Abstract
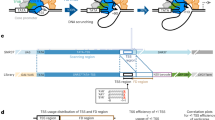
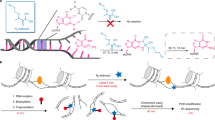
Recent studies using high-throughput sequencing protocols have uncovered the complexity of mammalian transcription by RNA polymerase II, helping to define several initiation patterns in which transcription start sites (TSSs) cluster in both narrow and broad genomic windows. Here we describe a paired-end sequencing strategy, which enables more robust mapping and characterization of capped transcripts. We used this strategy to explore the transcription initiation landscape in the Drosophila melanogaster embryo. Extending the previous findings in mammals, we found that fly promoters exhibited distinct initiation patterns, which were linked to specific promoter sequence motifs. Furthermore, we identified many 5′ capped transcripts originating from coding exons; our analyses support that they are unlikely the result of alternative TSSs, but rather the product of post-transcriptional modifications. We demonstrated paired-end TSS analysis to be a powerful method to uncover the transcriptional complexity of eukaryotic genomes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Juven-Gershon, T. & Kadonaga, J.T. Regulation of gene expression via the core promoter and the basal transcriptional machinery. Dev. Biol. 339, 225–229 (2010).
Ohler, U. & Wassarman, D.A. Promoting developing transcription. Development 137, 15–26 (2010).
Butler, J.E. & Kadonaga, J.T. Enhancer-promoter specificity mediated by DPE or TATA core promoter motifs. Genes Dev. 15, 2515–2519 (2001).
Hochheimer, A., Zhou, S., Zheng, S., Holmes, M.C. & Tjian, R. TRF2 associates with DREF and directs promoter-selective gene expression in Drosophila. Nature 420, 439–445 (2002).
Holmes, M.C. & Tjian, R. Promoter-selective properties of the TBP-related factor TRF1. Science 288, 867–870 (2000).
Isogai, Y., Keles, S., Prestel, M., Hochheimer, A. & Tjian, R. Transcription of histone gene cluster by differential core-promoter factors. Genes Dev. 21, 2936–2949 (2007).
Shiraki, T. et al. Cap analysis gene expression for high-throughput analysis of transcriptional starting point and identification of promoter usage. Proc. Natl. Acad. Sci. USA 100, 15776–15781 (2003).
Suzuki, Y. & Sugano, S. Construction of a full-length enriched and a 5′-end enriched cDNA library using the oligo-capping method. Methods Mol. Biol. 221, 73–91 (2003).
Zhang, Z. & Dietrich, F.S. Mapping of transcription start sites in Saccharomyces cerevisiae using 5′ SAGE. Nucleic Acids Res. 33, 2838–2851 (2005).
Ahsan, B. et al. MachiBase: a Drosophila melanogaster 5′-end mRNA transcription database. Nucleic Acids Res. 37, D49–D53 (2009).
Carninci, P. et al. Genome-wide analysis of mammalian promoter architecture and evolution. Nat. Genet. 38, 626–635 (2006).
Suzuki, H. et al. The transcriptional network that controls growth arrest and differentiation in a human myeloid leukemia cell line. Nat. Genet. 41, 553–562 (2009).
Valen, E. et al. Genome-wide detection and analysis of hippocampus core promoters using DeepCAGE. Genome Res. 19, 255–265 (2009).
Affymetrix ENCODE Transcriptome Project & Cold Spring Harbor Laboratory ENCODE Transcriptome Project. Post-transcriptional processing generates a diversity of 5′-modified long and short RNAs. Nature 457, 1028–1032 (2009).
Esteban, J.A., Salas, M. & Blanco, L. Fidelity of phi 29 DNA polymerase. Comparison between protein-primed initiation and DNA polymerization. J. Biol. Chem. 268, 2719–2726 (1993).
Carninci, P. et al. The transcriptional landscape of the mammalian genome. Science 309, 1559–1563 (2005).
Wilhelm, B.T. et al. Dynamic repertoire of a eukaryotic transcriptome surveyed at single-nucleotide resolution. Nature 453, 1239–1243 (2008).
Ohler, U., Liao, G.C., Niemann, H. & Rubin, G.M. Computational analysis of core promoters in the Drosophila genome. Genome Biol. 3, 0087 (2002).
Purnell, B.A., Emanuel, P.A. & Gilmour, D.S. TFIID sequence recognition of the initiator and sequences farther downstream in Drosophila class II genes. Genes Dev. 8, 830–842 (1994).
Burke, T.W. & Kadonaga, J.T. Drosophila TFIID binds to a conserved downstream basal promoter element that is present in many TATA-box-deficient promoters. Genes Dev. 10, 711–724 (1996).
FitzGerald, P.C., Sturgill, D., Shyakhtenko, A., Oliver, B. & Vinson, C. Comparative genomics of Drosophila and human core promoters. Genome Biol. 7, R53 (2006).
