Abstract
Assessing similarity between data sets with the reduced χ2 test requires the estimation of experimental errors, which, if incorrect, may render statistical comparisons invalid. We report a goodness-of-fit test, Correlation Map (CorMap), for assessing differences between one-dimensional spectra independently of explicit error estimates, using only data point correlations. Using small-angle X-ray scattering data, we demonstrate that CorMap maintains the power of the reduced χ2 test; moreover, CorMap is also applicable to other physical experiments.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
References
Bevington, P.R. & Robinson, K.D. in Data Reduction and Error Analysis for the Physical Sciences 3rd edn. 36–51 (McGraw-Hill, 2002).
Svergun, D.I., Koch, M.H.J., Timmins, P.A. & May, R.P. Small Angle X-Ray and Neutron Scattering from Solutions of Biological Macromolecules (Oxford Univ. Press, 2013).
Jacques, D.A., Gus, J.M., Svergun, D.I. & Trewhella, J. Acta Crystallogr. D Biol. Crystallogr. 68, 620–626 (2012).
Pearson, K. Philos. Mag. 50, 157–175 (1900).
Andrae, R., Schulze-Hartung, T. & Melchior, P. Preprint at http://arxiv.org/abs/1012.3754 (2010).
Schilling, M.F. Coll. Math. J. 21, 196–207 (1990).
Johnson, V.E. Proc. Natl. Acad. Sci. USA 110, 19313–19317 (2013).
Rambo, R.P. & Tainer, J.A. Nature 496, 477–481 (2013).
Trewhella, J. et al. Structure 21, 875–881 (2013).
Amato, A. et al. Phys. Rev. B Condens. Matter Mater. Phys. 89, 184425 (2014).
Petoukhov, M.V. et al. J. Appl. Crystallogr. 45, 342–350 (2012).
Franke, D. & Svergun, D.I. J. Appl. Crystallogr. 42, 342–346 (2009).
Varga, A. et al. FEBS Lett. 580, 2698–2706 (2006).
Round, A. et al. Acta Crystallogr. D Biol. Crystallogr. 71, 67–75 (2015).
Gasteiger, E. et al. in The Proteomics Protocols Handbook (ed. Walker, J.M.) 571–607 (Humana Press, 2005).
Franke, D., Kikhney, A.G. & Svergun, D.I. Nucl. Inst. Methods Phys. Res. A 689, 52–59 (2012).
Svergun, D., Barberato, C. & Koch, M.H.J. J. Appl. Crystallogr. 28, 768–773 (1995).
Jeffries, C.M., Graewert, M.A., Svergun, D.I. & Blanchet, C.E. J. Synchrotron Radiat. 22, 273–279 (2015).
Clopper, C.J. & Pearson, E.S. Biometrika 26, 404–413 (1934).
Acknowledgements
We thank E. Morenzoni of the Laboratory for Muon-Spin Spectroscopy, Paul Scherrer Institute, for providing the ZF-μSR data, taken at the GPS instrument of the Swiss Muon Source, Villigen, Switzerland. We thank R.P. Rambo for providing the original implementation of the χ2free test for our analysis and H. Mertens and J. Trewhella for many useful discussions. This work was supported by the Bundesministerium für Bildung und Forschung (BMBF) project BIOSCAT, grant 05K12YE1, and by the European Commission, BioStruct-X grant 283570.
Author information
Authors and Affiliations
Contributions
The initial idea was conceived of and simulation studies were done by D.F. Experimental data were collected by C.M.J. D.F, C.M.J. and D.I.S. participated in critical discussion and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Empirical and theoretical distributions of the reduced χ2 test.
The histogram shows the empirical distribution of 5,000 reduced χ2 values computed from 5,000 independent pair-wise comparisons of 10,000 SAXS data frames obtained from water (bars) together with their expected values from a reduced χ2 distribution (line) assuming no differences. The good agreement between the observed and expected distributions indicates accurate error estimates for this data set. Under the assumptions of correct errors and frame similarity, the acceptable range of values of χ2 is approximately 0.9 to 1.1, values less or greater are indicative of either differences in the data or miscalculated errors.
Supplementary Figure 2 The statistical properties of SAXS intensities recorded using a photon-counting detector.
