Abstract
RNA-protein complexes play pivotal roles in many central biological processes. Although methods based on high-throughput sequencing have advanced our ability to identify the specific RNAs bound by a particular protein, there is a need for precise and systematic ways to identify RNA interaction sites on proteins. We have developed an experimental and computational workflow combining photo-induced cross-linking, high-resolution mass spectrometry and automated analysis of the resulting mass spectra for the identification of cross-linked peptides, cross-linking sites and the cross-linked RNA oligonucleotide moieties of such RNA-binding proteins. The workflow can be applied to any RNA-protein complex of interest or to whole proteomes. We applied the approach to human and yeast mRNA-protein complexes in vitro and in vivo, demonstrating its powerful utility by identifying 257 cross-linking sites on 124 distinct RNA-binding proteins. The open-source software pipeline developed for this purpose, RNPxl, is available as part of the OpenMS project.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Glisovic, T., Bachorik, J.L., Yong, J. & Dreyfuss, G. RNA-binding proteins and post-transcriptional gene regulation. FEBS Lett. 582, 1977–1986 (2008).
Matera, A.G., Terns, R.M. & Terns, M.P. Non-coding RNAs: lessons from the small nuclear and small nucleolar RNAs. Nat. Rev. Mol. Cell Biol. 8, 209–220 (2007).
Yates, L.A., Norbury, C.J. & Gilbert, R.J. The long and short of microRNA. Cell 153, 516–519 (2013).
van der Feltz, C., Anthony, K., Brilot, A. & Pomeranz Krummel, D.A. Architecture of the spliceosome. Biochemistry 51, 3321–3333 (2012).
Sabin, L.R., Delás, M.J. & Hannon, G.J. Dogma derailed: the many influences of RNA on the genome. Mol. Cell 49, 783–794 (2013).
Mercer, T.R. & Mattick, J.S. Structure and function of long noncoding RNAs in epigenetic regulation. Nat. Struct. Mol. Biol. 20, 300–307 (2013).
Castello, A. et al. Insights into RNA biology from an atlas of mammalian mRNA-binding proteins. Cell 149, 1393–1406 (2012).
Baltz, A.G. et al. The mRNA-bound proteome and its global occupancy profile on protein-coding transcripts. Mol. Cell 46, 674–690 (2012).
Mitchell, S.F., Jain, S., She, M. & Parker, R. Global analysis of yeast mRNPs. Nat. Struct. Mol. Biol. 20, 127–133 (2013).
Klass, D.M. et al. Quantitative proteomic analysis reveals concurrent RNA-protein interactions and identifies new RNA-binding proteins in Saccharomyces cerevisiae. Genome Res. 23, 1028–1038 (2013).
Hafner, M. et al. Transcriptome-wide identification of RNA-binding protein and microRNA target sites by PAR-CLIP. Cell 141, 129–141 (2010).
Kohlbacher, O. et al. TOPP—the OpenMS proteomics pipeline. Bioinformatics 23, e191–e197 (2007).
Sturm, M. et al. OpenMS—an open-source software framework for mass spectrometry. BMC Bioinformatics 9, 163 (2008).
Geer, L.Y. et al. Open mass spectrometry search algorithm. J. Proteome Res. 3, 958–964 (2004).
Kramer, K. et al. Mass-spectrometric analysis of proteins cross-linked to 4-thio-uracil- and 5-bromo-uracil-substituted RNA. Int. J. Mass Spectrom. 304, 184–194 (2011).
Perkins, D.N., Pappin, D.J., Creasy, D.M. & Cottrell, J.S. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20, 3551–3567 (1999).
Pourshahian, S. & Limbach, P.A. Application of fractional mass for the identification of peptide-oligonucleotide cross-links by mass spectrometry. J. Mass Spectrom. 43, 1081–1088 (2008).
UniProt Consortium. Reorganizing the protein space at the Universal Protein Resource (UniProt). Nucleic Acids Res. 40, D71–D75 (2012).
Hentze, M.W. Enzymes as RNA-binding proteins: a role for (di)nucleotide-binding domains? Trends Biochem. Sci. 19, 101–103 (1994).
Mackereth, C.D. et al. Multi-domain conformational selection underlies pre-mRNA splicing regulation by U2AF. Nature 475, 408–411 (2011).
