Abstract
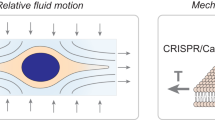
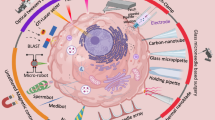
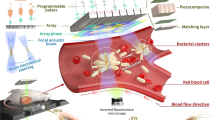
We report a high-throughput platform for delivering large cargo elements into 100,000 cells in 1 min. Our biophotonic laser-assisted surgery tool (BLAST) generates an array of microcavitation bubbles that explode in response to laser pulsing, forming pores in adjacent cell membranes through which cargo is gently driven by pressurized flow. The platform delivers large items including bacteria, enzymes, antibodies and nanoparticles into diverse cell types with high efficiency and cell viability. We used this platform to explore the intracellular lifestyle of Francisella novicida and discovered that the iglC gene is unexpectedly required for intracellular replication even after phagosome escape into the cell cytosol.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Rusk, N. Seamless delivery. Nat. Methods 8, 44 (2011).
Naldini, L. et al. In vivo gene delivery and stable transduction of nondividing cells by a lentiviral vector. Science 272, 263–267 (1996).
Akin, D. et al. Bacteria-mediated delivery of nanoparticles and cargo into cells. Nat. Nanotechnol. 2, 441–449 (2007).
Felgner, P.L. et al. Lipofection: a highly efficient, lipid-mediated DNA-transfection procedure. Proc. Natl. Acad. Sci. USA 84, 7413–7417 (1987).
De Smedt, S.C., Demeester, J. & Hennink, W.E. Cationic polymer based gene delivery systems. Pharm. Res. 17, 113–126 (2000).
Somiari, S. et al. Theory and in vivo application of electroporative gene delivery. Mol. Ther. 2, 178–187 (2000).
Guignet, E.G. & Meyer, T. Suspended-drop electroporation for high-throughput delivery of biomolecules into cells. Nat. Methods 5, 393–395 (2008).
Boukany, P.E. et al. Nanochannel electroporation delivers precise amounts of biomolecules into living cells. Nat. Nanotechnol. 6, 747–754 (2011).
Kim, H.J., Greenleaf, J.F., Kinnick, R.R., Bronk, J.T. & Bolander, M.E. Ultrasound-mediated transfection of mammalian cells. Hum. Gene Ther. 7, 1339–1346 (1996).
Mitragotri, S. Healing sound: the use of ultrasound in drug delivery and other therapeutic applications. Nat. Rev. Drug Discov. 4, 255–260 (2005).
Tirlapur, U.K. & König, K. Cell biology: targeted transfection by femtosecond laser. Nature 418, 290–291 (2002).
Tao, W., Wilkinson, J., Stanbridge, E.J. & Berns, M.W. Direct gene transfer into human cultured cells facilitated by laser micropuncture of the cell membrane. Proc. Natl. Acad. Sci. USA 84, 4180–4184 (1987).
Chakravarty, P., Qian, W., El-Sayed, M.A. & Prausnitz, M.R. Delivery of molecules into cells using carbon nanoparticles activated by femtos laser pulses. Nat. Nanotechnol. 5, 607–611 (2010).
Sharei, A. et al. A vector-free microfluidic platform for intracellular delivery. Proc. Natl. Acad. Sci. USA 110, 2082–2087 (2013).
Shalek, A.K. et al. Vertical silicon nanowires as a universal platform for delivering biomolecules into living cells. Proc. Natl. Acad. Sci. USA 107, 1870–1875 (2010).
Capecchi, M.R. High efficiency transformation by direct microinjection of DNA into cultured mammalian cells. Cell 22, 479–488 (1980).
Zhang, Y. & Yu, L.-C. Microinjection as a tool of mechanical delivery. Curr. Opin. Biotechnol. 19, 506–510 (2008).
Hurtig, J. & Orwar, O. Injection and transport of bacteria in nanotube-vesicle networks. Soft Matter 4, 1515–1520 (2008).
Wu, T.-H. et al. Photothermal nanoblade for large cargo delivery into mammalian cells. Anal. Chem. 83, 1321–1327 (2011).
Hartland, G.V. Optical studies of dynamics in noble metal nanostructures. Chem. Rev. 111, 3858–3887 (2011).
