Abstract
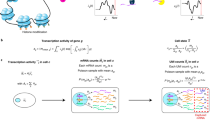
Single-cell RNA-seq can yield valuable insights about the variability within a population of seemingly homogeneous cells. We developed a quantitative statistical method to distinguish true biological variability from the high levels of technical noise in single-cell experiments. Our approach quantifies the statistical significance of observed cell-to-cell variability in expression strength on a gene-by-gene basis. We validate our approach using two independent data sets from Arabidopsis thaliana and Mus musculus.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Change history
11 October 2013
In the version of this article initially published online, the dilution for the ERCC Spike-In Control Mix added to the lysis mix was given as 1:40,000 in the Online Methods. The actual dilution used was 1:400. The error has been corrected for the PDF and HTML versions of this article.
References
Hashimshony, T., Wagner, F., Sher, N. & Yanai, I. Cell Rep. 2, 666–673 (2012).
Islam, S. et al. Genome Res. 21, 1160–1167 (2011).
Ramsköld, D. et al. Nat. Biotechnol. 30, 777–782 (2012).
Tang, F. et al. Nat. Protoc. 5, 516–535 (2010).
Tang, F. et al. Nat. Methods 6, 377–382 (2009).
Chambers, I. et al. Nature 450, 1230–1234 (2007).
Reynolds, N. et al. Cell Stem Cell 10, 583–594 (2012).
Chang, H.H., Hemberg, M., Barahona, M., Ingber, D.E. & Huang, S. Nature 453, 544–547 (2008).
Toyooka, Y., Shimosato, D., Murakami, K., Takahashi, K. & Niwa, H. Development 135, 909–918 (2008).
Shalek, A.K. et al. Nature 498, 236–240 (2013).
Marioni, J.C., Mason, C.E., Mane, S.M., Stephens, M. & Gilad, Y. Genome Res. 18, 1509–1517 (2008).
Brady, S.M. et al. Science 318, 801–806 (2007).
Jiang, L. et al. Genome Res. 21, 1543–1551 (2011).
Benjamini, Y. & Hochberg, Y. Stat. Soc. Series B Stat. Methodol. 57, 289–300 (1995).
Clough, S.J. & Bent, A.F. Plant J. 16, 735–743 (1998).
Birnbaum, K. et al. Nat. Methods 2, 615–619 (2005).
Wu, T.D. & Nacu, S. Bioinformatics 26, 873–881 (2010).
Irizarry, R.A. et al. Biostatistics 4, 249–264 (2003).
Anders, S. & Huber, W. Genome Biol. 11, R106 (2010).
Alexa, A., Rahnenfuhrer, J. & Lengauer, T. Bioinformatics 22, 1600–1607 (2006).
Acknowledgements
We thank E. Furlong and W. Huber for helpful discussions. We also acknowledge K. Birnbaum (New York University) for kindly providing pWOX5::GFP and pGl2::GFP seed. S.A. acknowledges partial funding from the European Union (FP7-Health, project Radiant); M.G.H. acknowledges the Australian Research Council for present funding. The EMBL Genomics Core Facility provided technical support for this work. We acknowledge A. Surani for the use of the C1 Single-Cell Auto Prep System in his lab and B. Jones for performing the experiment. We also acknowledge A. McKenzie (Medical Research Council Laboratory of Molecular Biology) for the Il13-GFP reporter mice and the Sanger-EBI Single Cell Centre for technical support. We acknowledge the support of European Research Council Starting Grant no. 260507, ThSWITCH.
Author information
Authors and Affiliations
Contributions
P.B. designed plant cell experiments, carried out experiments, interpreted results and wrote the paper; S.A. developed the statistical method, performed bioinformatics analyses and wrote the paper; J.K.K. performed bioinformatics analyses and helped write the paper; A.A.K. designed and carried out mouse cell experiments and helped write the paper; X.Z. designed and analyzed mouse cell experiments and helped write the paper; V.P. designed and carried out mouse cell experiments and helped write the paper; B.B. adapted an Illumina sequencing library preparation protocol; V.B. contributed to adapting the Illumina sequencing library preparation protocol and gave advice; S.A.T. designed mouse cell experiments and helped write the paper; J.C.M. contributed to the development of the statistical method, performed bioinformatics analyses, supervised the project and wrote the paper; M.G.H. initiated the project, designed plant cell experiments, interpreted results, supervised the project and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14 and Supplementary Notes 1–9 (PDF 8682 kb)
Supplementary Software
Accounting for technical noise in single-cell RNA-seq experiments – Supplement II. This file contains the R code used to perform the analysis described in the manuscript. (PDF 1938 kb)
Supplementary Table 1
Estimated amounts of total RNA in single plant cells. Content of total RNA from single cells was estimated based on the 50 pg HeLa total RNA spike-in. The calculation is based on the assumption that the fraction of polyadenylated RNA is comparable between HeLa and A. thaliana input material. For detailed description refer to the Methods section. (XLSX 8 kb)
Supplementary Table 2
List of highly variable genes in GL2 cells. The columns give the gene ID, the gene name, the normalized average read count, the cell with the strongest expression, and, for each cell, the log2 ratio of the cell's expression to the average. (XLSX 134 kb)
Supplementary Table 3
List of highly variable genes in QC cells. Columns are the same as in Supplementary Table 2. (XLSX 16 kb)
Supplementary Table 4
List of GO categories that are significantly enriched for highly variable genes (Online Methods). (XLSX 10 kb)
Supplementary Table 5
Read counts for the 91 mouse immune cells spiked with ERCC spike-ins. Each column corresponds to a cell, each row to gene or an ERCC spike-in molecule. Mouse gene names have been replaces by randomized identifiers (column 1). The second column contains the transcript lengths used for the analysis in Supplementary Note 5. The transcript lengths are computed from Ensembl annotation by taking the union of all exons within a gene, where the exons annotated as “retained introns” and “nonsense mediated decay” are excluded. (XLSX 14402 kb)
Supplementary Table 6
Full list of barcoded Illumina PE adapters used for multiplexing of cDNA libraries. The set of adapters used depends on the degree of multiplexing applied to the samples. 4-plex is the lower limit for multiplexing. (XLSX 10 kb)
Supplementary Table 7
List of qPCR primers used (XLSX 8 kb)
Supplementary Table 8
Read counts for A. thaliana experiments. Raw number of reads mapped to each gene for all samples. (XLSX 6101 kb)
Supplementary Table 9
Transcript lengths for the human genome used for the analysis described in Supplementary Note 5, computed as described above, in the legend for Supplementary Table 5. (XLSX 1227 kb)
Rights and permissions
About this article
Cite this article
Brennecke, P., Anders, S., Kim, J. et al. Accounting for technical noise in single-cell RNA-seq experiments. Nat Methods 10, 1093–1095 (2013). https://doi.org/10.1038/nmeth.2645
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2645
This article is cited by
-
Next-Generation Sequencing in Medicinal Plants: Recent Progress, Opportunities, and Challenges
Journal of Plant Growth Regulation (2024)
-
Acute stress reduces population-level metabolic and proteomic variation
BMC Bioinformatics (2023)
-
Heritable transcriptional defects from aberrations of nuclear architecture
Nature (2023)
-
Distortion-free PCA on sample space for highly variable gene detection from single-cell RNA-seq data
Frontiers of Computer Science (2023)
-
RNA-Seq analysis of duck embryo fibroblast cells gene expression during duck Tembusu virus infection
Veterinary Research (2022)