Ben-Shem, A. et al. The structure of the eukaryotic ribosome at 3.0 Å resolution. Science 334, 1524–1529 (2011).
Zhu, D., Stumpf, C.R., Krahn, J.M., Wickens, M. & Hall, T.M. A 5′ cytosine binding pocket in Puf3p specifies regulation of mitochondrial mRNAs. Proc. Natl. Acad. Sci. USA 106, 20192–20197 (2009).
Urlaub, H., Thiede, B., Müller, E.C., Brimacombe, R. & Wittmann-Liebold, B. Identification and sequence analysis of contact sites between ribosomal proteins and rRNA in Escherichia coli 30 S subunits by a new approach using matrix-assisted laser desorption/ionization-mass spectrometry combined with N-terminal microsequencing. J. Biol. Chem. 272, 14547–14555 (1997).
Kühn-Hölsken, E., Dybkov, O., Sander, B., Lührmann, R. & Urlaub, H. Improved identification of enriched peptide RNA cross-links from ribonucleoprotein particles (RNPs) by mass spectrometry. Nucleic Acids Res. 35, e95 (2007).
Luo, X. et al. Structural and functional analysis of the E. coli NusB-S10 transcription antitermination complex. Mol. Cell 32, 791–802 (2008).
Kühn-Hölsken, E. et al. Mapping the binding site of snurportin 1 on native U1 snRNP by cross-linking and mass spectrometry. Nucleic Acids Res. 38, 5581–5593 (2010).
Mozaffari-Jovin, S. et al. The Prp8 RNase H-like domain inhibits Brr2-mediated U4/U6 snRNA unwinding by blocking Brr2 loading onto the U4 snRNA. Genes Dev. 26, 2422–2434 (2012).
Ghalei, H., Hsiao, H.H., Urlaub, H., Wahl, M.C. & Watkins, N.J. A novel Nop5-sRNA interaction that is required for efficient archaeal box C/D sRNP formation. RNA 16, 2341–2348 (2010).
Müller, M. et al. A cytoplasmic complex mediates specific mRNA recognition and localization in yeast. PLoS Biol. 9, e1000611 (2011).
Schmidt, C., Kramer, K. & Urlaub, H. Investigation of protein-RNA interactions by mass spectrometry—techniques and applications. J. Proteomics 75, 3478–3494 (2012).
Allain, F.H. et al. Solution structure of the HMG protein NHP6A and its interaction with DNA reveals the structural determinants for non-sequence-specific binding. EMBO J. 18, 2563–2579 (1999).
Werner, E., Wende, W., Pingoud, A. & Heinemann, U. High resolution crystal structure of domain I of the Saccharomyces cerevisiae homing endonuclease PI-SceI. Nucleic Acids Res. 30, 3962–3971 (2002).
Leidig, C. et al. Structural characterization of a eukaryotic chaperone—the ribosome-associated complex. Nat. Struct. Mol. Biol. 20, 23–28 (2013).
Schmitzová, J. et al. Crystal structure of Cwc2 reveals a novel architecture of a multipartite RNA-binding protein. EMBO J. 31, 2222–2234 (2012).
Urlaub, H., Raker, V.A., Kostka, S. & Lührmann, R. Sm protein-Sm site RNA interactions within the inner ring of the spliceosomal snRNP core structure. EMBO J. 20, 187–196 (2001).
Bessonov, S., Anokhina, M., Will, C., Urlaub, H. & Lührmann, R. Isolation of an active step I spliceosome and composition of its RNP core. Nature 452, 846–850 (2008).
Deckert, J. et al. Protein composition and electron microscopy structure of affinity-purified human spliceosomal B complexes isolated under physiological conditions. Mol. Cell. Biol. 26, 5528–5543 (2006).
Dignam, J.D., Lebovitz, R.M. & Roeder, R.G. Accurate transcription initiation by RNA polymerase II in a soluble extract from isolated mammalian nuclei. Nucleic Acids Res. 11, 1475–1489 (1983).
Rigaut, G. et al. A generic protein purification method for protein complex characterization and proteome exploration. Nat. Biotechnol. 17, 1030–1032 (1999).
Creamer, T.J. et al. Transcriptome-wide binding sites for components of the Saccharomyces cerevisiae non-poly(A) termination pathway: Nrd1, Nab3, and Sen1. PLoS Genet. 7, e1002329 (2011).
