Abstract
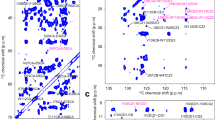
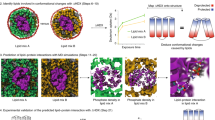
Determination of structure of integral membrane proteins, especially in their native environment, is a formidable challenge in structural biology. Here we demonstrate that magic angle spinning solid-state NMR spectroscopy can be used to determine structures of membrane proteins reconstituted in synthetic lipids, an environment similar to the natural membrane. We combined a large number of experimentally determined interatomic distances and local torsional restraints to solve the structure of an oligomeric membrane protein of common seven-helical fold, Anabaena sensory rhodopsin (ASR). We determined the atomic resolution detail of the oligomerization interface of the ASR trimer, and the arrangement of helices, side chains and the retinal cofactor in the monomer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Stroud, R.M. New tools in membrane protein determination. F1000 Biol. Rep. 3, 8 (2011).
Bill, R.M. et al. Overcoming barriers to membrane protein structure determination. Nat. Biotechnol. 29, 335–340 (2011).
Palczewski, K. et al. Crystal structure of rhodopsin: a G protein-coupled receptor. Science 289, 739–745 (2000).
Cherezov, V. et al. High-resolution crystal structure of an engineered human beta2-adrenergic G protein–coupled receptor. Science 318, 1258–1265 (2007).
Rasmussen, S.G. et al. Crystal structure of the human beta2 adrenergic G-protein-coupled receptor. Nature 450, 383–387 (2007).
Jaakola, V.P. et al. The 2.6 angstrom crystal structure of a human A2A adenosine receptor bound to an antagonist. Science 322, 1211–1217 (2008).
Shimamura, T. et al. Structure of the human histamine H1 receptor complex with doxepin. Nature 475, 65–70 (2011).
Kim, H.J., Howell, S.C., Van Horn, W.D., Jeon, Y.H. & Sanders, C.R. Recent advances in the application of solution NMR spectroscopy to multi-span integral membrane proteins. Prog. Nucl. Magn. Reson. Spectrosc. 55, 335–360 (2009).
Hiller, S. & Wagner, G. The role of solution NMR in the structure determinations of VDAC-1 and other membrane proteins. Curr. Opin. Struct. Biol. 19, 396–401 (2009).
Nietlispach, D. & Gautier, A. Solution NMR studies of polytopic alpha-helical membrane proteins. Curr. Opin. Struct. Biol. 21, 497–508 (2011).
Zhou, Y. et al. NMR solution structure of the integral membrane enzyme DsbB: functional insights into DsbB-catalyzed disulfide bond formation. Mol. Cell 31, 896–908 (2008).
Van Horn, W.D. et al. Solution nuclear magnetic resonance structure of membrane-integral diacylglycerol kinase. Science 324, 1726–1729 (2009).
Gautier, A., Mott, H.R., Bostock, M.J., Kirkpatrick, J.P. & Nietlispach, D. Structure determination of the seven-helix transmembrane receptor sensory rhodopsin II by solution NMR spectroscopy. Nat. Struct. Mol. Biol. 17, 768–774 (2010).
Reckel, S. et al. Solution NMR structure of proteorhodopsin. Angew. Chem. Int. Edn. Engl. 50, 11942–11946 (2011).
Berardi, M.J., Shih, W.M., Harrison, S.C. & Chou, J.J. Mitochondrial uncoupling protein 2 structure determined by NMR molecular fragment searching. Nature 476, 109–113 (2011).
Lange, A. et al. Toxin-induced conformational changes in a potassium channel revealed by solid-state NMR. Nature 440, 959–962 (2006).
Ahuja, S. et al. Helix movement is coupled to displacement of the second extracellular loop in rhodopsin activation. Nat. Struct. Mol. Biol. 16, 168–175 (2009).
Sharma, M. et al. Insight into the mechanism of the influenza A proton channel from a structure in a lipid bilayer. Science 330, 509–512 (2010).
Cady, S.D. et al. Structure of the amantadine binding site of influenza M2 proton channels in lipid bilayers. Nature 463, 689–692 (2010).
Shahid, S.A. et al. Membrane-protein structure determination by solid-state NMR spectroscopy of microcrystals. Nat. Methods 9, 1212–1217 (2012).
Park, S.H. et al. Structure of the chemokine receptor CXCR1 in phospholipid bilayers. Nature 491, 779–783 (2012).
Manolikas, T., Herrmann, T. & Meier, B.H. Protein structure determination from 13C spin-diffusion solid-state NMR spectroscopy. J. Am. Chem. Soc. 130, 3959–3966 (2008).
Loquet, A. et al. 3D structure determination of the Crh protein from highly ambiguous solid-state NMR restraints. J. Am. Chem. Soc. 130, 3579–3589 (2008).
