Abstract
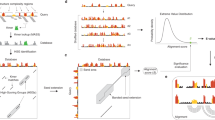
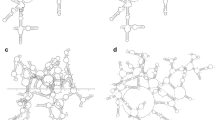
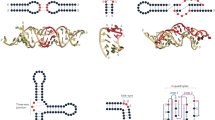
Structural RNA modules, sets of ordered non-Watson-Crick base pairs embedded between Watson-Crick pairs, have central roles as architectural organizers and sites of ligand binding in RNA molecules, and are recurrently observed in RNA families throughout the phylogeny. Here we describe a computational tool, RNA three-dimensional (3D) modules detection, or RMDetect, for identifying known 3D structural modules in single and multiple RNA sequences in the absence of any other information. Currently, four modules can be searched for: G-bulge loop, kink-turn, C-loop and tandem-GA loop. In control test sequences we found all of the known modules with a false discovery rate of 0.23. Scanning through 1,444 publicly available alignments, we identified 21 yet unreported modules and 141 known modules. RMDetect can be used to refine RNA 2D structure, assemble RNA 3D models, and search and annotate structured RNAs in genomic data.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Leontis, N.B., Stombaugh, J. & Westhof, E. The non-Watson-Crick base pairs and their associated isostericity matrices. Nucleic Acids Res. 30, 3497–3531 (2002).
Lescoute, A. et al. Recurrent structural RNA motifs, Isostericity Matrices and sequence alignments. Nucleic Acids Res. 33, 2395–2409 (2005).
Leontis, N.B. & Westhof, E. Analysis of RNA motifs. Curr. Opin. Struct. Biol. 13, 300–308 (2003).
Leontis, N.B. & Westhof, E. A common motif organizes the structure of multi-helix loops in 16 S and 23 S ribosomal RNAs. J. Mol. Biol. 283, 571–583 (1998).
Macke, T.J. et al. RNAMotif, an RNA secondary structure definition and search algorithm. Nucleic Acids Res. 29, 4724–4735 (2001).
Yao, Z., Weinberg, Z. & Ruzzo, W.L. CMfinder—a covariance model based RNA motif finding algorithm. Bioinformatics 22, 445–452 (2006).
Parisien, M. & Major, F. The MC-Fold and MC-Sym pipeline infers RNA structure from sequence data. Nature 452, 51–55 (2008).
Jossinet, F., Ludwig, T.E. & Westhof, E. Assemble: an interactive graphical tool to analyze and build RNA architectures at the 2D and 3D levels. Bioinformatics 26, 2057–2059 (2010).
Das, R., Karanicolas, J. & Baker, D. Atomic accuracy in predicting and designing noncanonical RNA structure. Nat. Methods 7, 291–294 (2010).
Westhof, E. The amazing world of bacterial structured RNAs. Genome Biol. 11, 108 (2010).
Moore, P.B. Structural Motifs in RNA. Annu. Rev. Biochem. 68, 287–300 (1999).
Klein, D.J. et al. The kink-turn: a new RNA secondary structure motif. EMBO J. 20, 4214–4221 (2001).
Djelloul, M. & Denise, A. Automated motif extraction and classification in RNA tertiary structures. RNA 14, 2489–2497 (2008).
Sarver, M. et al. FR3D: finding local and composite recurrent structural motifs in RNA 3D structures. J. Math. Biol. 56, 215–252 (2008).
Apostolico, A. et al. Finding 3D motifs in ribosomal RNA structures. Nucleic Acids Res. 37, e29 (2009).
Zhong, C., Tang, H. & Zhang, S. RNAMotifScan: automatic identification of RNA structural motifs using secondary structural alignment. Nucleic Acids Res. 38, e176 (2010).
Thébault, P. et al. Searching RNA motifs and their intermolecular contacts with constraint networks. Bioinformatics 22, 2074–2080 (2006).
Gautheret, D., Konings, D. & Gutell, R.R. A major family of motifs involving G? A mismatches in ribosomal RNA. J. Mol. Biol. 242, 1–8 (1994).
Leontis, N.B., Stombaugh, J. & Westhof, E. Motif prediction in ribosomal RNAs Lessons and prospects for automated motif prediction in homologous RNA molecules. Biochimie 84, 961–973 (2002).
Serganov, A., Huang, L. & Patel, D.J. Structural insights into amino acid binding and gene control by a lysine riboswitch. Nature 455, 1263–1267 (2008).
Golden, B.L., Kim, H. & Chase, E. Crystal structure of a phage Twort group I ribozyme-product complex. Nat. Struct. Mol. Biol. 12, 82–89 (2005).
Wang, J., Henkin, T.M. & Nikonowicz, E.P. NMR structure and dynamics of the specifier loop domain from the Bacillus subtilis tyrS T box leader RNA. Nucleic Acids Res. 38, 3388–3398 (2010).
Kuhn, J.F., Tran, E.J. & Maxwell, E.S. Archaeal ribosomal protein L7 is a functional homolog of the eukaryotic 15.5kD/Snu13p snoRNP core protein. Nucleic Acids Res. 30, 931–941 (2002).