(a) Histograms and limiting distributions of experimental intensities, Iexp(qk), of repeated measurements of water collected at a single value of q, in this case qk = 0.2012 nm−1 (wide histogram: 10,000 frames at 0.1s, medium histogram: 1,000 frames at 1.0s, narrow histogram: 500 frames at 10s). Generally, the distribution of intensities at any given qk is Gaussian and the respective standard deviations in this example decrease with √10 as expected by the Standard Error of the Mean. (b) Experimental error estimates of 10,000 frames of water according to Poisson counting statistics (dark gray) and the standard deviations of the Normals (light gray) across all of q. The spikes in the variations correspond to different numbers of pixels used to assess the errors caused by the gaps in the detector modules. (c) Example of a pair-wise joint normal distribution of two q-locations (qk, ql). (d) Correlation map of 10,000 frames of water, highlighting that data points are uncorrelated across the whole q-range.
Supplementary Figure 3 Application of CorMap to detect the onset of X-ray radiation damage to a protein sample during SAXS measurements.
Correlation map time series from experimental SAXS data frames of lysozyme (consecutive 50 ms exposures, 1 s total, n=1600 data points, unsubtracted data). The upper left panel shows an all-vs.-all frame comparison, indicating differences exist between the frames across the whole dataset. The top-left to bottom-right panels show the pair-wise correlation maps of the first frame relative to each subsequent frame together with Bonferroni adjusted p-values. Up to frame 13, the adjusted p-value is stable (1.00); frames 14-16 show a reduced p-value relative to frame 1 (0.0573-0.0143), while at frame 17 and later the adjusted p-value drops to < 0.01 indicating of statistically significant differences. The column to the right shows the overlay of 1D scattering profiles of selected frame pairs.
Supplementary Figure 4 Application of CorMap to detect concentration effects (repulsive interparticle interference).
(a) SAXS scattering patterns of RNAse collected at 3.7 mg/ml, 7.5 mg/ml and 15 mg/ml; (b)-(d) pair-wise correlation maps from the RNAse sample scattering at the respective concentrations do not reveal statistically significant differences across the profiles (n=1675, C=14, 14, 12, adjusted P-values: 0.1485, 0.1485 and 0.5525); (e) scattering patterns of human serum albumin at 5 mg/ml, 10 mg/ml and 20 mg/ml; (f)-(h) correlation maps of pair-wise comparisons of HSA at the three concentrations show concentration effects at low q and statistically significant differences between the SAXS data frames (n=1200, C=50, 162, 180, adjusted P <10e-6 in all cases).
Supplementary Figure 5 Empirical and theoretical distributions of the Correlation Map test.
Histogram of the edge lengths of maximum correlation patch sizes obtained from 5,000 independent experimental two-frame comparisons of water (bars), together with its expected distribution (dots). Here the number of available data points in the entire q-range, corresponds to coin tosses. In this figure, with n=1682 q-values, the expected largest edge length of the patches of similar correlation lies in the range of 8 to 20. Any larger lengths are extremely unlikely to occur by chance.
Supplementary Figure 6 Variation of the theoretical distribution with respect to its parameter n.
(a). The theoretical correlation map distributions calculated for n = 400, 800 and 1600 points. The maximum is located at log2(n). (b)-(d) Comparison of SAXS data sets comprised of 20 frames of water illustrating: (b) 1600 × 1600 data point comparison; (c) data re-binning of the same frames into 800 × 800 and (d) 400 × 400 data points. The white diagonal corresponds to each point's correlation to itself. The evaluation of differences using the correlation map takes into account the reduction in n, i.e., the expected edge length at a significance level α is dependent on the number of data points.
Supplementary Figure 7 Examples of experimental data and simulations thereof.
Overview of experimental data used to derive the empirical radiation damage components used in the simulations of (H3,H4,H5). The top row shows three frames each of the different experimental data sets (columns), the middle row depicts the extracted additive component for the simulation and the last row shows examples of the simulated data sets.
Supplementary Figure 8 Comparison of the statistical power of the CorMap and the reduced χ2 test.
Power comparison of the reduced χ2 test (dotted) and correlation map (line) at α = 0.01. The panels show the power for experimental frame comparisons where (a) represents systematic random shift errors, (b) systematic random scale errors, and (c)-(e) increasing contributions of modeled radiation damage. Effect sizes are in arbitrary units. True Positive proportions were estimated from 2,000 simulations each, the 99% Clopper-Pearson confidence intervals at each effect size are shown as vertical bars. Overlapping confidence intervals indicate equivalent tests at that effect size; fully separated intervals indicate significant differences between the tests. The corresponding count values are given in Supplementary Table 2.
Supplementary Figure 9 Models of bovine serum albumin used for statistical testing.
Backbone representations of the hypothetical BSA monomer modifications used to compare the False Positive rate and statistical power of the reduced χ2 test and correlation map for assessing SAXS data-model fits. The arrow from left-to-right indicates the rotation from native-to-rotated structure(s).
