The P value reported by tests is a probabilistic significance, not a biological one.
Bench scientists often perform statistical tests to determine whether an observation is statistically significant. Many tests report the P value to measure the strength of the evidence that a result is not just a likely chance occurrence. To make informed judgments about the observations in a biological context, we must understand what the P value is telling us and how to interpret it. This month we will develop the concept of statistical significance and tests by introducing the one-sample t-test.
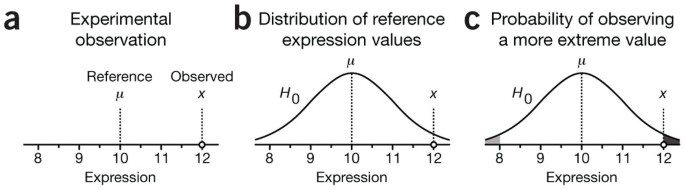
To help you understand how statistical testing works, consider the experimental scenario depicted in Figure 1 of measuring protein expression level in a cell line with a western blot. Suppose we measure an expression value of x = 12 and have good reason to believe (for example, from past measurements) that the reference level is μ = 10 (Fig. 1a). What can we say about whether this difference is due to random chance? Statistical testing can answer this question. But first, we need to mathematically frame our intuitive understanding of the biological and technical factors that disperse our measurements across a range of values.
(a–c) The significance of the difference between observed (x) and reference (μ) values (a) is calculated by assuming that observations are sampled from a distribution H0 with mean μ (b). The statistical significance of the observation x is the probability of sampling a value from the distribution that is at least as far from the reference, given by the shaded areas under the distribution curve (c). This is the P value.
We begin with the assumption that the random fluctuations in the experiment can be characterized by a distribution (Fig. 1b). This distribution is called the null distribution, and it embodies the null hypothesis (H0) that our observation is a sample from the pool of all possible instances of measuring the reference. We can think of constructing this distribution by making a large number of independent measurements of a protein whose mean expression is known to equal the reference value. This distribution represents the probability of observing a given expression level for a protein that is being expressed at the reference level. The mean of this distribution, μ, is the reference expression, and its spread is determined by reproducibility factors inherent to our experiment. The purpose of a statistical test is to locate our observation on this distribution to identify the extent to which it is an outlier.
Statistics quantifies the outlier status of an observation by the probability of sampling another observation from the null distribution that is as far or farther away from μ. In our example, this corresponds to measuring an expression value further from the reference than x. This probability is the P value, which is the output of common statistical tests. It is calculated from the area under the distribution curve in the shaded regions (Fig. 1c). In some situations we may care only if x is too big (or too small), in which case we would compute the area of only the dark (light) shaded region of Figure 1c.
Unfortunately, the P value is often misinterpreted as the probability that the null hypothesis (H0) is true. This mistake is called the 'prosecutor's fallacy', which appeals to our intuition and was so coined because of its frequent use in courtroom arguments. In the process of calculating the P value, we assumed that H0 was true and that x was drawn from H0. Thus, a small P value (for example, P = 0.05) merely tells us that an improbable event has occurred in the context of this assumption. The degree of improbability is evidence against H0 and supports the alternative hypothesis that the sample actually comes from a population whose mean is different than μ. Statistical significance suggests but does not imply biological significance.
At this point you may ask how we arrive at our assumptions about the null distribution in Figure 1b. After all, in order to calculate P, we need to know its precise shape. Because experimentally determining it is not practical, we need to make an informed guess. For the purposes of this column, we will assume that it is normal. We will discuss robustness of tests to this assumption of normality in another column. To complete our model of H0, we still need to estimate its spread. To do this we return to the concept of sampling.
To estimate the spread of H0, we repeat the measurement of our protein's expression. For example, we might make four additional independent measurements to make up a sample with n = 5 (Fig. 2a). We use the mean of expression values (\(\overline{x}\) = 10.85) as a measure of our protein's expression. Next, we make the key assumption that the s.d. of our sample (sx = 0.96) is a suitable estimate of the s.d. of the null distribution (Fig. 2b). In other words, regardless of whether the sample mean is representative of the null distribution, we assume that its spread is. This assumption of equal variances is common, and we will be returning to it in future columns.
(a) A sample of n = 5 observations is taken and characterized by the mean x̄, with error bars showing s.d. (sx) and s.e.m. (sx/√n). (b) The null distribution is assumed to be normal, and its s.d. is estimated by sx. As in Figure 1b, the population mean is assumed to be μ. (c) The average expression is located on the sampling distribution of sample means, whose spread is estimated by the s.e.m. and whose mean is also μ. The P value of x̄ is the shaded area under this curve.
