Abstract
Many biological processes are RNA-mediated, but higher-order structures for most RNAs are unknown, which makes it difficult to understand how RNA structure governs function. Here we describe selective 2′-hydroxyl acylation analyzed by primer extension and mutational profiling (SHAPE-MaP) that makes possible de novo and large-scale identification of RNA functional motifs. Sites of 2′-hydroxyl acylation by SHAPE are encoded as noncomplementary nucleotides during cDNA synthesis, as measured by massively parallel sequencing. SHAPE-MaP–guided modeling identified greater than 90% of accepted base pairs in complex RNAs of known structure, and we used it to define a new model for the HIV-1 RNA genome. The HIV-1 model contains all known structured motifs and previously unknown elements, including experimentally validated pseudoknots. SHAPE-MaP yields accurate and high-resolution secondary-structure models, enables analysis of low-abundance RNAs, disentangles sequence polymorphisms in single experiments and will ultimately democratize RNA-structure analysis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Sharp, P.A. The centrality of RNA. Cell 136, 577–580 (2009).
Dethoff, E.A., Chugh, J., Mustoe, A.M. & Al-Hashimi, H.M. Functional complexity and regulation through RNA dynamics. Nature 482, 322–330 (2012).
Weeks, K.M. Advances in RNA structure analysis by chemical probing. Curr. Opin. Struct. Biol. 20, 295–304 (2010).
Mathews, D.H. et al. Incorporating chemical modification constraints into a dynamic programming algorithm for prediction of RNA secondary structure. Proc. Natl. Acad. Sci. USA 101, 7287–7292 (2004).
Kertesz, M. et al. Genome-wide measurement of RNA secondary structure in yeast. Nature 467, 103–107 (2010).
Mauger, D.M. & Weeks, K.M. Toward global RNA structure analysis. Nat. Biotechnol. 28, 1178–1179 (2010).
Underwood, J.G. et al. FragSeq: transcriptome-wide RNA structure probing using high-throughput sequencing. Nat. Methods 7, 995–1001 (2010).
Lucks, J.B. et al. Multiplexed RNA structure characterization with selective 2′-hydroxyl acylation analyzed by primer extension sequencing (SHAPE-Seq). Proc. Natl. Acad. Sci. USA 108, 11063–11068 (2011).
Weeks, K.M. RNA structure probing dash seq. Proc. Natl. Acad. Sci. USA 108, 10933–10934 (2011).
Ding, Y. et al. In vivo genome-wide profiling of RNA secondary structure reveals novel regulatory features. Nature 505, 696–700 (2014).
Rouskin, S., Zubradt, M., Washietl, S., Kellis, M. & Weissman, J.S. Genome-wide probing of RNA structure reveals active unfolding of mRNA structures in vivo. Nature 505, 701–705 (2014).
Grohman, J.K. et al. A guanosine-centric mechanism for RNA chaperone function. Science 340, 190–195 (2013).
Wilkinson, K.A. et al. High-throughput SHAPE analysis reveals structures in HIV-1 genomic RNA strongly conserved across distinct biological states. PLoS Biol. 6, e96 (2008).
Gherghe, C. et al. Definition of a high-affinity Gag recognition structure mediating packaging of a retroviral RNA genome. Proc. Natl. Acad. Sci. USA 107, 19248–19253 (2010).
Tyrrell, J., McGinnis, J.L., Weeks, K.M. & Pielak, G.J. The cellular environment stabilizes adenine riboswitch RNA structure. Biochemistry 52, 8777–8785 (2013).
McGinnis, J.L. & Weeks, K.M. Ribosome RNA assembly intermediates visualized in living cells. Biochemistry 53, 3237–3247 (2014).
Mortimer, S.A. & Weeks, K.M. A fast-acting reagent for accurate analysis of RNA secondary and tertiary structure by SHAPE chemistry. J. Am. Chem. Soc. 129, 4144–4145 (2007).
Weeks, K.M. & Mauger, D.M. Exploring RNA structural codes with SHAPE chemistry. Acc. Chem. Res. 44, 1280–1291 (2011).
Rice, G.M., Leonard, C.W. & Weeks, K.M. RNA secondary structure modeling at consistent high accuracy using differential SHAPE. RNA 20, 846–854 (2014).
