Abstract
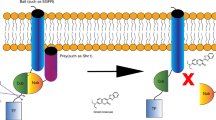
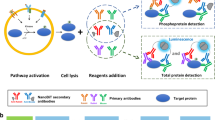
Cell signaling, one of key processes in both normal cellular function and disease, is coordinated by numerous interactions between membrane proteins that change in response to stimuli. We present a split ubiquitin–based method for detection of integral membrane protein-protein interactions (PPIs) in human cells, termed mammalian-membrane two-hybrid assay (MaMTH). We show that this technology detects stimulus (hormone or agonist)-dependent and phosphorylation-dependent PPIs. MaMTH can detect changes in PPIs conferred by mutations such as those in oncogenic ErbB receptor variants or by treatment with drugs such as the tyrosine kinase inhibitor erlotinib. Using MaMTH as a screening assay, we identified CRKII as an interactor of oncogenic EGFR(L858R) and showed that CRKII promotes persistent activation of aberrant signaling in non–small cell lung cancer cells. MaMTH is a powerful tool for investigating the dynamic interactomes of human integral membrane proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Stevens, T.J. & Arkin, I.T. Do more complex organisms have a greater proportion of membrane proteins in their genomes? Proteins 39, 417–420 (2000).
Jones, R.B., Gordus, A., Krall, J.A. & MacBeath, G. A quantitative protein interaction network for the ErbB receptors using protein microarrays. Nature 439, 168–174 (2006).
Schulze, W.X., Deng, L. & Mann, M. Phosphotyrosine interactome of the ErbB-receptor kinase family. Mol. Syst. Biol. 1, 2005.0008 (2005).
Lievens, S., Lemmens, I. & Tavernier, J. Mammalian two-hybrids come of age. Trends Biochem. Sci. 34, 579–588 (2009).
Michnick, S.W., Ear, P.H., Manderson, E.N., Remy, I. & Stefan, E. Universal strategies in research and drug discovery based on protein-fragment complementation assays. Nat. Rev. Drug Discov. 6, 569–582 (2007).
Rojo-Niersbach, E., Morley, D., Heck, S. & Lehming, N. A new method for the selection of protein interactions in mammalian cells. Biochem. J. 348, 585–590 (2000).
Kerppola, T.K. Visualization of molecular interactions by fluorescence complementation. Nat. Rev. Mol. Cell Biol. 7, 449–456 (2006).
Stefan, E. et al. Quantification of dynamic protein complexes using Renilla luciferase fragment complementation applied to protein kinase A activities in vivo. Proc. Natl. Acad. Sci. USA 104, 16916–16921 (2007).
Wehrman, T.S. et al. A system for quantifying dynamic protein interactions defines a role for Herceptin in modulating ErbB2 interactions. Proc. Natl. Acad. Sci. USA 103, 19063–19068 (2006).
Wehr, M.C. et al. Monitoring regulated protein-protein interactions using split TEV. Nat. Methods 3, 985–993 (2006).
Ciruela, F. Fluorescence-based methods in the study of protein-protein interactions in living cells. Curr. Opin. Biotechnol. 19, 338–343 (2008).
Johnsson, N. & Varshavsky, A. Split ubiquitin as a sensor of protein interactions in vivo. Proc. Natl. Acad. Sci. USA 91, 10340–10344 (1994).
Stagljar, I., Korostensky, C., Johnsson, N. & te Heesen, S. A genetic system based on split-ubiquitin for the analysis of interactions between membrane proteins in vivo. Proc. Natl. Acad. Sci. USA 95, 5187–5192 (1998).
Paumi, C.M. et al. Mapping protein-protein interactions for the yeast ABC transporter Ycf1p by integrated split-ubiquitin membrane yeast two-hybrid analysis. Mol. Cell 26, 15–25 (2007).
Snider, J. et al. Mapping the functional yeast ABC transporter interactome. Nat. Chem. Biol. 9, 565–572 (2013).
Emami, K.H. & Carey, M. A synergistic increase in potency of a multimerized VP16 transcriptional activation domain. EMBO J. 11, 5005–5012 (1992).
Sadowski, I., Ma, J., Triezenberg, S. & Ptashne, M. GAL4–VP16 is an unusually potent transcriptional activator. Nature 335, 563–564 (1988).
Schmitz, M.L. & Baeuerle, P.A. The p65 subunit is responsible for the strong transcription activating potential of NF-kappa B. EMBO J. 10, 3805–3817 (1991).
