Abstract
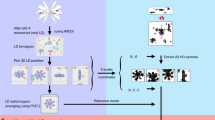
In recent work with large high-symmetry viruses, single-particle electron cryomicroscopy (cryo-EM) has achieved the determination of near-atomic-resolution structures by allowing direct fitting of atomic models into experimental density maps. However, achieving this goal with smaller particles of lower symmetry remains challenging. Using a newly developed single electron–counting detector, we confirmed that electron beam–induced motion substantially degrades resolution, and we showed that the combination of rapid readout and nearly noiseless electron counting allow image blurring to be corrected to subpixel accuracy, restoring intrinsic image information to high resolution (Thon rings visible to ∼3 Å). Using this approach, we determined a 3.3-Å-resolution structure of an ∼700-kDa protein with D7 symmetry, the Thermoplasma acidophilum 20S proteasome, showing clear side-chain density. Our method greatly enhances image quality and data acquisition efficiency—key bottlenecks in applying near-atomic-resolution cryo-EM to a broad range of protein samples.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Wolf, M., Garcea, R.L., Grigorieff, N. & Harrison, S.C. Subunit interactions in bovine papillomavirus. Proc. Natl. Acad. Sci. USA 107, 6298–6303 (2010)10.1073/pnas.0914604107.
Zhang, X., Jin, L., Fang, Q., Hui, W.H. & Zhou, Z.H. 3.3 A cryo-EM structure of a nonenveloped virus reveals a priming mechanism for cell entry. Cell 141, 472–482 (2010).
Chen, J.Z. et al. Molecular interactions in rotavirus assembly and uncoating seen by high-resolution cryo-EM. Proc. Natl. Acad. Sci. USA 106, 10644–10648 (2009).
Zhang, X. et al. Near-atomic resolution using electron cryomicroscopy and single-particle reconstruction. Proc. Natl. Acad. Sci. USA 105, 1867–1872 (2008).
Zhang, R. et al. 4.4 Å cryo-EM structure of an enveloped alphavirus Venezuelan equine encephalitis virus. EMBO J. 30, 3854–3863 (2011).
Zhang, J. et al. Mechanism of folding chamber closure in a group II chaperonin. Nature 463, 379–383 (2010).
Henderson, R. The potential and limitations of neutrons, electrons and X-rays for atomic resolution microscopy of unstained biological molecules. Q. Rev. Biophys. 28, 171–193 (1995).
McMullan, G., Chen, S., Henderson, R. & Faruqi, A.R. Detective quantum efficiency of electron area detectors in electron microscopy. Ultramicroscopy 109, 1126–1143 (2009)10.1016/j.ultramic.2009.04.002.
McMullan, G. et al. Experimental observation of the improvement in MTF from backthinning a CMOS direct electron detector. Ultramicroscopy 109, 1144–1147 (2009)10.1016/j.ultramic.2009.05.005.
McMullan, G., Clark, A.T., Turchetta, R. & Faruqi, A.R. Enhanced imaging in low dose electron microscopy using electron counting. Ultramicroscopy 109, 1411–1416 (2009)10.1016/j.ultramic.2010.07.012.
Bammes, B.E., Rochat, R.H., Jakana, J., Chen, D.H. & Chiu, W. Direct electron detection yields cryo-EM reconstructions at resolutions beyond 3/4 Nyquist frequency. J. Struct. Biol. 177, 589–601 (2012)10.1016/j.jsb.2012.01.008.
Milazzo, A.C. et al. Initial evaluation of a direct detection device detector for single particle cryo-electron microscopy. J. Struct. Biol. 176, 404–408 (2011)10.1016/j.jsb.2011.09.002.
Henderson, R. & Glaeser, R.M. Quantitative analysis of image contrast in electron micrographs of beam-sensitive crystals. Ultramicroscopy 16, 139–150 (1985).
Glaeser, R.M., McMullan, G., Faruqi, A.R. & Henderson, R. Images of paraffin monolayer crystals with perfect contrast: minimization of beam-induced specimen motion. Ultramicroscopy 111, 90–100 (2011)10.1016/j.ultramic.2010.10.010.
Typke, D., Gilpin, C.J., Downing, K.H. & Glaeser, R.M. Stroboscopic image capture: reducing the dose per frame by a factor of 30 does not prevent beam-induced specimen movement in paraffin. Ultramicroscopy 107, 106–115 (2007)10.1016/j.ultramic.2006.06.005.
Miyazawa, A., Fujiyoshi, Y., Stowell, M. & Unwin, N. Nicotinic acetylcholine receptor at 4.6 Å resolution: transverse tunnels in the channel wall. J. Mol. Biol. 288, 765–786 (1999).
Jensen, G.J. Alignment error envelopes for single particle analysis. J. Struct. Biol. 133, 143–155 (2001)10.1006/jsbi.2001.4334.
Rosenthal, P.B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003).
Saad, A. et al. Fourier amplitude decay of electron cryomicroscopic images of single particles and effects on structure determination. J. Struct. Biol. 133, 32–42 (2001).
Brilot, A.F. et al. Beam-induced motion of vitrified specimen on holey carbon film. J. Struct. Biol. 177, 630–637 (2012)10.1016/j.jsb.2012.02.003.
Campbell, M.G. et al. Movies of ice-embedded particles enhance resolution in electron cryo-microscopy. Structure 20, 1823–1828 (2012)10.1016/j.str.2012.08.026.
Bai, X.-c., Fernandez, I.S., McMullan, G. & Scheres, S.H.W. Ribosome structures to near-atomic resolution from thirty thousand cryo-EM particles. ELife 2, e00461 (2013)10.7554/eLife.00461.