Jung, K.H., Trivedi, V.D. & Spudich, J.L. Demonstration of a sensory rhodopsin in eubacteria. Mol. Microbiol. 47, 1513–1522 (2003).
Wang, S., Kim, S.Y., Jung, K.H., Ladizhansky, V. & Brown, L.S. A eukaryotic-like interaction of soluble cyanobacterial sensory rhodopsin transducer with DNA. J. Mol. Biol. 411, 449–462 (2011).
Vogeley, L. et al. Anabaena sensory rhodopsin: a photochromic color sensor at 2.0 Å. Science 306, 1390–1393 (2004).
Shi, L., Kawamura, I., Jung, K.H., Brown, L.S. & Ladizhansky, V. Conformation of a seven-helical transmembrane photosensor in the lipid environment. Angew. Chem. Int. Edn. Engl. 50, 1302–1305 (2011).
Wang, S. et al. Paramagnetic relaxation enhancement reveals oligomerization interface of a membrane protein. J. Am. Chem. Soc. 134, 16995–16998 (2012).
Wang, S., Shi, L., Kawamura, I., Brown, L.S. & Ladizhansky, V. Site-specific solid-state NMR detection of hydrogen-deuterium exchange reveals conformational changes in a 7-helical transmembrane protein. Biophys. J. 101, L23–L25 (2011).
Peng, X., Libich, D., Janik, R., Harauz, G. & Ladizhansky, V. Dipolar chemical shift correlation spectroscopy for homonuclear carbon distance measurements in proteins in the solid state: application to structure determination and refinement. J. Am. Chem. Soc. 130, 359–369 (2008).
Lange, A., Luca, S. & Baldus, M. Structural constraints from proton-mediated rare-spin correlation spectroscopy in rotating solids. J. Am. Chem. Soc. 124, 9704–9705 (2002).
Castellani, F. et al. Structure of a protein determined by solid-state magic-angle-spinning NMR spectroscopy. Nature 420, 98–102 (2002).
Brunger, A.T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Bardiaux, B., Malliavin, T. & Nilges, M. ARIA for solution and solid-state NMR. Methods Mol. Biol. 831, 453–483 (2012).
Nadaud, P.S., Helmus, J.J., Hofer, N. & Jaroniec, C.P. Long-range structural restraints in spin-labeled proteins probed by solid-state nuclear magnetic resonance spectroscopy. J. Am. Chem. Soc. 129, 7502–7503 (2007).
Essen, L., Siegert, R., Lehmann, W.D. & Oesterhelt, D. Lipid patches in membrane protein oligomers: crystal structure of the bacteriorhodopsin-lipid complex. Proc. Natl. Acad. Sci. USA 95, 11673–11678 (1998).
Dencher, N.A. & Heyn, M.P. Formation and properties of bacteriorhodopsin monomers in the non-ionic detergents octyl-beta-D-glucoside and Triton X-100. FEBS Lett. 96, 322–326 (1978).
Shi, L. et al. Three-dimensional solid-state NMR study of a seven-helical integral membrane proton pump—structural insights. J. Mol. Biol. 386, 1078–1093 (2009).
Emami, S., Fan, Y., Munro, R., Ladizhansky, V. & Brown, L.S. Yeast-expressed human membrane protein aquaporin-1 yields excellent resolution of solid-state MAS NMR spectra. J. Biomol. NMR 55, 147–155 (2013).
Klammt, C. et al. Facile backbone structure determination of human membrane proteins by NMR spectroscopy. Nat. Methods 9, 834–839 (2012).
daCosta, C.J. & Baenziger, J.E. A rapid method for assessing lipid:protein and detergent:protein ratios in membrane-protein crystallization. Acta Crystallogr. D Biol. Crystallogr. 59, 77–83 (2003).
Morcombe, C.R. & Zilm, K.W. Chemical shift referencing in MAS solid state NMR. J. Magn. Reson. 162, 479–486 (2003).
Janik, R., Peng, X. & Ladizhansky, V. (13)C-(13)C distance measurements in U-(13)C, (15)N-labeled peptides using rotational resonance width experiment with a homogeneously broadened matching condition. J. Magn. Reson. 188, 129–140 (2007).
Delaglio, F. et al. NMRPipe—a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Keller, R. The Computer Aided Resonance Assignment Tutorial. 1st edn. (CANTINA Verlag Goldau, 2004).
Wang, S. et al. Solid-state NMR (13)C and (15)N resonance assignments of a seven-transmembrane helical protein Anabaena Sensory Rhodopsin. Biomol. NMR Assign. doi:10.1007/s12104-012-9421-y (16 September 2012).
Shen, Y., Delaglio, F., Cornilescu, G. & Bax, A. TALOS+: a hybrid method for predicting protein backbone torsion angles from NMR chemical shifts. J. Biomol. NMR 44, 213–223 (2009).