Merino, E.J., Wilkinson, K.A., Coughlan, J.L. & Weeks, K.M. RNA structure analysis at single nucleotide resolution by selective 2′-hydroxyl acylation and primer extension (SHAPE). J. Am. Chem. Soc. 127, 4223–4231 (2005).
Spitale, R.C. et al. RNA SHAPE analysis in living cells. Nat. Chem. Biol. 9, 18–20 (2013).
Hajdin, C.E. et al. Accurate SHAPE-directed RNA secondary structure modeling, including pseudoknots. Proc. Natl. Acad. Sci. USA 110, 5498–5503 (2013).
Steen, K.-A., Rice, G.M. & Weeks, K.M. Fingerprinting noncanonical and tertiary RNA structures by differential SHAPE reactivity. J. Am. Chem. Soc. 134, 13160–13163 (2012).
Doty, P., Boedtker, H., Fresco, J.R., Haselkorn, R. & Litt, M. Secondary structure in ribonucleic acids. Proc. Natl. Acad. Sci. USA 45, 482–499 (1959).
Huynen, M., Gutell, R. & Konings, D. Assessing the reliability of RNA folding using statistical mechanics. J. Mol. Biol. 267, 1104–1112 (1997).
Mathews, D.H. Using an RNA secondary structure partition function to determine confidence in base pairs predicted by free energy minimization. RNA 10, 1178–1190 (2004).
Watts, J.M. et al. Architecture and secondary structure of an entire HIV-1 RNA genome. Nature 460, 711–716 (2009).
Staple, D.W. & Butcher, S.E. Pseudoknots: RNA structures with diverse functions. PLoS Biol. 3, e213 (2005).
Brierley, I., Pennell, S. & Gilbert, R.J.C. Viral RNA pseudoknots: versatile motifs in gene expression and replication. Nat. Rev. Microbiol. 5, 598–610 (2007).
Paillart, J.-C., Skripkin, E., Ehresmann, B., Ehresmann, C. & Marquet, R. In vitro evidence for a long range pseudoknot in the 5′-untranslated and matrix coding regions of HIV-1 genomic RNA. J. Biol. Chem. 277, 5995–6004 (2002).
Resch, W., Ziermann, R., Parkin, N., Gamarnik, A. & Swanstrom, R. Nelfinavir-resistant, amprenavir-hypersusceptible strains of human immunodeficiency virus type 1 carrying an N88S mutation in protease have reduced infectivity, reduced replication capacity, and reduced fitness and process the Gag polyprotein precursor aberrantly. J. Virol. 76, 8659–8666 (2002).
Matoulkova, E., Michalova, E., Vojtesek, B. & Hrstka, R. The role of the 3′ untranslated region in post-transcriptional regulation of protein expression in mammalian cells. RNA Biol. 9, 563–576 (2012).
Gilmartin, G.M., Fleming, E.S. & Oetjen, J. Activation of HIV-1 pre-mRNA 3′ processing in vitro requires both an upstream element and TAR. EMBO J. 11, 4419–4428 (1992).
Klasens, B.I., Thiesen, M., Virtanen, A. & Berkhout, B. The ability of the HIV-1 AAUAAA signal to bind polyadenylation factors is controlled by local RNA structure. Nucleic Acids Res. 27, 446–454 (1999).
Ghadessy, F.J. & Holliger, P. Compartmentalized self-replication: a novel method for the directed evolution of polymerases and other enzymes. Methods Mol. Biol. 352, 237–248 (2007).
Chen, T. & Romesberg, F.E. Directed polymerase evolution. FEBS Lett. 588, 219–229 (2014).
Pollom, E. et al. Comparison of SIV and HIV-1 genomic RNA structures reveals impact of sequence evolution on conserved and non-conserved structural motifs. PLoS Pathog. 9, e1003294 (2013).
Deigan, K.E., Li, T.W., Mathews, D.H. & Weeks, K.M. Accurate SHAPE-directed RNA structure determination. Proc. Natl. Acad. Sci. USA 106, 97–102 (2009).
Lorsch, J.R., Bartel, D.P. & Szostak, J.W. Reverse transcriptase reads through a 2′–5′ linkage and a 2′-thiphosphate in a template. Nucleic Acids Res. 23, 2811–2814 (1995).