Remy, I. & Michnick, S.W. Application of protein-fragment complementation assays in cell biology. Biotechniques 42, 137–141 (2007).
Johnsson, N. & Varshavsky, A. Ubiquitin-assisted dissection of protein transport across membranes. EMBO J. 13, 2686–2698 (1994).
Rhee, Y., Gurel, F., Gafni, Y., Dingwall, C. & Citovsky, V. A genetic system for detection of protein nuclear import and export. Nat. Biotechnol. 18, 433–437 (2000).
Deribe, Y.L. et al. Regulation of epidermal growth factor receptor trafficking by lysine deacetylase HDAC6. Sci. Signal. 2, ra84 (2009).
Moore, C.A., Milano, S.K. & Benovic, J.L. Regulation of receptor trafficking by GRKs and arrestins. Annu. Rev. Physiol. 69, 451–482 (2007).
DeWire, S.M., Ahn, S., Lefkowitz, R.J. & Shenoy, S.K. Beta-arrestins and cell signaling. Annu. Rev. Physiol. 69, 483–510 (2007).
Yarden, Y. & Sliwkowski, M.X. Untangling the ErbB signalling network. Nat. Rev. Mol. Cell Biol. 2, 127–137 (2001).
Prickett, T.D. et al. Analysis of the tyrosine kinome in melanoma reveals recurrent mutations in ERBB4. Nat. Genet. 41, 1127–1132 (2009).
da Cunha Santos, G., Shepherd, F.A. & Tsao, M.S. EGFR mutations and lung cancer. Annu. Rev. Pathol. 6, 49–69 (2011).
Pines, G., Kostler, W.J. & Yarden, Y. Oncogenic mutant forms of EGFR: lessons in signal transduction and targets for cancer therapy. FEBS Lett. 584, 2699–2706 (2010).
Carey, K.D. et al. Kinetic analysis of epidermal growth factor receptor somatic mutant proteins shows increased sensitivity to the epidermal growth factor receptor tyrosine kinase inhibitor, erlotinib. Cancer Res. 66, 8163–8171 (2006).
Shortt, J. & Johnstone, R.W. Oncogenes in cell survival and cell death. Cold Spring Harb. Perspect. Biol. 4, a009829 (2012).
Ahn, R. et al. The ShcA PTB domain functions as a biological sensor of phosphotyrosine signaling during breast cancer progression. Cancer Res. 73, 4521–4532 (2013).
Bisson, N. et al. Selected reaction monitoring mass spectrometry reveals the dynamics of signaling through the GRB2 adaptor. Nat. Biotechnol. 29, 653–658 (2011).
Barrios-Rodiles, M. et al. High-throughput mapping of a dynamic signaling network in mammalian cells. Science 307, 1621–1625 (2005).
Taipale, M. et al. Quantitative analysis of HSP90-client interactions reveals principles of substrate recognition. Cell 150, 987–1001 (2012).
Blakely, K., Ketela, T. & Moffat, J. Pooled lentiviral shRNA screening for functional genomics in mammalian cells. Methods Mol. Biol. 781, 161–182 (2011).
Sriram, G. & Birge, R.B. Emerging roles for crk in human cancer. Genes Cancer 1, 1132–1139 (2010).
Cheung, H.W. et al. Amplification of CRKL induces transformation and epidermal growth factor receptor inhibitor resistance in human non-small cell lung cancers. Cancer Discov. 1, 608–625 (2011).
Kosaka, T., Yamaki, E., Mogi, A. & Kuwano, H. Mechanisms of resistance to EGFR TKIs and development of a new generation of drugs in non-small-cell lung cancer. J. Biomed. Biotechnol. 2011, 165214 (2011).
Li, J. et al. Perturbation of the mutated EGFR interactome identifies vulnerabilities and resistance mechanisms. Mol. Syst. Biol. 9, 705 (2013).
Yang, X. et al. A public genome-scale lentiviral expression library of human ORFs. Nat. Methods 8, 659–661 (2011).
Lemercier, C. et al. mHDA1/HDAC5 histone deacetylase interacts with and represses MEF2A transcriptional activity. J. Biol. Chem. 275, 15594–15599 (2000).
Kurat, C.F. et al. Restriction of histone gene transcription to S phase by phosphorylation of a chromatin boundary protein. Genes Dev. 25, 2489–2501 (2011).
Brown, K.R. & Jurisica, I. Unequal evolutionary conservation of human protein interactions in interologous networks. Genome Biol. 8, R95 (2007).