Sandelin, A. et al. Mammalian RNA polymerase II core promoters: insights from genome-wide studies. Nat. Rev. Genet. 8, 424–436 (2007).
Megraw, M., Pereira, F., Jensen, S.T., Ohler, U. & Hatzigeorgiou, A.G. A transcription factor affinity-based code for mammalian transcription initiation. Genome Res. 19, 644–656 (2009).
Ng, P. et al. Gene identification signature (GIS) analysis for transcriptome characterization and genome annotation. Nat. Methods 2, 105–111 (2005).
Rach, E.A., Yuan, H.Y., Majoros, W.H., Tomancak, P. & Ohler, U. Motif composition, conservation and condition-specificity of single and alternative transcription start sites in the Drosophila genome. Genome Biol. 10, R73 (2009).
Akhtar, W. & Veenstra, G.J. TBP2 is a substitute for TBP in Xenopus oocyte transcription. BMC Biol. 7, 45 (2009).
Gazdag, E. et al. TBP2 is essential for germ cell development by regulating transcription and chromatin condensation in the oocyte. Genes Dev. 23, 2210–2223 (2009).
Shibuya, T., Tange, T.O., Sonenberg, N. & Moore, M.J. eIF4AIII binds spliced mRNA in the exon junction complex and is essential for nonsense-mediated decay. Nat. Struct. Mol. Biol. 11, 346–351 (2004).
Schoenberg, D.R. & Maquat, L.E. Re-capping the message. Trends Biochem. Sci. 34, 435–442 (2009).
Core, L.J., Waterfall, J.J. & Lis, J.T. Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science 322, 1845–1848 (2008).
Nechaev, S. et al. Global analysis of short RNAs reveals widespread promoter-proximal stalling and arrest of Pol II in Drosophila. Science 327, 335–338 (2010).
Manak, J.R. et al. Biological function of unannotated transcription during the early development of Drosophila melanogaster. Nat. Genet. 38, 1151–1158 (2006).
Tweedie, S. et al. FlyBase: enhancing Drosophila Gene Ontology annotations. Nucleic Acids Res. 37, D555–D559 (2009).
Boyle, A.P., Guinney, J., Crawford, G.E. & Furey, T.S. F-Seq: a feature density estimator for high-throughput sequence tags. Bioinformatics 24, 2537–2538 (2008).
Barrett, T. et al. NCBI GEO: archive for high-throughput functional genomic data. Nucleic Acids Res. 37, D885–D890 (2009).
Hertz, G.Z. & Stormo, G.D. Identifying DNA and protein patterns with statistically significant alignments of multiple sequences. Bioinformatics 15, 563–577 (1999).
Wilson, R.J., Goodman, J.L. & Strelets, V.B. FlyBase: integration and improvements to query tools. Nucleic Acids Res. 36, D588–D593 (2008).
Acknowledgements
We thank D. MacAlpine and S. Powell for their help in collecting fly embryos, B. Xie and Y. Bao for optimizing paired-end sequencing procedure and J. Kadonaga for helpful comments on the manuscript. This work was funded by US National Institutes of Health (R01 HG004065 to U.O. and J.Z.) and National Science Foundation (MCB0822033 to J.Z. and U.O.).
Author information
Authors and Affiliations
Contributions
U.O. and J.Z. oversaw the project. T.N., S.S. and J.Z. designed and performed experiments related to PEAT library construction, quality control and various validation assays. E.P.S. provided fly stock for collecting embryos. Y.G. performed Illumina sequencing. D.L.C., E.A.R. and U.O. analyzed data. T.N., D.L.C., E.A.R., U.O. and J.Z. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–16, Supplementary Tables 1–9 and Supplementary Results (PDF 1319 kb)
Supplementary Data 1
Genomic information on all TSS clusters. (XLS 818 kb)
Rights and permissions
About this article
Cite this article
Ni, T., Corcoran, D., Rach, E. et al. A paired-end sequencing strategy to map the complex landscape of transcription initiation. Nat Methods 7, 521–527 (2010). https://doi.org/10.1038/nmeth.1464
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1464
This article is cited by
-
Dynamic regulation of N6,2′-O-dimethyladenosine (m6Am) in obesity
Nature Communications (2021)
-
Epigenetic regulation of spurious transcription initiation in Arabidopsis
Nature Communications (2020)
-
Geochemical and Petrographic Analyses of the Cambrian Oncoids of the North China Platform: Implications for Their Paleogeography and Paleoenvironment
Arabian Journal for Science and Engineering (2020)
-
Petrographic and rare earth elemental characteristics of Cambrian Girvanella oncoids exposed in the North China Platform: Constraints on forming mechanism, REE sources, and paleoenvironments
Arabian Journal of Geosciences (2020)
-
Eukaryotic core promoters and the functional basis of transcription initiation
Nature Reviews Molecular Cell Biology (2018)