Abstract
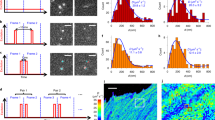
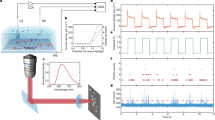
Biomolecular interactions are generally accompanied by modifications in size and charge of biomolecules at the nanometer scale. Here we describe a single-molecule method to sense these changes in real time based on statistical learning of diffusive and electric field–induced motion parameters of a trapped molecule in solution. We demonstrate the approach by resolving a monomer-trimer mixture along a protein dissociation pathway and visualizing the binding-unbinding kinetics of a single DNA molecule.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
van Oijen, A.M. Curr. Opin. Biotechnol. 22, 75–80 (2011).
Cisse, I.I., Kim, H. & Ha, T. Nat. Struct. Mol. Biol. 19, 623–627 (2012).
Jungmann, R. et al. Nano Lett. 10, 4756–4761 (2010).
Ulbrich, M.H. & Isacoff, E.Y. Nat. Methods 4, 319–321 (2007).
Ha, T. & Tinnefeld, P. Annu. Rev. Phys. Chem. 63, 595–617 (2012).
Magde, D., Webb, W.W. & Elson, E.L. Biopolymers 17, 361–367 (1978).
Muller, C.B. et al. Eur. Phys. Lett. 83, 46001 (2008).
Cohen, A.E. & Moerner, W.E. Proc. Natl. Acad. Sci. USA 103, 4362–4365 (2006).
Wang, Q. & Moerner, W.E. ACS Nano 5, 5792–5799 (2011).
Fields, A.P. & Cohen, A.E. Proc. Natl. Acad. Sci. USA 108, 8937–8942 (2011).
Wang, Q. & Moerner, W.E. J. Phys. Chem. B 117, 4641–4648 (2012).
Enderlein, J. Appl. Phys. B 71, 773–777 (2000).
Qi, Y. Extending Expectation Propagation for Graphical Models. PhD Thesis, Massachusetts Institute of Technology (2004).
Shumway, R.H. & Stoffer, D.S. J. Time Ser. Anal. 3, 253–264 (1982).
Huang, C., Berns, D.S. & MacColl, R. Biochemistry 26, 243–245 (1987).
Cohen-Bazire, G., Beguin, S., Rimon, S., Glazer, A.N. & Brown, D.M. Arch. Microbiol. 111, 225–238 (1977).
Markham, N.R. & Zuker, M. Nucleic Acids Res. 33, W577–W581 (2005).
Goldsmith, R.H. & Moerner, W.E. Nat. Chem. 2, 179–186 (2010).
Holzmeister, P., Acuna, G.P., Grohmann, D. & Tinnefeld, P. Chem. Soc. Rev. 10.1039/C3CS60207A (10 September 2013).
Tyagi, S. et al. Nat. Methods 10.1038/nmeth.2809 (19 January 2014).
Cohen, A.E. & Moerner, W.E. Opt. Express 16, 6941–6956 (2008).
Kartalov, E., Unger, M. & Quake, S.R. BioTechniques 34, 505–510 (2003).
Horvath, J. & Dolnik, V. Electrophoresis 22, 644–655 (2001).
Jordan, M.I. Stat. Sci. 19, 140–155 (2004).
Qi, Y. & Minka, T.P. IEEE Trans. Wirel. Comm. 6, 348–355 (2007).
Botev, Z.I., Grotowski, J.F. & Kroese, D.P. Ann. Stat. 38, 2916–2957 (2010).
Murphy, K.P. Machine Learning: a Probabilistic Perspective (The MIT Press, 2012).
Rasnik, I., McKinney, S.A. & Ha, T. Nat. Methods 3, 891–893 (2006).
Aitken, C.E., Marshall, R.A. & Puglisi, J.D. Biophys. J. 94, 1826–1835 (2008).
Acknowledgements
We thank Y. Jiang for help with high-performance liquid chromatography purification, and C. Calderon, G. Schlau-Cohen, H.-Y. Yang, S. Bockenhauer and S.J. Sahl for discussion. This work is funded in part by the Division of Chemical Sciences, Geosciences and Biosciences, Office of Basic Energy Sciences of the US Department of Energy through grant DE-FG02-07ER15892.
Author information
Authors and Affiliations
Contributions
Q.W. and W.E.M. conceived the project, discussed the results and wrote the manuscript. Q.W. designed and performed the experiments and data analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–19, Supplementary Tables 1–5 and Supplementary Notes 1–7 (PDF 7165 kb)
Real-time estimation of single-molecule transport coefficients in an anti-Brownian electrokinetic trap.
Screen recording during an experiment trapping single Alexa647 labeled 10nt-ssDNA molecules without complementary strand. Intensity (I) is photon counts every 10 ms, diffusion coefficient (D) and electrokinetic mobility (μx and μy) are estimated every 5,000 photons using the real-time EM algorithm. Occasional intensity spikes are indicative of transient co-occupancy of two objects in the trap. Time axis units: 10 ms (AVI 4611 kb)
Real-time visualization of single-DNA binding-unbinding dynamics.
Screen recording during an experiment trapping single 10nt-ssDNA in presence of unlabeled complementary strand (2 μM of 24nt-10comp). Frequent anti-correlated dynamics in diffusion coefficient (D) and electrokinetic mobility (μx and μy) visualize transitions between ssDNA (blue band) and dsDNA (red band). Time axis units: 10 ms (AVI 10418 kb)
Supplementary Software
C program implementation of the parameter estimation algorithm. (TXT 17 kb)
Rights and permissions
About this article
Cite this article
Wang, Q., Moerner, W. Single-molecule motions enable direct visualization of biomolecular interactions in solution. Nat Methods 11, 555–558 (2014). https://doi.org/10.1038/nmeth.2882
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2882
This article is cited by
-
ABEL-FRET: tether-free single-molecule FRET with hydrodynamic profiling
Nature Methods (2021)
-
Optical imaging of single-protein size, charge, mobility, and binding
Nature Communications (2020)
-
Recent advances in point spread function engineering and related computational microscopy approaches: from one viewpoint
Biophysical Reviews (2020)
-
Axial electrokinetic trapping of anisotropic particles
Scientific Reports (2019)
-
In situ monitoring of molecular aggregation using circular dichroism
Nature Communications (2018)