Patterson, J.T., Nickens, D.G. & Burke, D.H. HIV-1 reverse transcriptase pausing at bulky 2′ adducts is relieved by deletion of the RNase H domain. RNA Biol. 3, 163 (2006).
Roth, M.J., Tanese, N. & Goff, S.P. Purification and characterization of murine retroviral reverse transcriptase expressed in Escherichia coli. J. Biol. Chem. 260, 9326–9335 (1985).
Beckman, R.A., Mildvan, A.S. & Loeb, L.A. On the fidelity of DNA replication: manganese mutagenesis in vitro. Biochemistry 24, 5810–5817 (1985).
Wilkinson, K.A., Merino, E.J. & Weeks, K.M. Selective 2′-hydroxyl acylation analyzed by primer extension (SHAPE): quantitative RNA structure analysis at single nucleotide resolution. Nat. Protoc. 1, 1610–1616 (2006).
Williams, R. et al. Amplification of complex gene libraries by emulsion PCR. Nat. Methods 3, 545–550 (2006).
Langmead, B. & Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Hunter, J.D. Matplotlib: A 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Reuter, J.S. & Mathews, D.H. RNAstructure: software for RNA secondary structure prediction and analysis. BMC Bioinformatics 11, 129 (2010).
Byun, Y. & Han, K. PseudoViewer: web application and web service for visualizing RNA pseudoknots and secondary structures. Nucleic Acids Res. 34, W416–W422 (2006).
Zhang, J., Chung, T. & Oldenburg, K. A simple statistical parameter for use in evaluation and validation of high throughput screening assays. J. Biomol. Screen. 4, 67–73 (1999).
Wan, Y. et al. Landscape and variation of RNA secondary structure across the human transcriptome. Nature 505, 706–709 (2014).
Talkish, J., May, G., Lin, Y., Woolford, J.L. & McManus, C.J. Mod-seq: high-throughput sequencing for chemical probing of RNA structure. RNA 20, 713–720 (2014).
Adachi, A. et al. Production of acquired immunodeficiency syndrome-associated retrovirus in human and nonhuman cells transfected with an infectious molecular clone. J. Virol. 59, 284–291 (1986).
Derdeyn, C.A. et al. Sensitivity of human immunodeficiency virus type 1 to the fusion inhibitor T-20 is modulated by coreceptor specificity defined by the V3 loop of gp120. J. Virol. 74, 8358–8367 (2000).
Harrich, D. et al. Role of SP1-binding domains in in vivo transcriptional regulation of the human immunodeficiency virus type 1 long terminal repeat. J. Virol. 63, 2585–2591 (1989).
Jabara, C.B., Jones, C.D., Roach, J., Anderson, J.A. & Swanstrom, R. Accurate sampling and deep sequencing of the HIV-1 protease gene using a Primer ID. Proc. Natl. Acad. Sci. USA 108, 20166–20171 (2011).
Magoč, T. & Salzberg, S.L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Acknowledgements
We thank R.J. Gorelick for expert preparation of HIV-1 genomic RNA and many insightful discussions, C.E. Hajdin for extensive discussion and for initial pseudoknot mutant design, R. Swanstrom for critical discussions, K. Compliment for expert technical assistance and W. Resch for competition calculations. TZM-bl cells were obtained from J. Kappes and X. Wu (Tranzyme Inc.) via the US National Institutes of Health (NIH) AIDS Reagent Program. This work was supported by the NIH (AI068462 to K.M.W.) and the University of North Carolina Center for AIDS Research (P30 AI50410). N.A.S. was funded as a Lineberger Postdoctoral Fellow in the Basic Sciences and by a Ruth L. Kirschstein NRSA Fellowship (F32 GM010169). G.M.R. was supported in part by an NIH training grant in molecular and cellular biophysics (T32 GM08570).
Author information
Authors and Affiliations
Contributions
The SHAPE-MaP strategy was conceived and designed by N.A.S. and K.M.W. SHAPE experiments were performed by N.A.S. and G.M.R. HIV-1 replication assays were designed and performed by N.A.S. and J.A.E.N. The SHAPE-MaP data analysis pipeline was created by S.B. RNA folding and motif discovery analyses were conceived and created by G.M.R. and K.M.W. All authors collaborated in interpreting the experiments and writing the manuscript.