From our discussion about sampling1, we know that given that H0 is normal, the sampling distribution of means will also be normal, and we can use sx/√n to estimate its s.d. (Fig. 2c). We localize the mean expression on this distribution to calculate the P value, analogously to what was done with the single value in Figure 1c. To avoid the nuisance of dealing with a sampling distribution of means for each combination of population parameters, we can transform the mean x̄ to a value determined by the difference of the sample and population means D = x̄ − μ divided by the s.e.m. (sx/√n). This is called the test statistic.
It turns out, however, that the shape of this sampling distribution is close to, but not exactly, normal. The extent to which it departs from normal is known and given by the Student's t distribution (Fig. 3a), first described by William Gosset, who published under the pseudonym 'Student' (to avoid difficulties with his employer, Guinness) in his work on optimizing barley yields. The test statistic described above is compared to this distribution and is thus called the t statistic. The test illustrated in Figure 2 is called the one-sample t-test.
(a) The t distribution has higher tails that take into account that most samples will underestimate the variability in a population. The distribution is used to evaluate the significance of a t statistic derived from a sample of size n and is characterized by the degrees of freedom, d.f. = n − 1. (b) When n is small, P values derived from the t distribution vary greatly as n changes.
This departure in distribution shape is due to the fact that for most samples, the sample variance, sx2, is an underestimate of the variance of the null distribution. The distribution of sample variances turns out to be skewed. The asymmetry is more evident for small n, where it is more likely that we observe a variance smaller than that of the population. The t distribution accounts for this underestimation by having higher tails than the normal distribution (Fig. 3a). As n grows, the t distribution looks very much like the normal, reflecting that the sample's variance becomes a more accurate estimate.
As a result, if we do not correct for this—if we use the normal distribution in the calculation depicted in Figure 2c—we will be using a distribution that is too narrow and will overestimate the significance of our finding. For example, using the n = 5 sample in Figure 2b for which t = 1.98, the t distribution gives us P = 0.119. Without the correction built into this distribution, we would underestimate P using the normal distribution as P = 0.048 (Fig. 3b). When n is large, the required correction is smaller: the same t = 1.98 for n = 50 gives P = 0.054, which is now much closer to the value obtained from the normal distribution.
The relationship between t and P is shown in Figure 3b and can be used to express P as a function of the quantities on which t depends (D, sx, n). For example, if our sample in Figure 2b had a size of at least n = 8, the observed expression difference D = 0.85 would be significant at P < 0.05, assuming we still measured sx = 0.96 (t = 2.50, P = 0.041). A more general type of calculation can identify conditions for which a test can reliably detect whether a sample comes from a distribution with a different mean. This speaks to the test's power, which we will discuss in the next column.
Another way of thinking about reaching significance is to consider what population means would yield P < 0.05. For our example, these would be μ < 9.66 and μ > 12.04 and define the range of standard expression values (9.66–12.04) that are compatible with our sample. In other words, if the null distribution had a mean within this interval, we would not be able to reject H0 at P = 0.05 on the basis of our sample. This is the 95% confidence interval introduced last month, given by μ = x̄ ± t* × s.e.m. (a rearranged form of the one-sample t-test equation), where t* is the critical value of the t statistic for a given n and P. In our example, n = 5, P = 0.05 and t* = 2.78. We encourage readers to explore these concepts for themselves using the interactive graphs in Supplementary Table 1.
The one-sample t-test is used to determine whether our samples could come from a distribution with a given mean (for example, to compare the sample mean to a putative fixed value m) and for constructing confidence intervals for the mean. It appears in many contexts, such as measuring protein expression, the quantity of drug delivered by a medication or the weight of cereal in your cereal box. The concepts underlying this test are an important foundation for future columns in which we will discuss the comparisons across samples that are ubiquitous in the scientific literature.
References
Krzywinski, M. & Altman, N. Nat. Methods 10, 809–810 (2013).
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Table 1
Worksheets illustrate calculations that support statements in the column or provide interactive versions of its figures. Please note that the workbook requires that macros be enabled. (XLSM 477 kb)
Rights and permissions
About this article
Cite this article
Krzywinski, M., Altman, N. Significance, P values and t-tests. Nat Methods 10, 1041–1042 (2013). https://doi.org/10.1038/nmeth.2698
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2698
This article is cited by
-
Use of the p-values as a size-dependent function to address practical differences when analyzing large datasets
Scientific Reports (2021)
-
Investigation of phytoconstituents of Enicostemma littorale as potential glucokinase activators through molecular docking for the treatment of type 2 diabetes mellitus
In Silico Pharmacology (2021)
-
Association between microstructural white matter abnormalities and cognitive functioning in patients with type 2 diabetes mellitus: a diffusion tensor imaging study
The Egyptian Journal of Neurology, Psychiatry and Neurosurgery (2020)
-
Increasing high school teachers self-efficacy for integrated STEM instruction through a collaborative community of practice
International Journal of STEM Education (2020)
-
Tutorial: guidance for quantitative confocal microscopy
Nature Protocols (2020)