Abstract
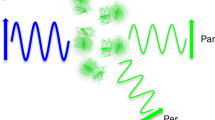
Single-molecule fluorescence imaging is often incompatible with physiological protein concentrations, as fluorescence background overwhelms an individual molecule's signal. We solve this problem with a new imaging approach called PhADE (PhotoActivation, Diffusion and Excitation). A protein of interest is fused to a photoactivatable protein (mKikGR) and introduced to its surface-immobilized substrate. After photoactivation of mKikGR near the surface, rapid diffusion of the unbound mKikGR fusion out of the detection volume eliminates background fluorescence, whereupon the bound molecules are imaged. We labeled the eukaryotic DNA replication protein flap endonuclease 1 with mKikGR and added it to replication-competent Xenopus laevis egg extracts. PhADE imaging of high concentrations of the fusion construct revealed its dynamics and micrometer-scale movements on individual, replicating DNA molecules. Because PhADE imaging is in principle compatible with any photoactivatable fluorophore, it should have broad applicability in revealing single-molecule dynamics and stoichiometry of macromolecular protein complexes at previously inaccessible fluorophore concentrations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Joo, C., Balci, H., Ishitsuka, Y., Buranachai, C. & Ha, T. Advances in single-molecule fluorescence methods for molecular biology. Annu. Rev. Biochem. 77, 51–76 (2008).
van Oijen, A.M. Single-molecule approaches to characterizing kinetics of biomolecular interactions. Curr. Opin. Biotechnol. 22, 75–80 (2011).
Klar, T.A. & Hell, S.W. Subdiffraction resolution in far-field fluorescence microscopy. Opt. Lett. 24, 954–956 (1999).
Hell, S. & Stelzer, E.H.K. Properties of a 4Pi confocal fluorescence microscope. J. Opt. Soc. Am. A 9, 2159–2166 (1992).
Levene, M.J. et al. Zero-mode waveguides for single-molecule analysis at high concentrations. Science 299, 682–686 (2003).
Boukobza, E., Sonnenfeld, A. & Haran, G. Immobilization in surface-tethered lipid vesicles as a new tool for single biomolecule spectroscopy. J. Phys. Chem. B 105, 12165–12170 (2001).
Habuchi, S., Tsutsui, H., Kochaniak, A.B., Miyawaki, A. & van Oijen, A.M. mKikGR, a monomeric photoswitchable fluorescent protein. PLoS ONE 3, e3944 (2008).
Masai, H., Matsumoto, S., You, Z., Yoshizawa-Sugata, N. & Oda, M. Eukaryotic chromosome DNA replication: Where, when, and how? Annu. Rev. Biochem. 79, 89–130 (2010).
Burgers, P.M.J. Polymerase dynamics at the eukaryotic DNA replication fork. J. Biol. Chem. 284, 4041–4045 (2009).
Herrick, J. & Bensimon, A. Introduction to molecular combing: genomics, DNA replication, and cancer. in DNA Replication: Methods and Protocols (eds. Vengrova, S. & Dalgaard, J.Z.) Ch. 5, 71–101 (Humana, Totowa, NJ, 2009).
Walter, J., Sun, L. & Newport, J. Regulated chromosomal DNA replication in the absence of a nucleus. Mol. Cell 1, 519–529 (1998).
Strausfeld, U.P. et al. Cip1 blocks the initiation of DNA replication in Xenopus extracts by inhibition of cyclin-dependent kinases. Curr. Biol. 4, 876–883 (1994).
Blow, J.J. Control of chromosomal DNA replication in the early Xenopus embryo. EMBO J. 20, 3293–3297 (2001).
Lebofsky, R., Takahashi, T. & Walter, J.C. DNA replication in nucleus-free Xenopus egg extracts. Methods Mol. Biol. 521, 229–252 (2009).
Yardimci, H., Loveland, A.B., Habuchi, S., van Oijen, A.M. & Walter, J.C. Uncoupling of sister replisomes during eukaryotic DNA replication. Mol. Cell 40, 834–840 (2010).
Gary, R., Kim, K., Cornelius, H.L., Park, M.S. & Matsumoto, Y. Proliferating cell nuclear antigen facilitates excision in long-patch base excision repair. J. Biol. Chem. 274, 4354–4363 (1999).