Corresponding author
Ethics declarations
Competing interests
N.A.S., S.B. and K.M.W. are listed as inventors on a US provisional patent application based on elements of this work.
Integrated supplementary information
Supplementary Figure 1 SHAPE-MaP data analysis pipeline.
Outline of software pipeline that fully automates calculations of per-nucleotide mutation rates, SHAPE reactivities, and standard error estimates given high-throughput sequencing data and at least one reference sequence. The software is executable on Unix-based platforms. See Online Methods for full description. This strategy is implemented in the ShapeMapper software.
Supplementary Figure 2 Overview of multiscale RNA secondary structure modeling.
Genome-scale RNA secondary structure modeling was performed in steps to increase computational efficiency and model accuracy and to facilitate incorporation of pseudoknot prediction into a global model. (top) The first step involved searching for pseudoknots in short windows. For all later steps, pseudoknot pairs were prohibited from forming base pairs. In the second step, the partition function was calculated in overlapping windows and averaged for Shannon entropy and pairing probability evaluations (see Online Methods). Base pairs with probabilities ≥99% were forced to form during calculation of a minimum free energy structure using Fold47, using overlapping 4000-nt windows. In the third step, a consensus structure from the overlapping windows was generated by retaining base pairs that appeared in more than half of possible windows. Finally, pseudoknotted helices were added to the final model. (bottom) Comparison of windowed folding versus one-step folding for calculating the partition functions and minimum free energy structures for RNAs of 1500 to 9200 nts. Wall-time for modeling the entire HIV-1 was estimated (asterisk). A small performance penalty is observed for splitting an RNA into overlapping windows. However, computation time for RNAs over 3000 nts will scale approximately linearly with sequence length. Folding times are reported both as wall clock times and as cpu cycles (2012 iMac with 3.1 GHz Intel Core i7 and 16 GB RAM). This strategy is implemented in the SHAPE-MaP Folding Pipeline.
Supplementary Figure 3 Strategies for the SHAPE-MaP experiment using either gene-specific primers or random fragmentation for sample analysis and sequencing library preparation.
SHAPE-MaP can be performed using gene-specific primers (for small RNAs or targeted areas in large RNAs and for analysis of scarce and low concentration RNAs) or random primers (for comprehensive analysis of large RNAs or complete transcriptomes) to create the initial cDNA pool. For both approaches, RNA is treated with a SHAPE reagent or with solvent under conditions of interest, and a sample of RNA is modified under denaturing conditions. For gene-specific samples, reverse transcription and PCR primers are designed based on the known target sequence. Large RNAs are randomly fragmented in a buffered Mg2+ solution. Single-stranded cDNA was synthesized using mutation-prone reverse transcription; misincorporation events in the nascent cDNA mark the location of SHAPE adducts in the subject RNA. Double-stranded cDNAs were created either by PCR (gene-specific approach) or second-strand synthesis (randomly fragmented samples). Sequence platform-specific sequences (including multiplexing barcodes) were added to the dsDNA libraries, either directly through a second PCR (gene-specific approach) or by a DNA-DNA ligation of adaptor sequences (random fragmented samples). Libraries prepared by either method were then sequenced, producing data that were processed into SHAPE reactivity profiles used in structure modeling applications. SHAPE-MaP is fully independent of sequencing platform and library generation scheme (once the initial cDNA has been synthesized). Thus, any platform and any library generation scheme can be used.
Supplementary Figure 4 Mutation rate histograms for paired and non-paired nucleotides in the 16S rRNA.
Nucleotides were separated into paired (upper panels) and non-paired (lower panels) groups based on their observed pairing in the E. coli 16S rRNA38. Mutation rate histograms for each experimental sample (SHAPE, untreated, and denatured) were calculated based on pairing status (left-hand panels). Distributions of mutation rates for the SHAPE-modified and untreated samples are similar for base-paired nucleotides; whereas nucleotides in non-paired conformations are much more reactive towards SHAPE probing. (right-hand panels) SHAPE-MaP reactivities are independent of nucleotide type.
Supplementary Figure 5 SHAPE-MaP replicates of E. coli 16S rRNA.