Link, S. & El-Sayed, M.A. Spectral properties and relaxation dynamics of surface plasmon electronic oscillations in gold and silver nanodots and nanorods. J. Phys. Chem. B 103, 8410–8426 (1999).
Kotaidis, V., Dahmen, C., von Plessen, G., Springer, F. & Plech, A. Excitation of nanoscale vapor bubbles at the surface of gold nanoparticles in water. J. Chem. Phys. 124, 184702 (2006).
Lukianova-Hleb, E. et al. Plasmonic nanobubbles as transient vapor nanobubbles generated around plasmonic nanoparticles. ACS Nano 4, 2109–2123 (2010).
Furlani, E.P., Karampelas, I.H. & Xie, Q. Analysis of pulsed laser plasmon-assisted photothermal heating and bubble generation at the nanoscale. Lab Chip 12, 3707–3719 (2012).
Yamane, D. et al. Electrical impedance monitoring of photothermal porated mammalian cells. J. Lab. Autom. 19, 50–59 (2014).
Marquis, H., Doshi, V. & Portnoy, D.A. The broad-range phospholipase C and a metalloprotease mediate listeriolysin O-independent escape of Listeria monocytogenes from a primary vacuole in human epithelial cells. Infect. Immun. 63, 4531–4534 (1995).
Clemens, D.L., Lee, B.-Y. & Horwitz, M.A. Virulent and avirulent strains of Francisella tularensis prevent acidification and maturation of their phagosomes and escape into the cytoplasm in human macrophages. Infect. Immun. 72, 3204–3217 (2004).
Nano, F.E. & Schmerk, C. The Francisella pathogenicity island. Ann. NY Acad. Sci. 1105, 122–137 (2007).
Barker, J.R. et al. The Francisella tularensis pathogenicity island encodes a secretion system that is required for phagosome escape and virulence. Mol. Microbiol. 74, 1459–1470 (2009).
Nano, F.E. et al. A Francisella tularensis pathogenicity island required for intramacrophage growth. J. Bacteriol. 186, 6430–6436 (2004).
de Bruin, O.M. et al. The biochemical properties of the Francisella pathogenicity island (FPI)-encoded proteins IglA, IglB, IglC, PdpB and DotU suggest roles in type VI secretion. Microbiology 157, 3483–3491 (2011).
Checroun, C., Wehrly, T.D., Fischer, E.R., Hayes, S.F. & Celli, J. Autophagy-mediated reentry of Francisella tularensis into the endocytic compartment after cytoplasmic replication. Proc. Natl. Acad. Sci. USA 103, 14578–14583 (2006).
Golovliov, I., Sjöstedt, A., Mokrievich, A. & Pavlov, V. A method for allelic replacement in Francisella tularensis. FEMS Microbiol. Lett. 222, 273–280 (2003).
Jones, J.W. et al. Absent in melanoma 2 is required for innate immune recognition of Francisella tularensis. Proc. Natl. Acad. Sci. USA 107, 9771–9776 (2010).
Acknowledgements
This work was supported by a University of California Discovery Biotechnology Award (178517), US National Institutes of Health grants AI065359, GM114188 and EB014456, and by NanoCav, LLC. This work was also funded by US National Science Foundation grant CBET-1404080. The authors thank K. Niazi and S. Rabizadeh for helpful discussions and support, D.A. Portnoy (University of California, Berkeley) for providing the L. monocytogenes strain DP-L2318, D.M. Monack (Stanford University) for the F. novicida ΔFPI strain, K.E. Klose (University of Texas at San Antonio) for the F. novicida U112 strain, F.E. Nano (University of Victoria) for the codon-optimized superfolder gfp and J.S. Hong (University of California, Los Angeles) for the stress gene assay.
Author information
Authors and Affiliations
Contributions
Y.-C.W., T.-H.W. and P.-Y.C. had the idea for the platform. Y.-C.W. designed and fabricated the BLAST device. T.-H.W. built the experimental setup. Y.-C.W., T.-H.W., D.L.C. and B.-Y.L. performed the experiments and analyzed the data. B.-Y.L. prepared mutant and complemented Francisella strains. X.W. ran the numerical simulations. P.-Y.C., M.A.T. and M.A.H. advised on experiments, data analysis and paper writing. All authors discussed the experimental results and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Fabrication of the BLAST platform.