Battaglia, M. et al. A rad-hard CMOS active pixel sensor for electron microscopy. Nucl. Instrum. Methods Phys. Res. A 598, 642–649 (2009)10.1016/j.nima.2008.09.029.
Bichsel, H. Straggling in thin silicon detectors. Rev. Mod. Phys. 60, 663–699 (1988)10.1103/RevModPhys.60.663.
Booth, C.R., Jakana, J. & Chiu, W. Assessing the capabilities of a 4kx4k CCD camera for electron cryo-microscopy at 300 kV. J. Struct. Biol. 156, 556–563 (2006)10.1016/j.jsb.2006.08.019.
Löwe, J. et al. Crystal structure of the 20S proteasome from the archaeon T. acidophilum at 3.4 Å resolution. Science 268, 533–539 (1995).
Rabl, J. et al. Mechanism of gate opening in the 20S proteasome by the proteasomal ATPases. Mol. Cell 30, 360–368 (2008).
Mindell, J.A. & Grigorieff, N. Accurate determination of local defocus and specimen tilt in electron microscopy. J. Struct. Biol. 142, 334–347 (2003).
Scheres, S.H. & Chen, S. Prevention of overfitting in cryo-EM structure determination. Nat. Methods 9, 853–854 (2012)10.1038/nmeth.2115.
Yu, Y. et al. Interactions of PAN's C-termini with archaeal 20S proteasome and implications for the eukaryotic proteasome-ATPase interactions. EMBO J. 29, 692–702 (2010).
Stark, H., Zemlin, F. & Boettcher, C. Electron radiation damage to protein crystals of bacteriorhodopsin at different temperatures. Ultramicroscopy 63, 75–79 (1996)10.1016/0304-3991(96)00045-9.
Mooney, P. Optimization of image collection for cellular electron microscopy. Methods Cell Biol. 79, 661–719 (2007)10.1016/S0091-679X(06)79027-6.
Meyer, R.R., Kirkland, A.I., Dunin-Borkowski, R.E. & Hutchison, J.L. Experimental characterisation of CCD cameras for HREM at 300 kV. Ultramicroscopy 85, 9–13 (2000).
Roseman, A.M. FindEM–a fast, efficient program for automatic selection of particles from electron micrographs. J. Struct. Biol. 145, 91–99 (2004).
Shaikh, T.R. et al. SPIDER image processing for single-particle reconstruction of biological macromolecules from electron micrographs. Nat. Protoc. 3, 1941–1974 (2008).
Li, X., Grigorieff, N. & Cheng, Y. GPU-enabled FREALIGN: accelerating single particle 3D reconstruction and refinement in Fourier space on graphics processors. J. Struct. Biol. 172, 407–412 (2010).
Pettersen, E.F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Trabuco, L.G., Villa, E., Mitra, K., Frank, J. & Schulten, K. Flexible fitting of atomic structures into electron microscopy maps using molecular dynamics. Structure 16, 673–683 (2008).
Acknowledgements
We thank K. Egami (UCSF) for purifying T. acidophilum 20S proteasome. We thank B. Lee for support in system optimization and DQE analysis and T. Sha for support in integrating the camera into the UCSF software environment. M. Lent was a principal architect of the camera and supported testing and troubleshooting of our prototype camera. This work is supported by the HHMI (D.A.A.) and US National Science Foundation grant DBI-0960271 to D.A.A and Y.C., which in part funded the development of the K2 camera in association with Gatan and P. Denes at Lawrence Berkeley Labs. An initial grant from the HHMI funded the first pixel prototype chip in collaboration with P. Denes. This work is also supported by the UCSF Program for Breakthrough Biomedical Research and US National Institutes of Health grants R01GM082893, R01GM098672 and S10RR026814 to Y.C. and P50GM082250 to A. Frankel.
Author information
Authors and Affiliations
Contributions
X.L., D.A.A. and Y.C. designed experiments. X.L. carried out all experiments. P.M. and C.K.B. determined DQE curves (Fig. 1a). S.Z. participated in implementing the K2 and dose fractionation. S.G. was the chief architect of the K2 project and, along with P.M., contributed significant scientific and technical insights throughout the project. All of the data in Figure 1a were collected at UCSF, and all of the other figures are based solely on experiments carried in the laboratories of Y.C. and D.A.A. M.B.B. provided technical assistance in operating the microscope. X.L., D.A.A. and Y.C. wrote the manuscript. All authors participated in discussion and revision of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
C.K.B., P.M. and S.G. are employees of Gatan Inc., which developed and is marketing the K2 camera.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12 (PDF 8693 kb)
Supplementary Software
Motion correction for dose fractionation (ZIP 502 kb)
Rights and permissions
About this article
Cite this article
Li, X., Mooney, P., Zheng, S. et al. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat Methods 10, 584–590 (2013). https://doi.org/10.1038/nmeth.2472
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2472
This article is cited by
-
Mechanism of anion exchange and small-molecule inhibition of pendrin
Nature Communications (2024)
-
One substrate many enzymes virtual screening uncovers missing genes of carnitine biosynthesis in human and mouse
Nature Communications (2024)
-
The AAA+ chaperone VCP disaggregates Tau fibrils and generates aggregate seeds in a cellular system
Nature Communications (2023)
-
Electron counting detectors in scanning transmission electron microscopy via hardware signal processing
Nature Communications (2023)
-
Reassessing chain tilt in the lamellar crystals of polyethylene
Nature Communications (2023)