Supplementary Figure 10 Application of CorMap as a tool to assess data-model fits.
Panel (a) shows a simulated BSA SAXS profile with a native model fit (p-value: 0.1848) and corresponding correlation map in panel (c). Panel (b) shows the same data with a model that does not fit, (20° rotation in theTyr496 to Val497 bond angle p-value: <10e-6). The insert highlights the region of the misfit that is more clearly visible in a disturbance of the randomness pattern in the correlation map in panel (d). The corresponding reduced χ2 values with correct errors for these cases are 1.0 and 1.7 respectively. In many publications a reduced χ2 of 1.7 might be considered indicative of a good fit, while the correlation map shows this may not actually be the case (d). Panel (e) indicates the power of the reduced χ2 test (dotted line) and the correlation map (solid line) to correctly classify model fits. The effect size in this instance corresponds to an increasing rotation of around a bond angle of several BSA models (Supplementary Fig. 9). True Positive proportions were estimated from 10,000 simulations at each point, the 99% Clopper-Pearson confidence intervals at each effect size are shown as vertical. Overlapping confidence intervals indicate equivalent tests at that effect size; fully separated intervals indicate significant differences between the tests.
Supplementary Figure 11 The reduced χ2 and χ2free tests are equivalent if the errors are correctly specified.
Comparison of results of reduced χ2 and χ2free test to evaluate data-model fitting. A total of 23,000 simulated BSA datasets with correctly specified errors were analyzed using both tests to assess the fits of the models shown in Supplementary Figure 9; ‘without effect' (black) and with increasingly larger effect (gray scale). The results of χ2 and χ2free tests are, up to sampling variation inc2free, essentially identical, but do not correspond precisely to the diagonal (black line); the values ofc2free are systematically larger than those of χ2.
Supplementary Figure 12 The reduced χ2 and χ2free tests are equivalent if the errors are correctly specified, regardless of the actual error values.
(a) Example of a simulated SAXS dataset with 3% constant relative errors in black and the model scattering in white on top; (b) Comparison of reduced χ2 and χ2free tests of 1,000 repetitions of (a). The outcome is identical to what is shown in Supplementary Fig. 11.
Supplementary Figure 13 Comparison of reduced χ2 statistic and χ2free with incorrectly specified errors.
A total of 23,000 BSA model datasets were analyzed as described in the main text, the only difference being the assignment of incorrect errors prior to analysis. Panel (a) correct error structure, but half the magnitude, (b) correct error structure but twice the magnitude, (c) a random permutation of the correct errors and (d) a constant 75% relative error across the data set. The circle shown in each panel indicates the location of the correct results shown in Supplementary Fig. 11.
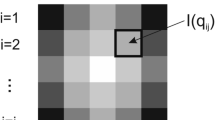
Supplementary Figure 14 Radial averaging of an idealized SAXS image.
Only pixels with a constant distance (black) from the beam center (red) are considered for each data point. Anti-aliasing must not be employed.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14 and Supplementary Tables 1 and 2 (PDF 3704 kb)
Goodness-of-fit of bead model refinement.
Dummy atom bead model refinement against lysozyme SAXS data. The left panel displays the progressive improvement of the fit (solid line) for the step-wise DAMMIF bead model refinement of the shape of lysozyme against lysozyme SAXS data (dots). As the fit improves, the correlation matrix (right panel) goes from having large contiguous areas of +1 or -1 correlations (i.e., large patches) to a randomized lattice pattern. The initial and finally-refined lysozyme models are shown in Figure 2 of the main text. (MPG 6868 kb)
Rights and permissions
About this article
Cite this article
Franke, D., Jeffries, C. & Svergun, D. Correlation Map, a goodness-of-fit test for one-dimensional X-ray scattering spectra. Nat Methods 12, 419–422 (2015). https://doi.org/10.1038/nmeth.3358
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3358
This article is cited by
-
Unconventional structure and mechanisms for membrane interaction and translocation of the NF-κB-targeting toxin AIP56
Nature Communications (2023)
-
Dynamics and structural changes of calmodulin upon interaction with the antagonist calmidazolium
BMC Biology (2022)
-
Production and characterisation of modularly deuterated UBE2D1–Ub conjugate by small angle neutron and X-ray scattering
European Biophysics Journal (2022)
-
Small-angle X-ray and neutron scattering
Nature Reviews Methods Primers (2021)
-
Estimation of the molecular weight of nanoparticles using a single small-angle X-ray scattering measurement on a relative scale
Scientific Reports (2021)