Zhu, C.Q. et al. Understanding prognostic gene expression signatures in lung cancer. Clin. Lung Cancer 10, 331–340 (2009).
Marcotte, R. et al. Essential gene profiles in breast, pancreatic, and ovarian cancer cells. Cancer Discov 2, 172–189 (2012).
Pinheiro, J.C. & Bates, D.M. Mixed-effects models in S and S-PLUS (Springer, 2000).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Chung, Y., Rabe-Hesketh, S., Dorie, V., Gelman, A. & Liu, J. A nondegenerate penalized likelihood estimator for variance parameters in multilevel models. Psychometrika 78, 685–709 (2013).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Acknowledgements
We thank B. Andrews for providing access to equipment and C. Nislow (University of British Columbia) for providing cDNA clones, J. Moffat (University of Toronto) for providing lentiviral plasmids, M.-S. Tsao (Princess Margaret Cancer Centre) for providing erlotinib and B. Neel (Princess Margaret Cancer Centre) for providing imatinib; S. Angers, K. Blakely, J. Jin, S. Kittanakom, A. Arnoldo, M. Mendoza and Y. Fedyshyn for experimental assistance and advice; and U. Petrovic, M. Ali, M. Lam, Z. Yao, K. Sokolina for advice and/or critical review of the manuscript. This work was supported by grants from the Ontario Genomics Institute (303547), Canadian Institutes of Health Research (Catalyst-NHG94491, PPP-125785), Canadian Foundation for Innovation (IOF-LOF), Natural Sciences and Engineering Research Council of Canada (RGPIN 372393-12), Canadian Cystic Fibrosis Foundation (300348), Canadian Cancer Society (2010-700406), Novartis and University Health Network (GL2-01-018) to I.S.; J.P. was funded by a Fonds zur Förderung der wissenschaftlichen Forschung–Erwin Schrödinger postdoctoral fellowship.
Author information
Authors and Affiliations
Contributions
I.S. conceived the project, provided guidance and assisted in manuscript preparation. J.P. coordinated, managed and was actively involved in all experiments, and wrote the bulk of the manuscript. B.G. was actively involved in the design and performance of the functional and screening part of MaMTH. M.K. and I.J. provided bioinformatics predictions and analysis of EGFR interactors. M.T., I.K. and S.L. were responsible for LUMIER assays. Y.Z. and T.P. generated co-IP and MS data. C.F.K. performed knockdown confirmations and provided technical help for all experiments. A.S., J.R.S., M.-S.T. and J.M. generated and analyzed shRNA knockdown data. J.S. was actively involved in manuscript preparation and provided project guidance. M.M.U. and A.N. were involved in data analysis, bait and prey generation, and fluorescence microscopy. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
A provisional patent application was filed by the University of Toronto on 2 July 2013 to the US Patent and Trademark Office, application 61/833,304. The intellectual property has been assigned to the University of Toronto and its commercialization is being managed by the Innovations and Partnership Office.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15, Supplementary Tables 1, 4, and 6–10 (PDF 15809 kb)
Supplementary Table 2
Previously identified EGFR interactors plus PubMed-IDs and number of preys identified in various publications. (XLSX 20 kb)
Supplementary Table 3
EGFR screening luciferase data and unrelated bait interaction data. (XLSX 73 kb)
Supplementary Table 5
Previously identified EGFR interactors not detected by MaMTH. (XLSX 10 kb)
Rights and permissions
About this article
Cite this article
Petschnigg, J., Groisman, B., Kotlyar, M. et al. The mammalian-membrane two-hybrid assay (MaMTH) for probing membrane-protein interactions in human cells. Nat Methods 11, 585–592 (2014). https://doi.org/10.1038/nmeth.2895
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2895
This article is cited by
-
Imaging strategies for receptor tyrosine kinase dimers in living cells
Analytical and Bioanalytical Chemistry (2023)
-
A method for Boolean analysis of protein interactions at a molecular level
Nature Communications (2022)
-
Prevalence of IGFBP3, NOS3 and TCF7L2 polymorphisms and their association with hypertension: a population-based study with Brazilian women of African descent
BMC Research Notes (2021)
-
Receptor tyrosine kinases and cancer: oncogenic mechanisms and therapeutic approaches
Oncogene (2021)
-
Application of the conventional and novel methods in testing EGFR variants for NSCLC patients in the last 10 years through different regions: a systematic review
Molecular Biology Reports (2021)