Sankaranarayanan, R. et al. The structure of threonyl-tRNA synthetase-tRNA(Thr) complex enlightens its repressor activity and reveals an essential zinc ion in the active site. Cell 97, 371–381 (1999).
Ben-Gal, I. et al. Identification of transcription factor binding sites with variable-order Bayesian networks. Bioinformatics 21, 2657–2666 (2005).
Matthews, B. Comparison of the predicted and observed secondary structure of T4 phage lysozyme. Biochim. Biophys. Acta 405, 442–451 (1975).
Gardner, P.P. et al. Rfam: updates to the RNA families database. Nucleic Acids Res. 37, D136–D140 (2009).
Wilm, A., Mainz, I. & Steger, G. An enhanced RNA alignment benchmark for sequence alignment programs. Algorithms Mol. Biol. 1, 19 (2006).
Zhou, Y. et al. GISSD: Group I Intron Sequence and Structure Database. Nucleic Acids Res. 36, D31–D37 (2008).
Weinberg, Z. et al. Exceptional structured noncoding RNAs revealed by bacterial metagenome analysis. Nature 462, 656–659 (2009).
Weinberg, Z. et al. Comparative genomics reveals 104 candidate structured RNAs from bacteria, archaea, and their metagenomes. Genome Biol. 11, R31 (2010).
Vitreschak, A.G. Regulation of the vitamin B12 metabolism and transport in bacteria by a conserved RNA structural element. RNA 9, 1084–1097 (2003).
Lee, H. et al. cis-acting RNA signals in the NS5B C-terminal coding sequence of the Hepatitis C virus genome. J. Virol. 78, 10865–10877 (2004).
Le Quesne, J.P. et al. Derivation of a structural model for the c-myc IRES. J. Mol. Biol. 310, 111–126 (2001).
Paul, AV. et al. Identification of an RNA hairpin in poliovirus RNA that serves as the primary template in the in vitro uridylylation of VPg. J. Virology 74, 10359–10370 (2000).
Wassarman, K.M. et al. Identification of novel small RNAs using comparative genomics and microarrays. Genes Dev. 1637–1651 (2001).
Bösl, M. & Kersten, H. A novel RNA product of the tyrT operon of Escherichia coli. Nucleic Acids Res. 19, 5863–5870 (1991).
Weinberg, Z. et al. Identification of 22 candidate structured RNAs in bacteria using the CMfinder comparative genomics pipeline. Nucleic Acids Res. 35, 4809–4819 (2007).
Klinck, R. et al. A potential RNA drug target in the hepatitis C virus internal ribosomal entry site. RNA 6, 1423–1431 (2000).
Gao, Y.-G. et al. The structure of the ribosome with elongation factor G trapped in the posttranslocational state. Science 326, 694–699 (2009).
Jang, S.B. et al. Structures of two RNA octamers containing tandem G.A base pairs. Acta Crystallogr. D Biol. Crystallogr. 60, 829–835 (2004).
Berman, H.M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Pruesse, E. et al. SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 35, 7188–7196 (2007).
Darty, K., Denise, A. & Ponty, Y. VARNA: Interactive drawing and editing of the RNA secondary structure. Bioinformatics 25, 1974–1975 (2009).
Durbin, R. et al. Biological Sequence Analysis: Probabilistic Models of Proteins and Nucleic Acids (Cambridge University Press, 1998).
Hofacker, I.L. et al. Fast folding and comparison of RNA secondary structures. Monatshefte für Chemie 125, 167–188 (1994).
Lindgreen, S., Gardner, P.P. & Krogh, A. Measuring covariation in RNA alignments: physical realism improves information measures. Bioinformatics 22, 2988–2995 (2006).
Acknowledgements
We thank R. Backofen for useful suggestions. J.A.C. is supported by the Ph.D. Program in Computational Biology of the Instituto Gulbenkian de Ciência, Portugal (sponsored by Fundação Calouste Gulbenkian, Siemens SA and Fundação para a Ciência e Tecnologia; SFRH/BD/33528/2008).
Author information
Authors and Affiliations
Contributions
J.A.C. conceived the algorithms, performed the computations and wrote the manuscript. E.W. conceived the research and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10, Supplementary Table 1, Supplementary Notes 1–6, Supplementary Data 1–3 (PDF 8744 kb)
Supplementary Software 1
Rmdetect (ZIP 940 kb)
Rights and permissions
About this article
Cite this article
Cruz, J., Westhof, E. Sequence-based identification of 3D structural modules in RNA with RMDetect. Nat Methods 8, 513–519 (2011). https://doi.org/10.1038/nmeth.1603
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1603
This article is cited by
-
Chemical reversible crosslinking enables measurement of RNA 3D distances and alternative conformations in cells
Nature Communications (2022)
-
PATTERNA: transcriptome-wide search for functional RNA elements via structural data signatures
Genome Biology (2018)
-
New in silico approach to assessing RNA secondary structures with non-canonical base pairs
BMC Bioinformatics (2015)