Abstract
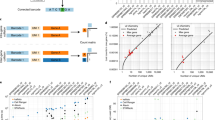
We present eXpress, a software package for efficient probabilistic assignment of ambiguously mapping sequenced fragments. eXpress uses a streaming algorithm with linear run time and constant memory use. It can determine abundances of sequenced molecules in real time and can be applied to ChIP-seq, metagenomics and other large-scale sequencing data. We demonstrate its use on RNA-seq data and show that eXpress achieves greater efficiency than other quantification methods.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Change history
04 December 2012
In the HTML version of this article initially published online, errors in mathematical terms were present in the Online Methods section. The errors have been corrected in the HTML version.
References
Lipman, D., Flicek, P., Salzberg, S., Gerstein, M. & Knight, R. Genome Biol. 12, 402 (2011).
Wold, B. & Myers, R.M. Nat. Methods 5, 19–21 (2008).
Hashimoto, T. et al. Bioinformatics 25, 2613–2614 (2009).
Li, B. & Dewey, C.N. BMC Bioinformatics 12, 323 (2011).
Trapnell, C. et al. Nat. Biotechnol. 28, 511–515 (2010).
Chung, D. et al. PLOS Comput. Biol. 7, e1002111 (2011).
Meinicke, P., Aßhauer, K.P. & Lingner, T. Bioinformatics 27, 1618–1624 (2011).
Taub, M., Lipson, D. & Speed, T.P. Commun. Inf. Syst. 10, 69–82 (2010).
Cappé, O. & Moulines, E. J. R. Stat. Soc. Series B Stat. Methodol. 71, 593–613 (2009).
Liang, P. & Klein, D. in Proc. Hum. Lang. Technol. Conf. North Am. Ch. Assoc. Comput. Linguist. 611–619 (ACL, 2009).
Hansen, K.D., Brenner, S.E. & Dudoit, S. Nucleic Acids Res. 38, e131 (2010).
Roberts, A. et al. Genome Biol. 12, R22 (2011).
Lee, S. et al. Nucleic Acids Res. 39, e9 (2011).
MAQC Consortium. et al. Nat. Biotechnol. 24, 1151–1161 (2006).
Anders, S. & Huber, W. Genome Biol. 11, R106 (2010).
Trapnell, C. et al. Nat. Biotechnol. (in the press).
Branton, D. et al. Nat. Biotechnol. 26, 1146–1153 (2008).
Stein, L.D. Genome Biol. 11, 207 (2010).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Genome Biol. 10, R25 (2009).
Ahmadi, A. et al. Nucleic Acids Res. 40, e41 (2011).
Acknowledgements
This work was supported by US National Institutes of Health grant R01HG006129. A.R. was supported in part by a National Science Foundation graduate research fellowship. We thank H. Pimentel for developing Map2GTF for converting genome mappings to transcriptome mappings and incorporating it into TopHat to help with our analysis.
Author information
Authors and Affiliations
Contributions
A.R. and L.P. developed the mathematics and statistics and designed the algorithms. A.R. implemented the method in eXpress. A.R. and L.P. tested the software and performed the analysis. A.R. and L.P. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11 and Supplementary Tables 1 and 2 (PDF 3034 kb)
Supplementary Software
eXpress source code and compiled binary files. (ZIP 3800 kb)
Rights and permissions
About this article
Cite this article
Roberts, A., Pachter, L. Streaming fragment assignment for real-time analysis of sequencing experiments. Nat Methods 10, 71–73 (2013). https://doi.org/10.1038/nmeth.2251
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2251
This article is cited by
-
Multiple bHLH/MYB-based protein complexes regulate proanthocyanidin biosynthesis in the herbage of Lotus spp.
Planta (2024)
-
Transcriptomic insights into the acclimatization response of the cold-water Ophiuroid Ophiopholis mirabilis to elevated temperatures
Marine Biology (2024)
-
Genotype by sex interactions in ankylosing spondylitis
Nature Genetics (2023)
-
Individual lipid transfer proteins from Tanacetum parthenium show different specificity for extracellular accumulation of sesquiterpenes
Plant Molecular Biology (2023)
-
Pseudogenization of the Hair-Related Genes PADI3 and S100A3 in Cetaceans and Hippopotamus amphibius
Journal of Molecular Evolution (2023)