Abstract
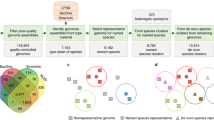
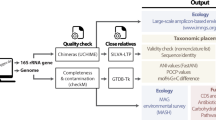
The exponentially increasing number of sequenced genomes necessitates fast, accurate, universally applicable and automated approaches for the delineation of prokaryotic species. We developed specI (species identification tool; http://www.bork.embl.de/software/specI/), a method to group organisms into species clusters based on 40 universal, single-copy phylogenetic marker genes. Applied to 3,496 prokaryotic genomes, specI identified 1,753 species clusters. Of 314 discrepancies with a widely used taxonomic classification, >62% were resolved by literature support.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Rosselló-Mora, R. & Amann, R. FEMS Microbiol. Rev. 25, 39–67 (2001).
Stackebrandt, E. et al. Int. J. Syst. Evol. Microbiol. 52, 1043–1047 (2002).
Kämpfer, P. & Glaeser, S. Environ. Microbiol. 14, 291–317 (2012).
Richter, M. & Rosselló-Móra, R. Proc. Natl. Acad. Sci. USA 106, 19126–19131 (2009).
Chun, J. et al. Int. J. Syst. Evol. Microbiol. 57, 2259–2261 (2007).
Stackebrandt, E. & Goebel, B.M. Int. J. Syst. Bacteriol. 44, 846–849 (1994).
Stackebrandt, E. & Ebers, J. Microbiol. Today 33, 152 (2006).
Konstantinidis, K. & Tiedje, J. Proc. Natl. Acad. Sci. USA 102, 2567–2572 (2005).
von Mering, C. et al. Science 315, 1126–1130 (2007).
Wu, M. & Scott, A. Bioinformatics 28, 1033–1034 (2012).
Ciccarelli, F. et al. Science 311, 1283–1287 (2006).
Creevey, C.J. et al. PLoS ONE 6, e22099 (2011).
Powell, S. et al. Nucleic Acids Res. 40, 9 (2012).
Murray, R.G.E. Int. J. Syst. Bacteriol. 46, 831 (1996).
Konstantinidis, K. & Tiedje, J. J. Bacteriol. 187, 6258–6264 (2005).
Kremer, K. et al. J. Clin. Microbiol. 37, 2607–2618 (1999).
McDonald, D. et al. ISME J. 6, 610–618 (2012).
Jousselin, E., Desdevises, Y. & Coeur d'acier, A. Proc. Royal Soc. B Biol. Sci. 276, 187–196 (2009).
Chen, X., Li, S. & Aksoy, S. J. Mol. Evol. 48, 49–58 (1999).
Brenner, D.J. Bergey's Manual of Systematic Bacteriology 1, 408–420 (The Williams & Wilkins Co., 1984).
Chisholm, S.W. et al. Nature. 334, 340–343 (1988).
Moore, L., Rocap, G. & Chisholm, S. Nature 393, 464–467 (1998).
Cowan, S. J. Gen. Microbiol. 67, 1–8 (1971).
Sorek, R. et al. Science 318, 1449–1452 (2007).
Arumugam, M., Harrington, E., Foerstner, K., Raes, J. & Bork, P. Bioinformatics 26, 2977–2978 (2010).
Altschul, S. et al. Nucleic Acids Res. 25, 3389–3402 (1997).
Huang, Y., Gilna, P. & Li, W. Bioinformatics 25, 1338–1340 (2009).
Finn, R., Clements, J. & Eddy, S. Nucleic Acids Res. 39, 37 (2011).
Pruesse, E. et al. Nucleic Acids Res. 35, 7188–7196 (2007).
Caporaso, J. et al. Bioinformatics 26, 266–267 (2010).
Jensen, L. et al. Nucleic Acids Res. 36, D250–D254 (2008).
Pearson, W. & Lipman, D. Proc. Natl. Acad. Sci. USA 85, 2444–2448 (1988).
Letunic, I. & Bork, P. Nucleic Acids Res. 39, W475–W478 (2011).
Edgar, R. Bioinformatics 26, 2460–2461 (2010).
Delcher, A., Salzberg, S. & Phillippy, A. Curr. Protoc. Bioinformatics 10, 10.3 (2003).
Muller, J., Creevey, C., Thompson, J., Arendt, D. & Bork, P. Bioinformatics 26, 263–265 (2010).
Talavera, G. & Castresana, J. Syst. Biol. 56, 564–577 (2007).
Stamatakis, A. Bioinformatics 22, 2688–2690 (2006).
Acknowledgements
We thank the members of the Bork group for helpful discussions and Y. Yuan and members of the European Molecular Biology Laboratory information technology core facility for managing the high-performance computing resources. We acknowledge funding provided by the CancerBiome project (European Research Council project reference 268985), the 'METACARDIS' project (FP7-HEALTH-2012-INNOVATION-I-305312) and the International Human Microbiome Standards project (HEALTH-F4-2010-261376).
Author information
Authors and Affiliations
Contributions
P.B., D.R.M., S.S. and G.Z. designed the study. D.R.M. developed and implemented the program, D.R.M. and G.Z. performed the experiments, D.R.M., S.S. and G.Z. analyzed the data, and D.R.M., S.S., G.Z. and P.B. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Tables 1–3, 5–7, 15, 17, 19 and 20, and Supplementary Note (PDF 1573 kb)
Supplementary Table 4
NCBI Taxonomy information of type strains listed on the list of prokaryotic names with standing in nomenclature (LPSN;http://www.bacterio.net/) that could be linked to NCBI, including their sequencing status (XLS 365 kb)
Supplementary Table 8
ANIb values of Prochlorococcusmarinus (XLS 19 kb)
Supplementary Table 9
ANIm values of Prochlorococcusmarinus (XLS 14 kb)
Supplementary Table 10
ANIb values of the Serratia and Rahnella clades (XLS 14 kb)
Supplementary Table 11
ANIm values of the Serratia and Rahnella clades (XLS 14 kb)
Supplementary Table 12
ANIb values of the Buchnera clade (XLS 15 kb)
Supplementary Table 13
ANIm values of the Buchnera clade (XLS 15 kb)
Supplementary Table 14
Cluster assignments for the 3,496 genomes used in this study (XLS 496 kb)
Supplementary Table 16
Literature-based reclassifications of species assignments of NCBI Taxonomy database (XLS 94 kb)
Supplementary Table 18
Assignments of genomes were previously not assigned to a named species to known species using the species clustering strategy presented in this publication (XLS 28 kb)
Rights and permissions
About this article
Cite this article
Mende, D., Sunagawa, S., Zeller, G. et al. Accurate and universal delineation of prokaryotic species. Nat Methods 10, 881–884 (2013). https://doi.org/10.1038/nmeth.2575
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2575
This article is cited by
-
Rhodopsin-mediated nutrient uptake by cultivated photoheterotrophic Verrucomicrobiota
The ISME Journal (2023)
-
Virus–pathogen interactions improve water quality along the Middle Route of the South-to-North Water Diversion Canal
The ISME Journal (2023)
-
Systematic analysis of drug combinations against Gram-positive bacteria
Nature Microbiology (2023)
-
Population-level impacts of antibiotic usage on the human gut microbiome
Nature Communications (2023)
-
Ecological divergence of syntopic marine bacterial species is shaped by gene content and expression
The ISME Journal (2023)