Abstract
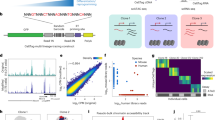
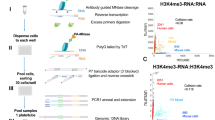
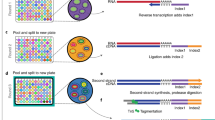
We have developed a quantitative technique for sorting cells on the basis of endogenous RNA abundance, with a molecular resolution of 10–20 transcripts. We demonstrate efficient and unbiased RNA extraction from transcriptionally sorted cells and report a high-fidelity transcriptome measurement of mouse induced pluripotent stem cells (iPSCs) isolated from a heterogeneous reprogramming culture. This method is broadly applicable to profiling transcriptionally distinct cellular states without requiring antibodies or transgenic fluorescent proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Chalfie, M., Tu, Y., Euskirchen, G., Ward, W.W. & Prasher, D.C. Science 263, 802–805 (1994).
Chambers, I. et al. Nature 450, 1230–1234 (2007).
Dietrich, J.-E. & Hiiragi, T. Development 134, 4219–4231 (2007).
Rufer, N., Dragowska, W., Thornbury, G., Roosnek, E. & Lansdorp, P. Nat. Biotechnol. 16, 743–747 (1998).
Prigodich, A.E. et al. Anal. Chem. 84, 2062–2066 (2012).
Robertson, K.L., Verhoeven, A.B., Thach, D.C. & Chang, E.L. RNA 16, 1679–1685 (2010).
Rhee, W.J. & Bao, G. BMC Biotechnol. 9, 30 (2009).
Masuda, N., Ohnishi, T., Kawamoto, S., Monden, M. & Okubo, K. Nucleic Acids Res. 27, 4436–4443 (1999).
Specht, K. et al. Am. J. Pathol. 158, 419–429 (2001).
Yamada, H. et al. Cytometry A 77, 1032–1037 (2010).
Maruo, R. et al. Mol. Biotechnol. 49, 42–47 (2011).
Larsson, H.M. et al. PLoS ONE 7, e49874 (2012).
Femino, A.M., Fay, F.S., Fogarty, K. & Singer, R.H. Science 280, 585–590 (1998).
Raj, A., Bogaard, P.V.D., Rifkin, S.A., van Oudenaarden, A. & Tyagi, S. Nat. Methods 5, 877–879 (2008).
Beard, C., Hochedlinger, K., Plath, K., Wutz, A. & Jaenisch, R. Genesis 44, 23–28 (2006).
Takahashi, K. & Yamanaka, S. Cell 126, 663–676 (2006).
Hanna, J. et al. Nature 462, 595–601 (2009).
Buganim, Y. et al. Cell 150, 1209–1222 (2012).
Golipour, A. et al. Cell Stem Cell 11, 769–782 (2012).
Wernig, M. et al. Nat. Biotechnol. 26, 916–924 (2008).
Acknowledgements
We thank M. Lou for performing the microarray experiments and M. Bienko, N. Crosetto and N. Slavov for critical reading of the manuscript. This work was supported by the US National Institutes of Health (NIH) National Cancer Institute Physical Sciences Oncology Center at Massachusetts Institute of Technology (U54CA143874), an NIH Pioneer award (8 DP1 CA174420-05), a Nederlandse Organisatie voor Wetenschappelijk Onderzoek (NWO) Vici award to A.v.O. and an NWO Rubicon award to S.S. R.J. was supported by NIH grants HD 045022 and R37CA084198. D.A.F. was supported by a Vertex Scholarship, a US National Science Foundation Graduate Research Fellowship and Jerome and Florence Brill Graduate Student Fellowship. Support for S.K. was provided by the Koch Institute for Integrative Cancer Research Graduate Fellowship.
Author information
Authors and Affiliations
Contributions
A.v.O., S.K. and S.S. developed the idea of transcriptionally profiling RNA-sorted cells. S.S. and S.K. demonstrated the compatibility of smFISH with flow cytometry. S.K. performed all of the experiments; K.W. collaborated on the RNA integrity measurements in Supplementary Figure 4 and the reverse cross-linking controls in Figure 2. S.K., S.S., K.W. and A.v.O. developed the reverse cross-linking protocol. S.K. and D.M. optimized the RNA flow sorting procedure. S.K. conceived and experimentally validated the RNA-preserving hybridization buffer (RPHB). D.A.F. and R.J. produced the 2° reprogrammable MEFs. S.K. analyzed the data, developed the analytic estimate of the molecular resolution, prepared the figures, and wrote the manuscript in collaboration with A.v.O., who guided the project. All authors read and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Note (PDF 8051 kb)
Supplementary Data
Probe sequences of smFISH libraries (XLS 35 kb)
Rights and permissions
About this article
Cite this article
Klemm, S., Semrau, S., Wiebrands, K. et al. Transcriptional profiling of cells sorted by RNA abundance. Nat Methods 11, 549–551 (2014). https://doi.org/10.1038/nmeth.2910
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2910
This article is cited by
-
Comparison of bias and resolvability in single-cell and single-transcript methods
Communications Biology (2021)
-
Microdroplet-based one-step RT-PCR for ultrahigh throughput single-cell multiplex gene expression analysis and rare cell detection
Scientific Reports (2021)
-
ClampFISH detects individual nucleic acid molecules using click chemistry–based amplification
Nature Biotechnology (2019)
-
RNA-Seq following PCR-based sorting reveals rare cell transcriptional signatures
BMC Genomics (2016)
-
Fixed single-cell transcriptomic characterization of human radial glial diversity
Nature Methods (2016)