Harrington, J.J. & Lieber, M.R. The characterization of a mammalian DNA structure-specific endonuclease. EMBO J. 13, 1235–1246 (1994).
Shen, B., Nolan, J.P., Sklar, L.A. & Park, M.S. Functional analysis of point mutations in human flap endonuclease-1 active site. Nucleic Acids Res. 25, 3332–3338 (1997).
Blow, J.J. & Laskey, R.A. Initiation of DNA replication in nuclei and purified DNA by a cell-free extract of Xenopus eggs. Cell 47, 577–587 (1986).
Walter, J. & Newport, J.W. Regulation of replicon size in Xenopus egg extracts. Science 275, 993–995 (1997).
Herrick, J., Stanislawski, P., Hyrien, O. & Bensimon, A. Replication fork density increases during DNA synthesis in X. laevis egg extracts. J. Mol. Biol. 300, 1133–1142 (2000).
Lucas, I., Chevrier-Miller, M., Sogo, J.M. & Hyrien, O. Mechanisms ensuring rapid and complete DNA replication despite random initiation in Xenopus early embryos. J. Mol. Biol. 296, 769–786 (2000).
Blow, J.J., Gillespie, P.J., Francis, D. & Jackson, D.A. Replication origins in Xenopus egg extract are 5–15 kb apart and are activated in clusters that fire at different times. J. Cell Biol. 152, 15–26 (2001).
Marheineke, K. & Hyrien, O. Control of replication origin density and firing time in Xenopus egg extracts. J. Biol. Chem. 279, 28071–28081 (2004).
Marheineke, K. & Hyrien, O. Aphidicolin triggers a block to replication origin firing in Xenopus egg extracts. J. Biol. Chem. 276, 17092–17100 (2001).
Ridelis, I. et al. Use of Kikume green-red fusions to study the influence of pharmacological chaperones on trafficking of G protein–coupled receptors. FEBS Lett. 586, 784–791 (2012).
Herrick, J., Jun, S., Bechhoefer, J. & Bensimon, A. Kinetic model of DNA replication in eukaryotic organisms. J. Mol. Biol. 320, 741–750 (2002).
Ayyagari, R., Gomes, X.V., Gordenin, D.A. & Burgers, P.M.J. Okazaki fragment maturation in yeast. I. Distribution of functions between FEN1 and DNA2. J. Biol. Chem. 278, 1618–1625 (2003).
Hoskins, A.A., Gelles, J. & Moore, M.J. New insights into the spliceosome by single molecule fluorescence microscopy. Curr. Opin. Chem. Biol. 15, 864–870 (2011).
Jain, A. et al. Probing cellular protein complexes using single-molecule pull-down. Nature 473, 484–488 (2011).
Thompson, M.A., Biteen, J.S., Lord, S.J., Conley, N.R. & Moerner, W.E. Molecules and methods for super-resolution imaging. in Single Molecule Tools, Part B: Super-Resolution, Particle Tracking, Multiparameter, and Force Based Methods (ed. Walter, N.G.) Ch. 2, 27–59 (Academic Press, 2010).
Lord, S.J. et al. Azido push-pull fluorogens photoactivate to produce bright fluorescent labels. J. Phys. Chem. B 114, 14157–14167 (2010).
Bates, M., Blosser, T.R. & Zhuang, X. Short-range spectroscopic ruler based on a single-molecule optical switch. Phys. Rev. Lett. 94, 108101 (2005).
Conley, N.R., Biteen, J.S. & Moerner, W.E. Cy3-Cy5 covalent heterodimers for single-molecule photoswitching. J. Phys. Chem. B 112, 11878–11880 (2008).
Patterson, G., Davidson, M., Manley, S. & Lippincott-Schwartz, J. Superresolution imaging using single-molecule localization. Annu. Rev. Phys. Chem. 61, 345–367 (2010).
Loparo, J.J., Kulczyk, A.W., Richardson, C.C. & van Oijen, A.M. Simultaneous single-molecule measurements of phage T7 replisome composition and function reveal the mechanism of polymerase exchange. Proc. Natl. Acad. Sci. USA 108, 3584–3589 (2011).
Havens, C.G. & Walter, J.C. Docking of a specialized PIP box onto chromatin-bound PCNA creates a degron for the ubiquitin ligase CRL4Cdt2. Mol. Cell 35, 93–104 (2009).