Data correspond to full biological replicates performed six months apart by different individuals. The inset for nucleotides 1350-1450 (bottom right) shows standard errors.
Supplementary Figure 6 Error analysis for SHAPE-MaP.
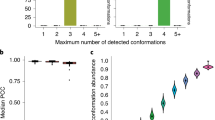
Deep bootstrapping of highly sequenced TPP riboswitch samples (see Fig. 2). Individual sequencing reads from a large pool (150,000) were sampled with replacement 100 times per simulated depth. The standard error of the SHAPE reactivity was calculated at each depth from each bootstrap. Consistent with a Poisson model, the standard error of the SHAPE measurement decreased as the -1/2 power of read depth across all nucleotides.
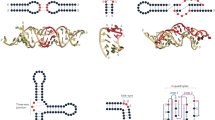
Supplementary Figure 7 Secondary structure models for regions of HIV-1NL4-3, identified de novo, with low SHAPE reactivities and low Shannon entropies.
Nucleotides are colored by SHAPE reactivity. Structures are the same as shown in Fig. 4c, except that nucleotide identities are shown explicitly.
Supplementary Figure 8 Pseudoknot mutants.
Green arrows indicate sites of disruptive mutations.
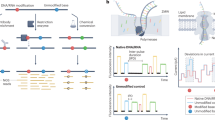
Supplementary Figure 10 Pseudoknot SHAPE-MaP profiles for ENVPK and CAPK.
(upper panel) SHAPE-MaP and structure profiles for ENVPK and direct growth competition and viral spread data. (lower panel) SHAPE-MaP and structure profiles for CAPK, located in a high entropy region of the RNA genome and thus served as a negative control. Also displayed are competition and viral spread assay data.
Supplementary Figure 11 Detection of 2'-O-adducts by mutational profiling.
Shown are rates for sequence changes and unambiguously aligned deletions, above background for the E. coli 16S rRNA. Nucleotides were defined as non-paired or paired based on the accepted secondary structure. The letter in the lower right of each panel indicates the expected nucleotide based on the coding strand, and the letters on the vertical axes indicate the nucleotide detected by sequencing or “del” for deletion. Rates are shown for the (a) 1M7, (b) 1M6, and (c) NMIA reagents. Nucleotide misincorporation and deletion rates were similar for the three SHAPE reagents.
Supplementary Figure 12 Primer design for SHAPE-MaP.
Sequences with low or unevenly distributed GC-content benefit from the newly designed LNA-based primers used to analyze HIV-1 sequences in this work.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12 and Supplementary Tables 1–3 (PDF 5911 kb)
Supplementary Data 1
Full SHAPE dataset for the HIV-1 RNA genome. (XLSX 752 kb)
Supplementary Data 2
Structure models for each well-defined region in the HIV-1 RNA genome, in connect-table format. (ZIP 46 kb)
Supplementary Data 3
Pairing probabilities for HIV-1 nucleotides, in tab-delimited text format. (TXT 190 kb)
Supplementary Data 4
Complete differential SHAPE-Map data for the model RNAs reported in Figure 3. (ZIP 233 kb)
Supplementary Software 1
Software pipeline for analyzing MaP data. Illustrated in Supplementary Figure 1. (ZIP 76 kb)
Supplementary Software 2
Pipeline for automated windowed modeling approach for folding long RNAs and calculating pairing probabilities and Shannon entropies. Illustrated in Supplementary Figure 2. (ZIP 205 kb)
Rights and permissions
About this article
Cite this article
Siegfried, N., Busan, S., Rice, G. et al. RNA motif discovery by SHAPE and mutational profiling (SHAPE-MaP). Nat Methods 11, 959–965 (2014). https://doi.org/10.1038/nmeth.3029
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3029
This article is cited by
-
RIP-PEN-seq identifies a class of kink-turn RNAs as splicing regulators
Nature Biotechnology (2024)
-
NAP-seq reveals multiple classes of structured noncoding RNAs with regulatory functions
Nature Communications (2024)
-
Dengue and Zika RNA-RNA interactomes reveal pro- and anti-viral RNA in human cells
Genome Biology (2023)
-
Probing the dynamic RNA structurome and its functions
Nature Reviews Genetics (2023)
-
Nano-DMS-MaP allows isoform-specific RNA structure determination
Nature Methods (2023)