Castello, A. et al. System-wide identification of RNA-binding proteins by interactome capture. Nat. Protoc. 8, 491–500 (2013).
Martens, L. et al. mzML—a community standard for mass spectrometry data. Mol. Cell. Proteomics 10, R110.000133 (2011).
Chambers, M.C. et al. A cross-platform toolkit for mass spectrometry and proteomics. Nat. Biotechnol. 30, 918–920 (2012).
Lange, E. et al. A geometric approach for the alignment of liquid chromatography-mass spectrometry data. Bioinformatics 23, i273–i281 (2007).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Sturm, M. & Kohlbacher, O. TOPPView: an open-source viewer for mass spectrometry data. J. Proteome Res. 8, 3760–3763 (2009).
Michalski, A., Neuhauser, N., Cox, J. & Mann, M. A systematic investigation into the nature of tryptic HCD spectra. J. Proteome Res. 11, 5479–5491 (2012).
Olsen, J.V. et al. Parts per million mass accuracy on an Orbitrap mass spectrometer via lock mass injection into a C-trap. Mol. Cell. Proteomics 4, 2010–2021 (2005).
Vizcaíno, J.A. et al. The PRoteomics IDEntifications (PRIDE) database and associated tools: status in 2013. Nucleic Acids Res. 41, D1063–D1069 (2013).
Acknowledgements
We dedicate this article to the memory of our colleague and friend Andreas Bertsch. The authors thank M. Raabe, U. Pleßmann and T. Conrad for technical assistance, U. Zaman for providing unpublished data, R. Hofele for helpful discussions, R. Lührmann for support and providing infrastructure, S. Klinge for help with the yeast ribosome structure in PyMol, and S. Aiche and C. Bielow for discussions on workflow implementation. This work was supported by a German Research Foundation (DFG) grant to H.U. (SFB860, INST 186/859-1), a Higher Education Commission, Pakistan (HEC)/German Academic Exchange Service (DAAD) stipend to S.Q., funding from the European Research Council (ERC) under the Union's Seventh Framework Programme (FP7/2007-2013)/ERC-2011-ADG_20110310 to M.W.H. and a Marie Curie Fellowship (FP7/2007-2013)/MC-IEF-301031 to B.M.B.
Author information
Authors and Affiliations
Contributions
K.K., B.M.B., S.Q., K.-L.B., M.W.H. and H.U. designed biochemical experiments. K.-L.B. and K.K. designed and transformed the yeast strain. K.K. and B.M.B. carried out experiments for the yeast systems; K.K. analyzed the resulting data. S.Q. performed experiments in the human system; K.K. and S.Q. analyzed the resulting data. K.K., T.S., O.K. and H.U. designed data analysis strategy; T.S. implemented it. K.K. and T.S. tested the data analysis tools. K.K., T.S., B.M.B., M.W.H., O.K. and H.U. wrote the paper; all authors contributed comments throughout all stages of the manuscript. K.K., T.S. and S.Q. compiled the supplementary materials.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Details on data preparation in the data analysis workflow.
The raw data, e.g. in Thermo’s raw format, is converted by the appropriate external data conversion tool, e.g. msconvert, into the open mzML format. Next, data in profile mode is centroided. Finally, the data of the control experiment is aligned relative to that of the UV irradiated sample to correct for retention time (RT) shifts. The data preparation steps yield the centroided data of the UV irradiation experiment in mzML format and the aligned, centroided data of the control experiment.
Supplementary Figure 2 Details on data reduction in the data analysis workflow.
The sample data is submitted into the ID filter pipeline. Here, a search is performed against the target-decoy version of a protein database containing the proteome of the species of interest as well as contaminant sequences. Peptides below a certain false discovery rate (FDR; default 1%) are considered as good matches and reported in an idXML file. Corresponding MS/MS spectra are removed from the mzML file. Next, the RNPxlXICFilter tool automatically screens whether the precursors of remaining MS/MS spectra also appear in the non-irradiated control. To this end, extracted ion chromatograms of those precursors are calculated in UV and control sample. MS/MS spectra of precursors appearing in both samples with comparable intensities, i.e. less than double in the UV sample, are not written into the output mzML file.
Supplementary Figure 3 Details on the different data analysis steps performed by the RNPxl tool.
