Abstract
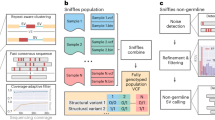
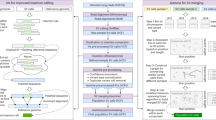
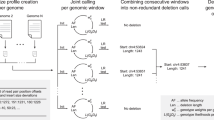
We present an open-source algorithm, Scalpel (http://scalpel.sourceforge.net/), which combines mapping and assembly for sensitive and specific discovery of insertions and deletions (indels) in exome-capture data. A detailed repeat analysis coupled with a self-tuning k-mer strategy allows Scalpel to outperform other state-of-the-art approaches for indel discovery, particularly in regions containing near-perfect repeats. We analyzed 593 families from the Simons Simplex Collection and demonstrated Scalpel's power to detect long (≥30 bp) transmitted events and enrichment for de novo likely gene-disrupting indels in autistic children.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
DePristo, M.A. et al. Nat. Genet. 43, 491–498 (2011).
O'Rawe, J. et al. Genome Med. 5, 28 (2013).
Zook, J.M. et al. Nat. Biotechnol. 32, 246–251 (2014).
Mullaney, J.M., Mills, R.E., Pittard, W.S. & Devine, S.E. Hum. Mol. Genet. 19, R131–R136 (2010).
Pearson, C.E., Edamura, N.K. & Cleary, J.D. Nat. Rev. Genet. 6, 729–742 (2005).
Iossifov, I. et al. Neuron 74, 285–299 (2012).
Li, H., Ruan, J. & Durbin, R. Genome Res. 18, 1851–1858 (2008).
Montgomery, S.B. et al. Genome Res. 23, 749–761 (2013).
Albers, C.A. et al. Genome Res. 21, 961–973 (2011).
Ye, K., Schulz, M.H., Long, Q., Apweiler, R. & Ning, Z. Bioinformatics 25, 2865–2871 (2009).
Karakoc, E. et al. Nat. Methods 9, 176–178 (2012).
Li, Y. et al. Nat. Biotechnol. 29, 723–730 (2011).
Li, H. Bioinformatics 28, 1838–1844 (2012).
Li, S. et al. Genome Res. 23, 195–200 (2013).
Iqbal, Z., Caccamo, M., Turner, I., Flicek, P. & McVean, G. Nat. Genet. 44, 226–232 (2012).
Chen, K. et al. Genome Res. 24, 310–317 (2014).
Li, H. et al. Bioinformatics 25, 2078–2079 (2009).
Garrison, E. & Marth, G. Haplotype-based variant detection from short-read sequencing. Preprint at http://arxiv.org/abs/1207.3907v2 (2012).
Gymrek, M., Golan, D., Rosset, S. & Erlich, Y. Genome Res. 22, 1154–1162 (2012).
Highnam, G. et al. Nucleic Acids Res. 41, e32 (2013).
MacArthur, D.G. & Tyler-Smith, C. Hum. Mol. Genet. 19, R125–R130 (2010).
Sjödin, P., Bataillon, T. & Schierup, M.H. PLoS ONE 5, e8650 (2010).
Sanders, S.J. et al. Nature 485, 237–241 (2012).
O'Roak, B.J. et al. Nature 485, 246–250 (2012).
Neale, B.M. et al. Nature 485, 242–245 (2012).
Darnell, J.C. et al. Cell 146, 247–261 (2011).
Nagarajan, N. & Pop, M. Nat. Rev. Genet. 14, 157–167 (2013).
Li, H. & Durbin, R. Bioinformatics 26, 589–595 (2010).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. Preprint at http://arxiv.org/abs/1303.3997v1 (2013).
Smith, T.F. & Waterman, M.S. J. Mol. Biol. 147, 195–197 (1981).
Medvedev, P., Georgiou, K., Myers, G. & Brudno, M. Lect. Notes Comput. Sci. 4645, 289–301 (2007).
Jackson, B.G. & Aluru, S. in 37th Int. Conf. Parallel Process. 346–353 (ICPP, 2008).
Narzisi, G. & Mishra, B. Bioinformatics 27, 153–160 (2011).
Langmead, B. & Salzberg, S. Nat. Methods 9, 357–359 (2012).
Fischbach, G.D. & Lord, C. Neuron 68, 192–195 (2010).
Acknowledgements
The project was supported in part by the US National Institutes of Health (R01-HG006677) and US National Science Foundation (DBI-1350041) to M.C.S. and by the Cold Spring Harbor Laboratory (CSHL) Cancer Center Support Grant (5P30CA045508), the Stanley Institute for Cognitive Genomics and the Simons Foundation (SF51 and SF235988) to M.W. The DNA samples used in this work are included within SSC release 13. Approved researchers can obtain the SSC population data set described in this study by applying at https://base.sfari.org/. We thank S. Eskipehlivan for the technical assistance with the MiSeq validation experiments. We thank M. Bekritsky, S. Neuburgerand, M. Ronemus, D. Levy, B. Yamron and B. Mishra for helpful discussions and comments on the paper. We thank R. Aboukhalil for testing the software.
Author information
Authors and Affiliations
Contributions
G.N. developed the software and conducted the computational experiments. G.N. and M.C.S. designed and analyzed the experiments. Y.W. assisted in designing the primers and performed the MiSeq validation experiments. J.A.O. designed the primers and analyzed the MiSeq data. H.F. and J.A.O. assisted with the computational experiments for the comparative analysis between different variant-detection pipelines. G.J.L. planned and supervised the experimental design for indel validation. Z.W. designed the primers and performed experiments for the validation of de novo and transmitted indels in the SSC. I.I., Y.-h.L. and M.W. assisted with the analysis of the SSC. G.N. and M.C.S. wrote the manuscript with input from all authors. All of the authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–20, Supplementary Tables 1–11 and Supplementary Notes 1–6 (PDF 3587 kb)
Supplementary Data 1
Scripts and data used to generate Figure 3b (ZIP 225 kb)
Supplementary Data 2
List of 97 high-quality de novo indels in 593 SSC families (XLS 47 kb)
Rights and permissions
About this article
Cite this article
Narzisi, G., O'Rawe, J., Iossifov, I. et al. Accurate de novo and transmitted indel detection in exome-capture data using microassembly. Nat Methods 11, 1033–1036 (2014). https://doi.org/10.1038/nmeth.3069
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3069
This article is cited by
-
A comparative study of next-generation sequencing and fragment analysis for the detection and allelic ratio determination of FLT3 internal tandem duplication
Diagnostic Pathology (2022)
-
ESR1 hotspot mutations in endometrial stromal sarcoma with high-grade transformation and endocrine treatment
Modern Pathology (2022)
-
Low neoantigen expression and poor T-cell priming underlie early immune escape in colorectal cancer
Nature Cancer (2021)
-
TERT promoter hotspot mutations and gene amplification in metaplastic breast cancer
npj Breast Cancer (2021)
-
Massively parallel sequencing analysis of 68 gastric-type cervical adenocarcinomas reveals mutations in cell cycle-related genes and potentially targetable mutations
Modern Pathology (2021)