The silicon-based device was fabricated on 2” double-sided polished silicon wafers of 300 μm thickness as follows: 1) A 1.5 μm thick thermal oxide layer was grown on both sides of a wafer. 2) After surface treatment with hexamethyldisilazane, photoresist AZ4620 (AZ Electronic Materials) was spin coated on one side of the wafer and exposed to UV light through a chrome mask with the design of fluid channels. The patterns were developed in dilute AZ400K (AZ Electronic Materials) solution for 5 min. 3) After 100°C baking for 10 min, the wafer was etched by advanced oxide etching (Surface Technology Systems) to remove the oxide layer and by deep reactive ion etching (Unaxis USA) to etch through the silicon substrate. 4) Another photolithography process was repeated on the opposite side of the wafer to pattern a 3 μm hole array and aligned to the channel structures. 5) Advanced oxide etching was used again to etch through holes on the oxide thin film. There was still a residue photoresist layer on the top surface. 6, 7) A 100 nm thick titanium thin film was deposited at a tilted angle by an e-beam evaporator (CHA Industries). 8) Finally, the device was immersed in acetone to detach the titanium thin film on the photoresist such that only the sidewalls of the small holes retained the metal coating.
Supplementary Figure 2 Time-resolved images showing the dynamics of rapidly expanding bubbles triggered under different laser fluence and polarization (bubbles are the darker area of these images).
Cavitation bubbles are initiated at the two tips of the tapered titanium film due to the lighting-rod enhancement effect near sharp metal tips. This is a non-resonant broadband effect and is also insensitive to light polarization on our platform. The bubble size is dependent on the laser fluence. Higher pulse energy generates larger sized cavitation bubbles with longer lifetimes. The lifetime of bubbles triggered by nanosecond laser pulses typically lasts for a few hundred nanoseconds. Scale bar, 3 μm.
Supplementary Figure 3 40-kDa FITC-dextran delivered into HeLa and three types of primary human cells (3 d after delivery).
(a-d) Bright-field microscopy images of (a) HeLa cells, (b) normal human dermal fibroblasts (NHDFs), (c) peripheral blood monocyte derived macrophages (PB-MDMs), and (d) renal proximal tubule epithelial cells (RPTECs). Scale bar, 50 μm. (e-h) 40kDa FITC-dextran (green) was delivered into (e) HeLa cells, (f) NHDFs, (g) PB-MDMs, and (h) RPTECs and incubated on chips for 3 d after delivery. Cells were stained by CellTracker™ Blue CMAC (blue, top row); propidium iodide (red) was used to identify the dead cells (bottom row). Scale bar, 100 μm.
Supplementary Figure 4 Additional large cargo successfully delivered into cells with BLAST.
(a) 2 μm green fluorescent polystyrene beads were delivered into HeLa cells. Cell membranes were stained with WGA 350 (blue), and propidium iodide (red) was used to identify dead cells. Scale bar, 50 μm. (b) 200 nm magnetic beads (red) were delivered into HeLa cells and attracted by a nearby micromagnet. Scale bars, 50 μm. (c) Mouse anti-α-tubulin monoclonal antibody with Alexa 488 conjugate was delivered into NHDFs, labeling the cytoskeleton (green). After 5 h incubation, cells were fixed and permeabilized to remove background fluorescence. Scale bar, 30 μm. (d) Delivery efficiency of 5 different sizes of polystyrene beads (20 nm, 200 nm, 500 nm, 1 μm, 2 μm) and cell viability. The data shows that the cell viability is similar and above 90% with delivery of all bead sizes. The delivery efficiency decreases with size from 93% for 20 nm beads to 62% for 2 µm beads. Error bars, s.d. (n = 4). (e) Co-delivery of 100 nm and 500 nm beads at a 6:1 ratio (180 beads/pl for 100 nm beads and 30 beads/pl for 500 nm beads). This ratio remains close to 6 after the beads were delivered into cells [92.4 ± 36.2 beads/cell for 100 nm beads (red) and 16.3 ± 7.4 beads/cell for 500 nm beads (green)], which proves that BLAST can deliver multiple payloads and the delivery process is not strongly size and diffusion speed dependent. Scale bar, 50 μm. (f) A control experiment shows that with fluid pumping but no laser pulsing, few 100 nm and 500 nm beads enter cells. Scale bar, 50 μm. (p < 0.0001 for beads/cell in e vs. f)
Supplementary Figure 5 Evaluation of stress levels of HeLa cells after BLAST delivery.
