Abstract
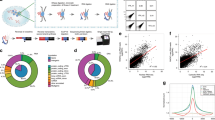
Classical approaches to determine structures of noncoding RNA (ncRNA) probed only one RNA at a time with enzymes and chemicals, using gel electrophoresis to identify reactive positions. To accelerate RNA structure inference, we developed fragmentation sequencing (FragSeq), a high-throughput RNA structure probing method that uses high-throughput RNA sequencing of fragments generated by digestion with nuclease P1, which specifically cleaves single-stranded nucleic acids. In experiments probing the entire mouse nuclear transcriptome, we accurately and simultaneously mapped single-stranded RNA regions in multiple ncRNAs with known structure. We probed in two cell types to verify reproducibility. We also identified and experimentally validated structured regions in ncRNAs with, to our knowledge, no previously reported probing data.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Gesteland, R., Cech, T. & Atkins, J. (eds). The RNA World 3rd edn. (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, New York, USA, 2005).
Affymetrix/Cold Spring Harbor Laboratory ENCODE Transcriptome Project. Post-transcriptional processing generates a diversity of 5′-modified long and short RNAs. Nature 457, 1028–1032 (2009).
Guttman, M. et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458, 223–227 (2009).
Ambros, V. microRNAs: tiny regulators with great potential. Cell 107, 823–826 (2001).
Kapranov, P. et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science 316, 1484–1488 (2007).
Knapp, G. Enzymatic approaches to probing of RNA secondary and tertiary structure. Methods Enzymol. 2, 192–212 (1989).
Low, J.T. & Weeks, K.M. SHAPE-directed RNA secondary structure prediction. Methods 52, 150–158 (2010).
Machado-Lima, A., del Portillo, H.A. & Durham, A.M. Computational methods in noncoding RNA research. J. Math. Biol. 56, 15–49 (2008).
Crawford, G.E. et al. Genome-wide mapping of DNase hypersensitive sites using massively parallel signature sequencing (MPSS). Genome Res. 16, 123–131 (2006).
Ying, Q.-L., Stavridis, M., Griffiths, D., Li, M. & Smith, A. Conversion of embryonic stem cells into neuroectodermal precursors in adherent monoculture. Nat. Biotechnol. 21, 183–186 (2003).
Desai, N.A. & Shankar, V. Single-strand-specific nucleases. FEMS Microbiol. Rev. 26, 457–491 (2003).
Cameron, V. & Uhlenbeck, O.C. 3′-phosphatase activity in T4 polynucleotide kinase. Biochemistry 16, 5120–5126 (1977).
Romier, C., Dominguez, R., Lahm, A., Dahl, O. & Suck, D. Recognition of single-stranded DNA by nuclease P1: high resolution crystal structures of complexes with substrate analogs. Proteins 32, 414–424 (1998).
Naik, A.K. & Raghavan, S.C. P1 nuclease cleavage is dependent on length of the mismatches in DNA. DNA Repair (Amst.) 7, 1384–1391 (2008).
Parker, K.A. & Steitz, J.A. Structural analyses of the human U3 ribonucleoprotein particle reveal a conserved sequence available for base pairing with pre-rRNA. Mol. Cell. Biol. 7, 2899–2913 (1987).
Mougin, A., Gottschalk, A., Fabrizio, P., Lührmann, R. & Branlant, C. Direct probing of RNA structure and RNA-protein interactions in purified HeLa cell's and yeast spliceosomal U4/U6.U5 tri-snRNP particles. J. Mol. Biol. 317, 631–649 (2002).
Granneman, S. et al. Role of pre-rRNA base pairing and 80S complex formation in subnucleolar localization of the U3 snoRNP. Mol. Cell. Biol. 24, 8600–8610 (2004).
Kass, S., Tyc, K., Steitz, J.A. & Sollner-Webb, B. The U3 small nucleolar ribonucleoprotein functions in the first step of preribosomal RNA processing. Cell 60, 897–908 (1990).
Peculis, B.A. & Steitz, J.A. Disruption of U8 nucleolar snRNA inhibits 5.8S and 28S rRNA processing in the Xenopus oocyte. Cell 73, 1233–1245 (1993).
Tycowski, K., Shu, M. & Steitz, J. Requirement for intron-encoded U22 small nucleolar RNA in 18S ribosomal RNA maturation. Science 266, 1558–1561 (1994).
Wang, Z., Gerstein, M. & Snyder, M. RNA-Seq: a revolutionary tool for transcriptomics. Nat. Rev. Genet. 10, 57–63 (2009).
Kertesz, M. et al. Genome-wide measurement of RNA secondary structure in yeast. Nature 467, 103–107 (2010).
Reuter, J.S. & Mathews, D.H. RNAstructure: software for RNA secondary structure prediction and analysis. BMC Bioinformatics 11, 129 (2010).
Mandal, M. & Breaker, R.R. Gene regulation by riboswitches. Nat. Rev. Mol. Cell Biol. 5, 451–463 (2004).