First, spectra of small precursors and those corresponding to short RNA oligonucleotides according to their fractional mass are filtered from the input mzML file. Next, the masses for all possible nucleotide combinations as defined in the tool’s parameters are calculated and subsequently subtracted from all precursor masses to yield the precursor mass variants. One mzML file for each spectrum is created, containing the original experimental precursor mass as well as all its precursor mass variants together with the unaltered MS/MS fragment information. These are submitted into the database search engine. Then, the tool summarizes the search results, retaining the best scoring hit per spectrum and annotates the cross-linked RNA moiety according to the mass difference between experimental precursor mass and the precursor mass variant that gave rise to the database search result. Finally, the results are summarized in csv and idXML format.
Supplementary Figure 4 Alignment of RRM motifs of human proteins in which cross-linked peptides have been identified.
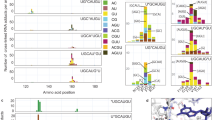
Proteins are identified by their gene names. The corresponding recommended protein names can be found in Supplementary Table 1. The number of the respective RRM and the positions of the first and the last residue within the RRM are given next to the sequences. The RNP2 and RNP1 consensus sequences marked with light gray, with conserved aromatic residues in dark gray. Peptides found cross-linked to RNA are underlined, cross-linked regions are highlighted in yellow. If the cross-linked amino acid was identified from the MS/MS fragment spectrum it is shown in red. Overall, eleven cross-linked residues align with the conserved aromatic residues in the RNP2 and RNP1 consensus sequences that are frequently connected to direct protein–RNA contacts.
Supplementary Figure 5 Alignment of KH motifs of human proteins in which cross-linked peptides have been identified.
Proteins are identified by their gene names, the corresponding recommended protein names can be found in Supplementary Table 1. The number of the respective KH and the positions of the first and the last residue within the KH are given next to the sequences. The consensus VIGXXGXXI map is marked with light gray. Peptides found cross-linked to RNA are underlined, cross-linked regions are highlighted in yellow, if the MS/MS fragment spectrum allowed to narrow down the cross-linking site to a single amino acid residue is shown in red.
Supplementary Figure 6 Alignment of RRM motifs of yeast proteins in which cross-linked peptides have been identified after isolation with TAP purification of Cbp20.
More precisely, alignment of RRM1 and RRM2 of Nucleolar protein 3 (NPL3), RRM2 of Nucleolar protein 13 (NOP13), RRM4 of Polyadenylate-binding protein (PAB1), and RRM2 of Single-stranded nucleic-acid binding protein (SBP1). Positions of the first and the last residue within the RRM are given next to the sequences. All RRMs show good alignment to the RNP2 and RNP1 consensus sequences marked with light gray, with conserved aromatic residues in dark gray. Peptides found cross-linked to RNA are underlined, cross-linked regions are highlighted in yellow; if the cross-linked amino acid was identified from the MS/MS fragment spectrum it is shown in red. Cross-linking in RRM1 of Npl3 occurred through a phenylalanine residue; however the spectrum (Spectrum Y12 in Supplementary Data) does not allow distinguishing whether Phe160, Phe162, or Phe165 is the actual cross-linked amino acid. Phe160 and Phe162 as well as Phe242 of Nop13 and Phe325 of Pab1 correspond to conserved aromatic residues in the respective consensus sequences.
Supplementary Figure 7 RRM motif aligment of yeast proteins cross-linked to 4SU.
Proteins are identified by their gene names, the corresponding recommended protein names can be found in Supplementary Table 3. The number of the respective RRM and the positions of the first and the last residue within the RRM are given next to the sequences. The RNP2 and RNP1 consensus sequences marked with light gray, with conserved aromatic residues in dark gray. Peptides found cross-linked to RNA are underlined, cross-linked regions are highlighted in yellow, if the cross-linked amino acid was identified from the MS/MS fragment spectrum it is shown in red.
Supplementary Figure 8 KH motif alignment of yeast proteins cross-linked to 4SU.
Proteins are identified by their gene names, the corresponding recommended protein names can be found in Supplementary Table 3. The number of the respective KH and the positions of the first and the last residue within the KH are given next to the sequences. The consensus VIGXXGXXI map is marked with light gray. Peptides found cross-linked to RNA are underlined, cross-linked regions are highlighted in yellow, if the MS/MS fragment spectrum allowed to narrow down the cross-linking site to a single amino acid residue is shown in red.