Bibikova, M. et al. Characterization of FEN-1 from Xenopus laevis. J. Biol. Chem. 273, 34222–34229 (1998).
Walter, J.C. Evidence for sequential action of cdc7 and cdk2 protein kinases during initiation of DNA replication in Xenopus egg extracts. J. Biol. Chem. 275, 39773–39778 (2000).
Wohlschlegel, J.A. et al. Inhibition of eukaryotic DNA replication by geminin binding to Cdt1. Science 290, 2309–2312 (2000).
Yardimci, H., Loveland, A.B., van Oijen, A.M. & Walter, J.C. Single-molecule analysis of DNA replication in Xenopus egg extracts. Methods. published online, doi:10.1016/j.ymeth.2012.03.033 (6 April 2012).
Lebofsky, R., van Oijen, A.M. & Walter, J.C. DNA is a co-factor for its own replication in Xenopus egg extracts. Nucleic Acids Res. 39, 545–555 (2011).
Edwards, M.C. et al. MCM2–7 complexes bind chromatin in a distributed pattern surrounding the origin recognition complex in Xenopus egg extracts. J. Biol. Chem. 277, 33049–33057 (2002).
Abràmoff, M.D., Magalhães, P.J. & Ram, S.J. Image processing with ImageJ. Biophotonics Int. 11, 36–42 (2004).
Acknowledgements
We thank H. Yardimci and J.J. Loparo for assistance with experiments and critical reading of the manuscript and C.G. Havens (Harvard University) for sharing advice and reagents. This work was supported by grants from the US National Institutes of Health (NIH) (GM077248), American Cancer Society (RSG0823401GMC) and Netherlands Organization for Scientific Research (NWO; Vici 680-47-607) to A.M.v.O. as well as by grants from the NIH (GM62267) and American Cancer Society (RSG0823401GMC) to J.C.W. A.B.L. was supported by the NIH and National Institute of General Medical Sciences Molecular Biophysics Training Grant (T32 GM008313).
Author information
Authors and Affiliations
Contributions
A.B.L. performed all experiments. A.M.v.O. and S.H. designed the microscope. A.B.L., A.M.v.O. and J.C.W. designed experiments and performed data analysis. A.B.L., A.M.v.O. and J.C.W. prepared the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Tables 1–3 and Supplementary Methods (PDF 1007 kb)
PhADE imaging of Fen1KikR under conditions when only one origin fires reveals the growth rate of individual replication bubbles.
D179A Fen1KikR was imaged (as in Fig. 2b) using PhADE between 5 and 25 min after NPE addition (time stamps). During this time, D179A Fen1KikR signals appeared, grew in length, and some eventually split as forks traveled far apart. Some D179A Fen1KikR signals were lost as DNA tethers broke and DNA compacted. The extracts were washed out 25 min after NPE addition, dig-dUTP incorporation was detected and DNA was stained with an intercalating dye. (AVI 5321 kb)
PhADE imaging of D179A Fen1KikR reveals the timing and distribution of initiation events.
D179A Fen1KikR was imaged using PhADE (as in Fig. 3a) between 2.5 and 10 min after NPE addition (time stamps). During this time, D179A Fen1KikR signals appeared, grew and merged, which we interpret as replication initiation, elongation and termination. Some molecules were lost as DNA tethers broke and DNA compacted. A kymograph of the molecule labeled #1 in the first frame of the movie is shown in Figure 3a. All molecules highlighted with cyan boxes in the first frame were analyzed for Fig. 3b–d. Arrows indicate QDot 605-Biotin that are used as fiduciary markers. (AVI 1942 kb)
Rights and permissions
About this article
Cite this article
Loveland, A., Habuchi, S., Walter, J. et al. A general approach to break the concentration barrier in single-molecule imaging. Nat Methods 9, 987–992 (2012). https://doi.org/10.1038/nmeth.2174
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2174
This article is cited by
-
Control of DNA replication in vitro using a reversible replication barrier
Nature Protocols (2024)
-
The mechanism of DNA replication termination in vertebrates
Nature (2015)
-
Probing molecular choreography through single-molecule biochemistry
Nature Structural & Molecular Biology (2015)
-
An improved surface passivation method for single-molecule studies
Nature Methods (2014)
-
Bacterial replication, transcription and translation: mechanistic insights from single-molecule biochemical studies
Nature Reviews Microbiology (2013)