, respectively (XLS 44 kb)
Supplementary Table 10
GPCA validation results (XLSX 79 kb)
Supplementary Table 11
Search for PDZ-containing targets of E6 that are potentially involved in cancer (XLS 120 kb)
Supplementary Data 1
Exported Caliper files (source data for all holdup data) (ZIP 110976 kb)
Supplementary Data 2
Parameters and binding intensities (ZIP 593 kb)
Supplementary Data 3
2 alignments in FASTA format (ZIP 16 kb)
Rights and permissions
About this article
Cite this article
Vincentelli, R., Luck, K., Poirson, J. et al. Quantifying domain-ligand affinities and specificities by high-throughput holdup assay. Nat Methods 12, 787–793 (2015). https://doi.org/10.1038/nmeth.3438
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3438
This article is cited by
-
Large-scale phage-based screening reveals extensive pan-viral mimicry of host short linear motifs
Nature Communications (2023)
-
Quantitative fragmentomics allow affinity mapping of interactomes
Nature Communications (2022)
-
Promiscuity mapping of the S100 protein family using a high-throughput holdup assay
Scientific Reports (2022)
-
Molecular basis of the interaction of the human tyrosine phosphatase PTPN3 with the hepatitis B virus core protein
Scientific Reports (2021)
-
Role of PDZ-binding motif from West Nile virus NS5 protein on viral replication
Scientific Reports (2021)