Tajkhorshid, E., Paizs, B. & Suhai, S. Conformational effects on the proton affinity of the Schiff base in bacteriorhodopsin: a density functional study. J. Phys. Chem. B 101, 8021–8028 (1997).
Tajkhorshid, E., Baudry, J., Schulten, K. & Suhai, S. Molecular dynamics study of the nature and origin of retinal's twisted structure in bacteriorhodopsin. Biophys. J. 78, 683–693 (2000).
Tajkhorshid, E. & Suhai, S. Influence of the methyl groups on the structure, charge distribution, and proton affinity of the retinal Schiff base. J. Phys. Chem. B 103, 5581–5590 (1999).
Baudry, J., Crouzy, S., Roux, B. & Smith, J.C. Quantum chemical and free energy simulation analysis of retinal conformational energetics. J. Chem. Inf. Comput. Sci. 37, 1018–1024 (1997).
Farrar, M.R. et al. Solid-state NMR-study of [Epsilon-C-13]Lys-Bacteriorhodospin—Schiff-base photoisomerization. Biophys. J. 65, 310–315 (1993).
Smith, S.O. et al. Structure and protein environment of the retinal chromophore in light-adapted and dark-adapted bacteriorhodopsin studied by solid-state NMR. Biochemistry 28, 8897–8904 (1989).
Sammalkorpi, M. & Lazaridis, T. Modeling a spin-labeled fusion peptide in a membrane: implications for the interpretation of EPR experiments. Biophys. J. 92, 10–22 (2007).
Higman, V.A. et al. Assigning large proteins in the solid state: a MAS NMR resonance assignment strategy using selectively and extensively 13C-labelled proteins. J. Biomol. NMR 44, 245–260 (2009).
Zech, S.G., Wand, A.J. & McDermott, A.E. Protein structure determination by high-resolution solid-state NMR spectroscopy: application to microcrystalline ubiquitin. J. Am. Chem. Soc. 127, 8618–8626 (2005).
Dominguez, C., Boelens, R. & Bonvin, A.M. HADDOCK: a protein-protein docking approach based on biochemical or biophysical information. J. Am. Chem. Soc. 125, 1731–1737 (2003).
de Vries, S.J. et al. HADDOCK versus HADDOCK: new features and performance of HADDOCK2.0 on the CAPRI targets. Proteins 69, 726–733 (2007).
Laskowski, R.A., Macarthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK—a program to check the stereochemical quality of protein structures. J. Appl. Cryst. 26, 283–291 (1993).
Vriend, G. WHAT IF: a molecular modeling and drug design program. J. Mol. Graphics 8, 52–56 (1990).
Rodriguez, R., Chinea, G., Lopez, N., Pons, T. & Vriend, G. Homology modeling, model and software evaluation: three related resources. Bioinformatics 14, 523–528 (1998).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Acknowledgements
This research was funded by Natural Science and Engineering Research Council of Canada, National Research Foundation of Korea (Global Research Network Program), Canada Foundation for Innovation and Ontario Ministry of Economic Development and Innovation. S.W. is a recipient of the Canadian Institutes for Health Research Postdoctoral Fellowship. V.L. is supported by Canada Research Chair in Biophysics (Tier II). We thank B. Bardiaux (Leibniz-Institut für Molekulare Pharmakologie, Berlin) for providing the ARIA 2.3 program before publication, and C.P. Jaroniec and J. Lanyi for carefully reading the manuscript.
Author information
Authors and Affiliations
Contributions
S.W., L.S., I.K. and R.A.M. prepared samples; R.A.M. and L.S.B. performed biochemical and FTIR characterization; V.L. and S.W. collected SSNMR spectroscopy data; S.W. processed and analyzed the data, and calculated the structures; T.O. and A.W. synthesized isotopically labeled retinal; S.-Y.K. and K.-H.J. produced ASR mutants; L.S.B. and V.L. designed the study; S.W., L.S.B. and V.L. wrote the paper. All authors discussed the results of the study.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1–5 (PDF 3286 kb)
Rights and permissions
About this article
Cite this article
Wang, S., Munro, R., Shi, L. et al. Solid-state NMR spectroscopy structure determination of a lipid-embedded heptahelical membrane protein. Nat Methods 10, 1007–1012 (2013). https://doi.org/10.1038/nmeth.2635
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2635
This article is cited by
-
Solid-state NMR spectroscopy
Nature Reviews Methods Primers (2021)
-
Structure of membrane diacylglycerol kinase in lipid bilayers
Communications Biology (2021)
-
Extensively sparse 13C labeling to simplify solid-state NMR 13C spectra of membrane proteins
Journal of Biomolecular NMR (2021)
-
Solid-state NMR spectroscopy based atomistic view of a membrane protein unfolding pathway
Nature Communications (2019)
-
X-ray Crystallographic Structure and Oligomerization of Gloeobacter Rhodopsin
Scientific Reports (2019)