Maroney, P., Romfo, C. & Nilsen, T. Nuclease protection of RNAs containing site-specific labels: a rapid method for mapping RNA-protein interactions. RNA 6, 1905–1909 (2000).
Kiss-László, Z., Henry, Y., Bachellerie, J.P., Caizergues-Ferrer, M. & Kiss, T. Site-specific ribose methylation of preribosomal RNA: a novel function for small nucleolar RNAs. Cell 85, 1077–1088 (1996).
Beard, C., Hochedlinger, K., Plath, K., Wutz, A. & Jaenisch, R. Efficient method to generate single-copy transgenic mice by site-specific integration in embryonic stem cells. Genesis 44, 23–28 (2006).
Skarnes, W.C. Gene trapping methods for the identification and functional analysis of cell surface proteins in mice. Methods Enzymol. 328, 592–615 (2000).
Sobczak, K., Michlewski, G., de Mezer, M., Krol, J. & Krzyzosiak, W.J. Trinucleotide repeat system for sequence specificity analysis of RNA structure probing reagents. Anal. Biochem. 402, 40–46 (2010).
Mortazavi, A., Williams, B.A., Mccue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 5, 621–628 (2008).
Acknowledgements
A.V.U. was supported in part by US National Institutes of Health (NIH) bioinformatics training grant 1 T32 GM070386-01 and by a US National Science Foundation Graduate Research fellowship. S.K. was supported in part by NIH National Human Genome Research Institute grant U41 HG004568-01. C.S.O. was supported by California Institute for Regenerative Medicine training grant T3-00006. This study was funded in part by NIH R01HG004002 to D.H.M. and NIH 1R03DA026061-01 to S.R.S. We thank D. Bernick, S. Kuersten and O. Uhlenbeck for helpful discussions; Y. Ponty for adding the feature to display enzymatic/chemical modifications to VARNA, the program used to visualize our probing data; E. Farias-Hesson and N. Pourmand of the University of California Santa Cruz Genome Sequencing Center for preparing samples; workers at ABI for carrying out the sequencing; and M. Storm and F. Ng of ABI for facilitating that sequencing run.
Author information
Authors and Affiliations
Contributions
J.G.U. designed and carried out the experiments. A.V.U. designed and carried out the bioinformatics analysis, except for preparing the read mappings, which S.K. did, with C.S.O. contributing data. J.E.M. programmed additional features in the RNAstructure software. J.G.U., A.V.U. and S.R.S. wrote the manuscript. S.R.S., D.H.M., T.M.L. and D.H. directed the research.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–12, Supplementary Table 1, Supplementary Notes 1–3, Supplementary Discussion (PDF 9761 kb)
Supplementary Data 1
Stockholm-format (machine-readable) multiple alignment of U15b C/D box snoRNA homologs, containing structure models that were evaluated. See file for detailed comments. (ZIP 1 kb)
Supplementary Data 2
Stockholm-format (machine-readable) multiple alignment of U22 C/D box snoRNA homologs, containing structure models that were evaluated. See file for detailed comments. (ZIP 1 kb)
Supplementary Data 3
Stockholm-format (machine-readable) multiple alignment of U97 C/D box snoRNA homologs, containing structure models that were evaluated. See file for detailed comments. (ZIP 1 kb)
Supplementary Data 4
FASTA-format file of sequences used for filtering out sequencing reads prior to mapping to genome (see Methods). (ZIP 11 kb)
Supplementary Data 5
Six-column BED-format file containing genomic coordinates (mm9 genome assembly) of all RNAs examined in this study. This can be uploaded to the UCSC Genome Browser as a custom track. (ZIP 0 kb)
Supplementary Software
FragSeq algorithm implementation, configuration files and Readme. All FragSeq algorithm software, scripts and configuration files needed to reproduce the analysis in this paper are provided. The Readme file contains complete instructions on how to rerun our analysis. However, read mappings are not provided owing to their large size and have to be downloaded from the GEO (accession number is listed in the paper; see the Readme file). The script dpToVarna.py is also provided (Supplementary Note 3). (ZIP 45 kb)
Rights and permissions
About this article
Cite this article
Underwood, J., Uzilov, A., Katzman, S. et al. FragSeq: transcriptome-wide RNA structure probing using high-throughput sequencing. Nat Methods 7, 995–1001 (2010). https://doi.org/10.1038/nmeth.1529
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1529
This article is cited by
-
Systematic detection of tertiary structural modules in large RNAs and RNP interfaces by Tb-seq
Nature Communications (2023)
-
Comparative genomics identifies thousands of candidate structured RNAs in human microbiomes
Genome Biology (2021)
-
Expression profile of circular RNA and construction of circular RNA-Micro RNA network in salivary adenoid cystic carcinoma
Cancer Cell International (2021)
-
Global in situ profiling of RNA-RNA spatial interactions with RIC-seq
Nature Protocols (2021)
-
Improved recognition of splice sites in A. thaliana by incorporating secondary structure information into sequence-derived features: a computational study
3 Biotech (2021)