The HSPA6 gene expression level indicates the cellular response to heat shock. Gene expression levels were normalized to a GAPDH housekeeping gene for all cell samples including (A) laser pulsing with fluid pumping, (B) laser pulsing only, (C) fluid pumping only and (D) control without laser pulsing and fluid pumping. BLAST delivered cells (A-C) showed expression levels that were statistically similar to the cells without laser and flow applied (D). A positive control was provided by incubating HeLa cells in culture media at 42°C for 1 h. Error bars, s.d. (n = 3). Primers for gene expression were generated using Roche’s Universal Probe Library Assay Design Center online.
HSPA6 Fwd: (TCATGAAGCCGAGCAGTACA) Rev: (GTTTTTGGCAGCCACTCTGT).
GAPDH Fwd: (GCTCTCTGCTCCTCCTGTTC) Rev: (ACGACCAAATCCGTTGACTC).
Supplementary Figure 6 Low-magnification data showing the uniformity of enzyme delivery.
β-lactamase at 50 units/ml (a,c) and 1 unit/ml (b,d) in PBS was delivered into NHDFs using BLAST under the condition of fluid pumping with (a,b) or without (c,d) laser pulsing. The β-lactamase FRET substrate, CCF4 is added to the cells and under UV excitation - fluoresces green in the absence of cytosolic β-lactamase (c,d). β-lactamase delivered to the cells by laser pulsing cleaves the CCF4, causing a loss of FRET and a shift in fluorescence from green to blue (a,b). Scale bar, 100 μm.
Supplementary Figure 7 GFP-expressing parental L. monocytogenes, but not the Δhly ΔplcB mutant strain, recruits actin and forms actin tails after phagocytic uptake by THP-1 cells.
THP-1 cells were incubated with GFP-expressing parental or mutant strains L. monocytogenes at an MOI of 4:1 for 30 min in DMEM with 10% FBS, washed, and incubated in culture medium containing 5 μg/ml gentamicin to kill extracellular bacteria. After 6 h, the monolayers were fixed with 4% paraformaldehyde in PBS and actin filaments were stained with phalloidin-AlexaFluor 549 and nuclei were stained with DAPI. GFP-fluorescent wild-type (a) and mutant (d) bacteria fluoresce green (shown in gray scale). Actin filaments stained by phalloidin-AlexaFluor 549 fluoresce red (shown in gray scale) are recruited to the parental L. monocytogenes (b) and frequently form comet tails (arrowheads), but no recruitment or comet tails are seen with the escape incompetent mutant strain (e). In marked contrast, as shown in Fig. 5a, this escape incompetent mutant strain recruits actin and forms abundant comet tails after direct cytosolic delivery by BLAST. Merged color images are shown in panels (c) and (f). Scale bars, 10 μm.
Supplementary Figure 8 Characterization of F. novicida mutants: growth in HeLa cells and macrophages and IglC expression.
