Abstract
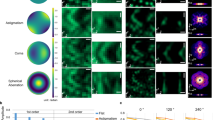
Three-dimensional (3D) structured-illumination microscopy (SIM) can double the lateral and axial resolution of a wide-field fluorescence microscope but has been too slow for live imaging. Here we apply 3D SIM to living samples and record whole cells at up to 5 s per volume for >50 time points with 120-nm lateral and 360-nm axial resolution. We demonstrate the technique by imaging microtubules in S2 cells and mitochondria in HeLa cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Westphal, V. et al. Science 320, 246–249 (2008).
Betzig, E. et al. Science 313, 1642–1645 (2006).
Rust, M.J., Bates, M. & Zhuang, X. Nat. Methods 3, 793–795 (2006).
Huang, B., Wang, W., Bates, M. & Zhuang, X. Science 319, 810–813 (2008).
Juette, M.F. et al. Nat. Methods 5, 527–529 (2008).
Shtengel, G. et al. Proc. Natl. Acad. Sci. USA 106, 3125–3130 (2009).
Huang, B., Jones, S.A., Brandenburg, B. & Zhuang, X. Nat. Methods 5, 1047–1052 (2008).
York, A.G., Ghitani, A., Vaziri, A., Davidson, M.W. & Shroff, H. Nat. Methods 8, 327–333 (2011).
Shroff, H., Galbraith, C.G., Galbraith, J.A. & Betzig, E. Nat. Methods 5, 417–423 (2008).
Wombacher, R. et al. Nat. Methods 7, 717–719 (2010).
Jones, S.A., Shim, S., He, J. & Zhuang, X. Nat. Methods 8, 499–508 (2011).
Lauterbach, M.A. et al. J. Biophoton. 3, 417–424 (2010).
Gustafsson, M.G.L. et al. Biophys. J. 94, 4957–4970 (2008).
Kner, P., Chhun, B.B., Griffis, E.R., Winoto, L. & Gustafsson, M.G. Nat. Methods 6, 339–342 (2009).
Hirvonen, L.M., Wicker, K., Mandula, O. & Heintzmann, R. Eur. Biophys. J. 38, 807–812 (2009).
Bereiter-Hahn, J. & Jendrach, M. Int. Rev. Cell Mol. Biol. 284, 1–65 (2010).
Rogers, S.L., Rogers, G.C., Sharp, D.J. & Vale, R.D. J. Cell Biol. 158, 873–884 (2002).
Maiato, H., Rieder, C.L. & Khodjakov, A. J. Cell Biol. 167, 831–840 (2004).
Acknowledgements
We dedicate this paper to the late Dr. Mats Gustafsson. We thank H. White and A. Arnold for assistance with sample preparation; S. Goodwin, S.A. Ribeiro and R. Vale (University of California, San Francisco), for providing the S2 cells; R. Fiolka and E. Betzig for valuable discussions, and L. Winoto for sharing the design of the optical fiber shaker.
Author information
Authors and Affiliations
Contributions
L.S. and P.K. built the optical hardware and wrote the software for the control system and image acquisition. P.K. built the electronics circuit that interfaces the digital signal processing board with the rest of the system. L.S. prepared the samples, and acquired and processed data. E.H.R. provided guidance on biological applications. M.G.L.G. made the conceptual design. L.S. and E.H.R. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8, Supplementary Tables 1–2, Supplementary Note, Supplementary Discussion (PDF 2889 kb)
Supplementary Video 1
Maximum-intensity projection from different angles of a live 3D SIM time series of a Drosophila S2 cell stably expressing α-tubulin–EGFP (Fig. 1). Volume thickness, 2.88 μm. Number of time points, 150. Time needed per 3D volume, 5 s. Time inserted between time points, 5.5 s. Ratio between this video's frame rate and the real-time frame rate, ∼62. Scale bar, 2 μm. (MOV 9025 kb)
Supplementary Video 2
Maximum-intensity projection from different angles of a live 3D SIM time series of a HeLa cell stained with 50 nM MitoTracker Green (Fig. 2). Volume thickness, 6.1 μm. Number of time points, 50. Time needed per 3D volume, 20.3 s. Time inserted between time points, 6 s. Scale bar, 2 μm. (MOV 7508 kb)
Supplementary Video 3
Maximum-intensity projection along z axis of a live 3D SIM time series of a HeLa cell stained with 50 nM MitoTracker Green (Fig. 2). Volume thickness, 6.1 μm. Number of time points, 50. Time needed per 3D volume, 20.3 s. Time inserted between time points, 6 s. Scale bar, 2 μm. (MOV 7395 kb)
Supplementary Video 4
HeLa cell mitochondria dynamics under live 3D SIM. Each frame was cropped from one axial plane at each time point of the same dataset shown in Figure 2 and Supplementary Video 2, and contains interesting events of mitochondrial fission-fusion and other morphological changes. Scale bar, 1 μm. (MOV 253 kb)
Supplementary Video 5
Dynamics of fine internal mitochondrial structures of a HeLa cell under live 3D SIM. Each frame is a maximum-intensity projection over a thickness of 1.3 μm for a subregion at each time point of the same dataset shown in Figure 2 and Supplementary Video 2 that contains the featured 'Y'-shaped mitochondrion. Besides the overall motion, one can observe the dynamics of the fine internal mitochondrial structures that resemble cristae. Scale bar, 1 μm. (MOV 390 kb)
Supplementary Video 6
Maximum-intensity projection from different angles of another live 3D SIM time series of a HeLa cell stained with 50 nM MitoTracker Green. Volume thickness, 6.4 μm. Number of time points, 75. Time needed per 3D volume, 20.8 s. Time inserted between time points, 2.5 s. Scale bar, 2 μm. (MOV 9539 kb)
Supplementary Video 7
An example of motion artifact. This is a mitochondrion from the same dataset shown in Fig. 2 and Supplementary Video 2. Scale bar: 1 μm. (MOV 877 kb)
Supplementary Video 8
Variation in zero-order illumination over time owing to insufficient time-averaging of the speckle pattern in the laser output from the multimode fiber. To generate spatially incoherent and smooth illumination light in 3D SIM, a mechanical fiber vibrator was used for shaking the fiber rapidly (Online Methods). This dataset, taken with only the zero-order diffraction beam to simulate uniform (that is, not structured) illumination, is for determining the non-uniformity in the nominally uniform illumination. Sample, thin film (< 300 nm thick) of fluorescein. Exposure time, 10 ms. The bright spots in the video were presumably where fluorescein molecules aggregated. It was determined (Supplementary Discussion) that the non-uniformity of the illumination using this exposure time was much less than 5%. Scale bar, 1 μm. (MOV 2571 kb)
Rights and permissions
About this article
Cite this article
Shao, L., Kner, P., Rego, E. et al. Super-resolution 3D microscopy of live whole cells using structured illumination. Nat Methods 8, 1044–1046 (2011). https://doi.org/10.1038/nmeth.1734
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1734
This article is cited by
-
Parameter estimation of the structured illumination pattern based on principal component analysis (PCA): PCA-SIM
Light: Science & Applications (2023)
-
MAxSIM: multi-angle-crossing structured illumination microscopy with height-controlled mirror for 3D topological mapping of live cells
Communications Biology (2023)
-
Superresolution structured illumination microscopy reconstruction algorithms: a review
Light: Science & Applications (2023)
-
Three-dimensional structured illumination microscopy with enhanced axial resolution
Nature Biotechnology (2023)
-
Open-3DSIM: an open-source three-dimensional structured illumination microscopy reconstruction platform
Nature Methods (2023)