Abstract
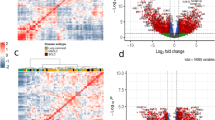
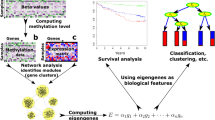
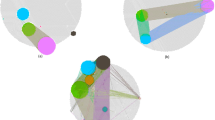
Recent technologies have made it cost-effective to collect diverse types of genome-wide data. Computational methods are needed to combine these data to create a comprehensive view of a given disease or a biological process. Similarity network fusion (SNF) solves this problem by constructing networks of samples (e.g., patients) for each available data type and then efficiently fusing these into one network that represents the full spectrum of underlying data. For example, to create a comprehensive view of a disease given a cohort of patients, SNF computes and fuses patient similarity networks obtained from each of their data types separately, taking advantage of the complementarity in the data. We used SNF to combine mRNA expression, DNA methylation and microRNA (miRNA) expression data for five cancer data sets. SNF substantially outperforms single data type analysis and established integrative approaches when identifying cancer subtypes and is effective for predicting survival.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Monti, S., Tamayo, P., Mesirov, J. & Golub, T. Consensus clustering: a resampling-based method for class discovery and visualization of gene expression microarray data. Mach. Learn. 52, 91–118 (2003).
Verhaak, R.G.W. et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 17, 98–110 (2010).
Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012).
Kirk, P., Griffin, J.E., Savage, R.S., Ghahramani, Z. & Wild, D.L. Bayesian correlated clustering to integrate multiple datasets. Bioinformatics 28, 3290–3297 (2012).
Cancer Genome Atlas Research Network. Comprehensive genomic characterization of squamous cell lung cancers. Nature 489, 519–525 (2012).
Cancer Genome Atlas Network. Comprehensive molecular characterization of human colon and rectal cancer. Nature 487, 330–337 (2012).
Shen, R., Olshen, A.B. & Ladanyi, M. Integrative clustering of multiple genomic data types using a joint latent variable model with application to breast and lung cancer subtype analysis. Bioinformatics 25, 2906–2912 (2009).
Goldenberg, A., Zheng, A.X., Fienberg, S.E. & Airoldi, E.M. A survey of statistical network models. Foundations and Trends in Machine Learning. 2, 129–233 (2010).
Barabási, A.-L. Network medicine -from obesity to the 'diseasome. N. Engl. J. Med. 357, 404–407 (2007).
Pearl, J. Probabilistic Reasoning in Intelligent Systems: Networks of Plausible Inference (Morgan Kaufmann, 1988).
Nigro, J.M. et al. Integrated array-comparative genomic hybridization and expression array profiles identify clinically relevant molecular subtypes of glioblastoma. Cancer Res. 65, 1678–1686 (2005).
Sturm, D. et al. Hotspot mutations in H3F3A and IDH1 define distinct epigenetic and biological subgroups of glioblastoma. Cancer Cell 22, 425–437 (2012).
Sun, S. et al. Protein alterations associated with temozolomide resistance in subclones of human glioblastoma cell lines. J. Neurooncol. 107, 89–100 (2012).
Hosmer Jr, D.W., Lemeshow, S. & May, S. Applied Survival Analysis: Regression Modeling of Time to Event Data (Wiley, 2011).
Rousseeuw, P. Silhouettes: a graphical aid to the interpretation and validation of cluster analysis. J. Comput. Appl. Math. 20, 53–65 (1987).
Tusher, V.G., Tibshirani, R. & Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proc. Natl. Acad. Sci. USA 98, 5116–5121 (2001).
Curtis, C. et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486, 346–352 (2012).
Margolin, A.A. et al. Systematic analysis of challenge-driven improvements in molecular prognostic models for breast cancer. Sci. Transl. Med. 5, 181 (2013).
Friend, S.H. & Ideker, T. Point: Are we prepared for the future doctor visit? Nat. Biotechnol. 29, 215–218 (2011).
Wang, K. et al. PennCNV: an integrated hidden Markov model designed for high-resolution copy number variation detection in whole-genome SNP genotyping data. Genome Res. 17, 1665–1674 (2007).
Troyanskaya, O. et al. Missing value estimation methods for DNA microarrays. Bioinformatics 17, 520–525 (2001).
Wang, B., Jiang, J., Wang, W., Zhou, Z.-H. & Tu, Z. Unsupervised metric fusion by cross diffusion. in Proc. IEEE Comput. Soc. Conf. Comput. Vis. Pattern Recognit. 2997–3004 (IEEE, 2012).
Ng, A.Y., Jordan, M.I. & Weiss, Y. On spectral clustering: analysis and an algorithm. Adv. Neural Inf. Process. Syst. 2, 849–856 (2002).
Wei, Y.C. & Cheng, C.K. Towards efficient hierarchical designs by ratio cut partitioning. in Proc. Int. Conf. Computer-Aided Design 298–301 (ICCAD, 1989).
Luxburg, U. A tutorial on spectral clustering. Stat. Comput. 17, 395–416 (2007).
Zhang, W. et al. Network-based survival analysis reveals subnetwork signatures for predicting outcomes of ovarian cancer treatment. PLoS Comput. Biol. 9, e1002975 (2013).
Acknowledgements
This study used data generated by TCGA and METABRIC; we thank TCGA, the Cancer Research UK and the British Columbia Cancer Agency Branch for sharing these invaluable data with the scientific community. We thank N. Jabado, M. Wilson and J. Rommens for feedback on the manuscript, and B. Sousa for help with the figures. This study was partially funded by the Government of Canada through Genome Canada and the Ontario Genomics Institute (OGI-068) to M.B.; A.G. is funded by the SickKids Research Institute. Z.T. was supported by NSF IIS-1360568.
Author information
Authors and Affiliations
Contributions
B.W. and A.G. conceived of and designed the approach. B.W. performed the data analysis, implemented the method in Matlab and performed all computational experiments. A.M.M. performed data preparation. F.D. wrote the R code that is distributed with the paper. M.F. assisted with network visualization and analysis. Z.T. helped with method design and theoretical framework. B.H.-K. assisted in preparation and analysis of the METABRIC data. B.W., M.B. and A.G. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–20, Supplementary Table 1, Supplementary Notes 1 –3 and Supplementary Results (PDF 6804 kb)
Supplementary Software
Similarity Network Fusion for aggregating multiple data types (ZIP 415 kb)
Supplementary Data
TCGA cancer datasets after pre-processing (ZIP 81276 kb)
Rights and permissions
About this article
Cite this article
Wang, B., Mezlini, A., Demir, F. et al. Similarity network fusion for aggregating data types on a genomic scale. Nat Methods 11, 333–337 (2014). https://doi.org/10.1038/nmeth.2810
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2810
This article is cited by
-
Multi-omics integration with weighted affinity and self-diffusion applied for cancer subtypes identification
Journal of Translational Medicine (2024)
-
A semi-supervised approach for the integration of multi-omics data based on transformer multi-head self-attention mechanism and graph convolutional networks
BMC Genomics (2024)
-
wMKL: multi-omics data integration enables novel cancer subtype identification via weight-boosted multi-kernel learning
British Journal of Cancer (2024)
-
Dynamic network curvature analysis of gene expression reveals novel potential therapeutic targets in sarcoma
Scientific Reports (2024)
-
Mononuclear phagocyte system-related multi-omics features yield head and neck squamous cell carcinoma subtypes with distinct overall survival, drug, and immunotherapy responses
Journal of Cancer Research and Clinical Oncology (2024)