(a,b) Francisella Pathogenicity Island is required for cytosolic replication. Macrophages are the natural primary host cells for Francisella, and it is well established that Francisella is engulfed by macrophages into membrane-bound phagocytic vacuoles from which the bacterium must escape into the cytosol to replicate intracellularly. As a control, experiments in which THP-1 cells were infected with Francisella were carried out in parallel with BLAST experiments in which HeLa cells were infected with Francisella. F. novicida wild-type or F. novicida mutant with deletion of the entire Francisella Pathogenicity Island (FPI) were delivered into HeLa cells by BLAST (a) or ingested by phorbol ester-differentiated human macrophage-like THP-1 cells (b), as described. At 1, 6, and 24 h post-infection, the monolayers were lysed with 1% saponin in PBS and plated on chocolate agar for enumeration of bacterial colony forming units (CFU). While the wild-type F. novicida multiply in both HeLa and THP-1cells, the ΔFPI strain is not able to multiply either in macrophages, in which it is unable to escape from the phagosome, or in HeLa cells, even when delivered directly into the cytosol. Error bars, s.d. (n = 3). (c) Complementation restores the ability of F. novicida ΔiglC mutant to multiply intracellularly. Human monocytic THP-1 cells (1×105/well) were differentiated with phorbol ester and infected with F. novicida expressing sfGFP (1×106/well) for 1.5, 6, or 24 h, as indicated. Three F. novicida strains were studied: F. novicida wild-type, F. novicida ΔiglC, and F. novicida ΔiglC complemented with iglC on a plasmid. The infected monolayers were then fixed with 4% formaldehyde, and the nuclei stained with DAPI. High resolution images of the infected monolayers were acquired by ImageXpress (Molecular Probes) and analyzed by MetaXpress High Content Image Analysis Software. From 1.5 to 24 h, the number of green fluorescent bacteria per nucleus increased about 10-fold for the F. novicida wild-type and complemented F. novicida ΔiglC strains; in contrast, the number of F. novicida ΔiglC per nucleus decreased by 3-fold. Error bars, s.d. (n = 8, 3, and 8 wells for all strains at 1.5, 6, and 24 h, respectively). (d) Immunoblot analysis confirms the lack of IglC expression in the F. novicida mutant strains. Bacterial lysates from 1.5×107 GFP expressing F. novicida wild type (lane 1), ΔFPI (lane 2), ΔiglC (lane 3), and ΔiglC complemented with iglC on a plasmid (lane 4) strains were separated by SDS-polyacrylamide gel electrophoresis and transferred to nitrocellulose membrane for probing with monoclonal antibody to IglB (1:1,000; BEI Resources) or polyclonal antibody to IglC (1:1,000), bacterioferritin (Bfr, 1:1,000), or GFP (1:2,000; Assay Designs), as indicated to the right of the figure. IglC expression is absent from F. novicida ΔFPI and ΔiglC strains (lanes 2 and 3) and restored in the iglC complemented strain (lane 4).
Supplementary Figure 9 Differential digitonin permeabilization assay for F. novicida localization in HeLa cells.
GFP-expressing F. novicida wild-type strain (a-e), ΔiglC-mutant strain (f-j), or iglC-complemented strain (k-o) were delivered by BLAST into HeLa cells and the cells were fixed and stained by the differential digitonin method at 30 min (a-e), or 1 d (f-o) after laser assisted delivery. Cell membranes were stained with WGA-AlexaFluor 594 (near infrared, shown in gray scale in b,g,l). HeLa cells were permeabilized with digitonin (which permeabilizes plasma membrane but not intracellular vesicles) and antibody accessible F. novicida (i.e. those not enclosed within vacuoles) were stained with a chicken anti-F. novicida antibody followed by a goat anti-chicken Texas Red-X conjugated secondary antibody (red, d,i,n). GFP-expressing F. novicida fluoresce green (c,h,m) and HeLa cell nuclei are stained blue with DAPI (a,f,k). Merged images are shown in e, j and o. At 30 min after BLAST (a-e), bacteria stain both green and red, indicating cytosolic localization. By 1 d after BLAST, most of the ΔiglC mutant strain have been repackaged into vacuoles (f-j) and fluoresce green but not red (arrowheads), with only occasional bacteria fluorescing both green and red (arrow). In contrast, by 1 d after BLAST delivery, the iglC-complemented strain (k-o) has multiplied extensively within the HeLa cell and is stained by the red fluorescent antibody, confirming cytosolic localization in the HeLa cells. Scale bars, 20 μm.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9, Supplementary Tables 1 and 2 and Supplementary Note (PDF 2099 kb)
Rights and permissions
About this article
Cite this article
Wu, YC., Wu, TH., Clemens, D. et al. Massively parallel delivery of large cargo into mammalian cells with light pulses. Nat Methods 12, 439–444 (2015). https://doi.org/10.1038/nmeth.3357
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3357
This article is cited by
-
Electroactive nanoinjection platform for intracellular delivery and gene silencing
Journal of Nanobiotechnology (2023)
-
The cellular response to plasma membrane disruption for nanomaterial delivery
Nano Convergence (2022)
-
Role of actin cytoskeleton in cargo delivery mediated by vertically aligned silicon nanotubes
Journal of Nanobiotechnology (2022)
-
Advanced tools and methods for single-cell surgery
Microsystems & Nanoengineering (2022)
-
Computational modelling of membrane gating in capsule translocation through microchannel with variable section
Microfluidics and Nanofluidics (2021)