Supplementary Figure 9 Cumulative increase of results depending on RNA combinations considered.
Dependencies are shown for each cross-linking experiment – (a) human RNPs, (b) yeast RNPs isolated with TAP purification of Cbp20, (c) yeast RNPs labeled with 4SU and isolated with oligo d(T). The order of cross-linked (oligo)nucleotides from left to right is first depending on increasing length and then on decreasing number of observed cross-links. (a) Around 31% of cross-links (58 of the total 189) were observed to a single U, which corresponds to 82% (49 of the total 60) of the cross-linking sites/regions identified. At least one modification of U, i.e. loss of water, and two dinucleotides (AU and UU) have to be considered to identify 59 of the 60 identified cross-linking sites/regions. (b) [U –H2O] is the most common RNA moiety, found in 25 cross-links (14%) and 21 unique cross-linking sites/regions (33% of the total results). In contrast to human, a much larger number of RNA combinations and modifications has to be taken into account to identify the cross-linking sites/regions reported in this study. (c) All cross-links to 4SU were observed with a net loss of H2S from the RNA. By taking into account a single 4SU as well as all dinucleotides of the cross-linked 4SU and the four native nucleotides, 269 of the 376 cross-links (72%) and 125 of the 133 unique cross-linking sites/regions (94%) could be identified. Overall, considering only a single nucleotide would only permit identification of 14-33% of the reported cross-links and 33-82% of the reported cross-linking sites/regions in the respective experiments, illustrating the benefit of the precursor variant approach over conventional database search treating one (or a few) cross-linked RNA moieties as a post-translational modification.
Supplementary Figure 10 Size-exclusion chromatograms of unsubstituted yeast (pre-)mRNPs.
The right panel shows the chromatogram of a UV irradiated sample, the left panel a chromatogram of a non-irradiated control. Absorption at 254 nm (red) and 280 nm (blue) were monitored. Fractions and elution time are indicated on the bottom. Fractions #3–#5 were further processed for subsequent LC-MS/MS analysis. No absorbance differences in the chromatograms between control and cross-linked sample are observed as previously described by Urlaub et al.1
1. Urlaub, H., Kruft, V., Bischof, O., Muller, E.C. & Wittmann-Liebold, B. Protein-rRNA binding features and their structural and functional implications in ribosomes as determined by cross-linking studies. EMBO J 14, 4578-4588 (1995).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10, Supplementary Note and Supplementary Data (PDF 14582 kb)
Supplementary Table 1
Cross-links of human RNPs. Detailed information about cross-links from human RNPs including protein and gene names, cross-link composition and positions, experimental and calculated masses, as well as a comparison to other MS-based cross-linking studies. (XLSX 59 kb)
Supplementary Table 2
Cross-links of yeast RNPs. Detailed information about cross-links from yeast RNPs including protein and gene names, cross-link composition and positions, experimental and calculated masses, as well as a comparison to other MS-based cross-linking studies. (XLSX 56 kb)
Supplementary Table 3
Cross-links of yeast RNPs to 4-thio-U substituted RNA. Detailed information about yeast RNPs cross-linked to 4SU substituted RNA including protein and gene names, cross-link composition and positions, experimental and calculated masses, as well as a comparison to other MS-based cross-linking studies. (XLSX 93 kb)
Supplementary Table 4
Comparison of cross-link identification by presented approach and common database search engines. Identification of all cross-links of human RNPs to a single U nucleotide is compared between the approach described in this work and the common database search engines OMSSA and Mascot. (XLSX 157 kb)
Supplementary Table 5
Comparison of cross-links found in unsubstituted yeast (pre-)mRNPs and 4-thio-U labelled yeast mRNPs. Cross-links identified in both yeast systems are compared on both protein and cross-linking region or site level. (XLSX 96 kb)
Supplementary Software
RNPxl software. Software updates and a FAQ can be found at www.openms.de/RNPxl. (ZIP 156890 kb)
Rights and permissions
About this article
Cite this article
Kramer, K., Sachsenberg, T., Beckmann, B. et al. Photo-cross-linking and high-resolution mass spectrometry for assignment of RNA-binding sites in RNA-binding proteins. Nat Methods 11, 1064–1070 (2014). https://doi.org/10.1038/nmeth.3092
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3092